Abstract
Breast cancer is one of the most widespread forms of cancer that affects a significant portion of the female population today. Its early detection and subsequent treatment can be life saving. However, it is difficult clinically and computationally to detect breast cancer and its subtypes in their early stages. On the other hand, Next Generation Sequencing (NGS) techniques have significantly accelerated the process of mapping the human genomes by providing high-throughput expression data of RNA. In this work, we study such NGS based expression data of mRNAs with the clinical data in order to (a) rank the genes based on their importance in survival of breast cancer subtypes and (b) find the relation between the up/down regulation of genes and survival probability of a population. In this regard, first volcano plot is used to find the differentially expressed genes for each subtype, and second, such genes are used to perform the Kaplan-Meier survival analysis with the integration of mRNA expression and clinical data to rank the genes by their importance in survival of breast cancer subtypes. These genes are ranked based on the p-value and significant genes are filtered out by considering the cut-off as p-value \(< 0.05\) for each breast cancer subtype. In our analysis, we have found a relation between gene regulation and survival probability, e.g. up and down regulated genes of a population show low rate of survival of that population. Moreover, for the biological significance, PPI network and KEGG Pathway analysis are conducted on a common set of genes that are present in all subtypes. The datasets, code and supplementary materials of this work are provided online (http://www.nitttrkol.ac.in/indrajit/projects/mrna-survival-breastcancer-subtypes/).
You have full access to this open access chapter, Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
Breast cancer is one of the most prolific types of cancer in females worldwide, and the most common cause of cancer related deaths regardless of race and ethnicity, responsible in 2018 for more than 626 thousand deaths worldwide [1]. One of the difficulties in seeking treatment is that breast cancer is a heterogeneous form of cancer with multiple subtypes, each requiring different treatment [3]. Thus, it is crucial to identify its subtype-specific biomarkers. To this end, computational approaches are being used, and tools and databases are being developed, yet it is still a challenging task, both computationally and clinically. The RNA-Seq method utilizes the Next Generation Sequencing (NGS) techniques to rapidly sequence and analyze whole transcriptome for potential breast cancer biomarkers. One of the possible biomarker types is mRNA: a type of RNA molecule which carries genetic information from the DNA nucleus to the ribosome, where the amino acid sequence of the protein products of gene expression is specified by mRNA’s sequence. The identification of subtype-wise biomarkers using NGS data is crucial for decision making in case of disease management, yet it is difficult to analyze this data because of its high dimensionality and complexity. However, some tools like Oncomine [8], BioXpress [9], etc. have been created to facilitate the search for biomarkers. Another class of tools can perform survival analysis. These include, for example, Kaplan–Meier Plotter [4], BreastMark [6], etc. Integrated solutions like KM-express [2] start to emerge that make use of NGS data and combine many approaches, are being developed as well.
This fact motivated us to study the NGS based high-throughput expression data of mRNAs. In this regard, we first analyze the datasets of breast cancer subtypes viz. Luminal A (LA), Luminal B (LB), HER2-Enriched (HER2-E), Basal-Like (BL) with Control samples separately in order to find the differentially expressed up and down regulated genes using volcano plot. Thereafter, Kaplan-Meier (KM) survival analysis [5] is performed on these genes to rank them based on p-value. Moreover, the significant genes from the ranked lists are identified based on the criterion, p-value \(< 0.05\), for each subtype. From the analysis, it is observed that the survival rate of a population is directly proportional with the up and down regulation of genes of same population. Furthermore, the common set of significant up and down regulated genes for all subtypes are considered to see their biological significance in terms of Protein-Protein Interaction (PPI) network and KEGG Pathway Analysis.
2 Material and Method
This section describes briefly the preparation of the datasets and the proposed framework.
2.1 Dataset Preparation
The RNA-Seq data in form of RSEM (RNA-Seq by Expectation Maximization) contains the expression values of mRNAs. It is obtained from The Cancer Genome Atlas (TCGA) [10]. The data is normalized by transforming to \(log_2\) scale. The dataset contains 17,987 mRNAs of 326 patients. Furthermore, the breast cancer subtype information is collected from [7]. Only the mRNAs that contains less than 1% zero expression values are kept, while the others are discarded in order to minimize the missing data. After completing this step, we obtain 14,831, 14,330, 14,609 and 14,267 mRNAs for LA, LB, HER2-E and BL subtype respectively. The number of samples, average age of patients and average follow-up days in each subtype are mentioned in Table 1.
