Abstract
Solid tumors are heterogeneous tissues composed of a mixture of cells and have special tissue architectures. However, cellular heterogeneity, the differences in cell types are generally not reflected in molecular profilers or in recent histopathological image-based analysis of lung cancer, rendering such information underused. This paper presents the development of a computational approach in H&E stained pathological images to quantitatively describe cellular heterogeneity from different types of cells. In our work, a deep learning approach was first used for cell subtype classification. Then we introduced a set of quantitative features to describe cellular information. Several feature selection methods were used to discover significant imaging biomarkers for survival prediction. These discovered imaging biomarkers are consistent with pathological and biological evidence. Experimental results on two lung cancer data sets demonstrated that survival models bsuilt from the clinical imaging biomarkers have better prediction power than state-of-the-art methods using molecular profiling data and traditional imaging biomarkers.
This work was partially supported by NSF IIS-1423056, CMMI-1434401, CNS-1405985.
You have full access to this open access chapter, Download conference paper PDF
Similar content being viewed by others
Keywords
- Squamous Cell Carcinoma
- Feature Selection Method
- Subtype Cell
- Cell Segmentation
- National Lung Screening Trial
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 Introduction
Lung cancer is the second most common cancer in both men and women. The non-small cell lung cancer (NSCLC) is the majority (80–85%) of lung cancer and two major NSCLC types are Adenocarcinoma (ADC) (40 %) and Squamous Cell Carcinoma (SCC) (25–30%).Footnote 1 The 5-year survival rate of lung cancer (17.7 %) is still significantly lower than most other cancers.Footnote 2 Therefore, predicting clinical outcome of lung cancer is an active field in today’s medical research.
Molecular profiling is a technique to query the expression of thousands of molecular data simultaneously. The information derived from molecular profiling can be used to classify tumors, and help to make clinical decisions [6, 15].
However, tumor microenvironment is a complex milieu that includes not only the cancer cells but also the stromal cells and immune cells. All this “extra” genomic information may muddle results and therefore make molecular analysis a challenging task for cancer prognosis [14].
Recently, Arne Warth et al. [10] showed that there exists connections between lung tumor morphology and prognosis. Advances in imaging have created a good chance to study such information using hispathological images to help tumor diagnosis [1, 14, 16]. In general, a pathologist can visually examine stained slides of a tumor to discover imaging biomarkers that can be used for diagnosis. For example, Fig. 1(A) shows two pathology images from ADC lung cancer patients. (A) is an image from one patient who had the worse survival outcome while (B) is captured from a patient who lived longer. A distinct pattern can be found in Fig. 1(A) as the more advanced tumor cells clustered in a larger more condensed area indicates a worse survival outcome than Fig. 1(B) where tumor cells are scattered into a smaller region with lymphocytes and stromal cells nearby. However, the process of manually searching for such imaging biomarkers is very labor-intensive and cannot be easily scaled to large number of samples. Wang et al. [9] proposed an automated image analysis to help pathologists find imaging biomarkers that could identify lung cancer survival characteristics. However, their results still remain some issues. First, they collected ADC and SCC samples together when looking for imaging biomarkers. According to lung cancer pathology [8], the two major types of NSCLC (ADC and SCC) are generally regarded as two different types of disease due to their distinct molecular mechanisms and pathological patterns. Second, spatial variations between the different types of cells (ADC and SCC) are associated with survival outcomes [14]. However, the study in [9] adopted a traditional cell segmentation which was unable to classify cell subtypes and achieve clinically interpretable imaging biomarkers in lung cancer.
In this paper, we introduced a computational image analysis to discover clinically interpretable imaging biomarkers for lung cancer survival prediction. Experiments on two lung cancer cohorts demonstrate that: (1) Two major subtypes of NSCLC should be treated separately since they have different key imaging biomarkers. (2) Spatial distribution of subtype cells are informative imaging biomarkers for lung cancer survival prediction. (3) The proposed framework can better describe tumor morphology and can provide powerful survival analysis than the state-of-the-art method with molecular profiling data.
2 Methodology
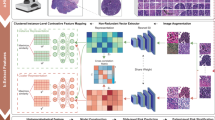
An overview of our method is presented in Fig. 2. An expert pathologist first labels regions of tissues. Several image tiles are extracted from the interested regions. Then a deep learning approach is applied to detect different types of cells (tumor, stroma and lymphocyte cells). A set of quantitative descriptors is used to cover granularity and subtype cellular heterogeneity. Our image analysis pipeline automatically segments H&E stained images, classifies cellular components into three categories (tumor, lymphocyte, stromal), and extracts features based on cell segmentation and detection results. Feature selection methods are used to find important features (image markers). These imaging biomarkers can then be applied for building survival models to predict patient clinical outcomes.
2.1 Deep Learning Approach for Cell Subtype Classification
The architecture of network can be seen in Fig. 3. Different cell types (cancer cells, stromal cells, lymphocytes) play different roles in tumor growth and metastasis, and accurately classifying cell types is a critical step to better characterization of tumor growth and outcome prediction [2, 14].
Due to the large appearance variation and high complexity of lung cancer tissues, traditional machine learning approaches do not clearly distinguish or define the different cell types. Motivated by recent deep learning method for cell detection [11, 12], we developed a two partially shared-weighted deep convolution neural networks (DCNNs) for cell subtype detection. The ground truth for cell subtype classification was annotated by an experienced pathologist. Then we built training samples with two annotations, one is for cell/non-cell classification and the other is for subtype cell. Each patch size is 40*40. We collected 48562 and 24281 patches for cell/noncell and subtype cell classification, respectively. Sparse kernels [5] are applied in the two DCNN models to eliminate all the redundant calculations for acceleration. In the final step, those two DCNN models are integrated into one model to achieve subtype cell detection. More details can be found in our research web page: http://ranger.uta.edu/~huang/R_Lung.htm.
2.2 