2.2 Method
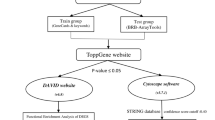
The proposed framework is discussed below and shown in Fig. 1.
Identification of Differentially Expressed Genes Using Volcano Plot: For the purpose of identifying up and down regulated genes, volcano plot technique is used. Volcano plot identifies differential genes using the t-test and fold-change (FC) methods. It plots log2 of fold-change value on the X-axis against -log10 of p-value from the t-test on the Y-axis. Genes having positive and negative fold change are called up and down regulated genes respectively. In the present experiment, the up and down regulated genes are obtained using volcano plot for each subtype.
Survival Analysis Using Kaplan-Meier Method: In order to assess the impact of mRNA expression on patient survival, we use the Kaplan-Meier method, which uses the patient status and days to last follow up information, to estimate the survival function S(t), which represents the probability of an individual from a given population to be alive at time t. The log-rank test can then be used to compare survival curves between groups. In our experiment, for each subtype, we divide the samples into high-expression and low-expression groups, by median split. Then we compute the subtype-wise p-value of the genes. This gives a p-value to each gene for each subtype. Moreover, KM plots are generated for such genes in each subtype. Thereafter, the genes are ranked subtype-wise by their p-value in ascending order.
Identification of Genes Common to All Subtypes: After obtaining subtype-wise rankings of genes, we seek to find those genes, that are important in all subtypes. This is performed by selecting those genes with p-value \(< 0.05\), hence they are statistically significant for the survival of patients. These four lists of genes are then used to plot a Venn diagram to find the significant common genes that are present in all four subtypes.
3 Experimental Results
3.1 Experimental Testbed
The experiment has been conducted with the use of MATLAB 2018a, Pandas 0.23 and Numpy 1.14 in Python 3.6.5. A computer with Intel i7 processor with 8 cores and 8 GB RAM are used for all computational purposes. An online toolFootnote 1 is used to plot the Venn diagram.
3.2 Results
The proposed framework has been used on the breast cancer subtype-specific datasets. As a result, volcano plot provides the subtype-wise up and down regulated genes as shown in Fig. 2. The number of obtained differentially expressed genes are reported in Table 2. These genes are used further for Kaplan-Meier estimator in order to perform the survival analysis. Table 3 reports top ten up and down regulated genes for each subtype while rest are mentioned in the supplementary. A number of genes well known for their association with breast cancer are seen in the table, such as CREB3L4, CD24, PYCR1, MAST1, etc. The listed genes in each subtype are used to find the common genes in all four subtypes and are shown in the Venn diagrams in Figs. 3(a) and (b). As a result, 149 and 224 common genes have been obtained.
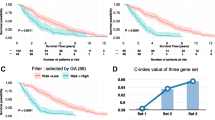
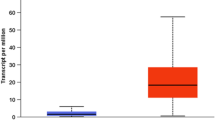
In order to find the relation between the up/down regulation of genes and survival probability of a population, KM plot of top genes and boxplot of expression for each subtype are shown in Fig. 4. It is found from the figure that the survival probability of a population is very low for the up regulated genes viz. CREB3L4, CELSR3, CD24 and MAST1. Similarly, for the down regulated genes viz. RBPMS2, CCRL1, RNF157 and SLC4A4, the same low survival probability of a population is observed. This is true for the up and down regulated genes.
Furthermore, the common 149 up and 224 down regulated genes are used to find their targeted Transcription Factors (TFs) from the TRRUST database. Using these TFs, a Protein-Protein Interaction network is plotted as shown in Fig. 5(a) with the help of STRING database. Here, a PPI enrichment p-value \(< 10^{-16}\) is obtained showing the significance of the network. The degree of top ten nodes of Fig. 5(a) is shown in Fig. 5(b). It is seen that TFs that are well known for their association with breast cancer such as ESR1, HDAC1, AR etc. are found on top. Apart from this, KEGG pathway analysis has also been performed using Enrichr tool for such common up and down regulated genes. The obtained pathways, as shown in Fig. 6 are Cell cycle, AMPK signaling pathway, Breast cancer, cGMP-PKG signaling pathway, cAMP signaling pathway, Thyroid cancer, etc. These pathways are significant for breast cancer. Therefore, the obtained genes are significant as they belong to the same pathways of breast cancer.
4 Conclusion
Detection of breast cancer subtypes in early stages is still a challenging task. It is thus important to identify biomarkers that can accurately help in early detection. NGS techniques can provide opportunities for new biomarker detection by providing high-throughput expression data of mRNA. We presented a systematic approach to identify genes as potential biomarkers for specific subtype of breast cancer using statistical analysis of population of patients’ survival data. In order to show our identified sets of genes to be enriched in breast cancer related pathways and functions, we validated them biologically. Moreover, it is seen that the up and down regulation of genes directly affect the survival probability of a population. As a scope of further research, this work can be extended to understand the biological mechanism, e.g., the effect of miRNAs, DNA Methylations and Enhancers to change the expression of such genes which are responsible for the change of survival probability of a population.
References
Bray, F., Ferlay, J., Soerjomataram, I., Siegel, R.L., Torre, L.A., Jemal, A.: Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 68(6), 394–424 (2018)
Chen, X., Miao, Z., Divate, M., Zhao, Z., Cheung, E.: KM-express: an integrated online patient survival and gene expression analysis tool for the identification and functional characterization of prognostic markers in breast and prostate cancers. Database (2018)
Dai, X., et al.: Breast cancer intrinsic subtype classification, clinical use and future trends. Am. J. Cancer Res. 5(10), 2929–2943 (2015)
Györffy, B., et al.: An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1,809 patients. Breast Cancer Res. Treat. 123(3), 725–731 (2010)
Jager, K.J., van Dijk, P.C., Zoccali, C., Dekker, F.W.: The analysis of survival data: the Kaplan-Meier method. Kidney Int. 74(5), 560–565 (2008)
Madden, S.F., et al.: BreastMark: an integrated approach to mining publicly available transcriptomic datasets relating to breast cancer outcome. Breast Cancer Res. 15(4), R52 (2013)
Cancer Genome Atlas Network, et al.: Comprehensive molecular portraits of human breast tumours. Nature 490(7418), 61–70 (2012)
Rhodes, D.R., et al.: Oncomine 3.0: genes, pathways, and networks in a collection of 18,000 cancer gene expression profiles. Neoplasia 9(2), 166–180 (2007)
Wan, Q., et al.: BioXpress: an integrated RNA-seq-derived gene expression database for pan-cancer analysis. Database (2015)
Weinstein, J.N., et al.: The cancer genome atlas pan-cancer analysis project. Nat. Genet. 45(10), 1113–1120 (2013)
Acknowledgements
This work has been supported by the Polish National Science Centre (2014/15/ B/ST6/05082), Foundation for Polish Science (TEAM to DP) and by the grant from the Department of Science and Technology, India under Indo-Polish/Polish-Indo project No.: DST/INT/POL/P-36/2016. Moreover, the work was cosupported by grant 1U54DK107967-01 “Nucleome Positioning System for Spatiotemporal Genome Organization and Regulation” within 4DNucleome NIH program.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this paper
Cite this paper
Saha, I. et al. (2019). Survival Analysis with the Integration of RNA-Seq and Clinical Data to Identify Breast Cancer Subtype Specific Genes. In: Deka, B., Maji, P., Mitra, S., Bhattacharyya, D., Bora, P., Pal, S. (eds) Pattern Recognition and Machine Intelligence. PReMI 2019. Lecture Notes in Computer Science(), vol 11942. Springer, Cham. https://doi.org/10.1007/978-3-030-34872-4_16
Download citation
DOI: https://doi.org/10.1007/978-3-030-34872-4_16
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-34871-7
Online ISBN: 978-3-030-34872-4
eBook Packages: Computer ScienceComputer Science (R0)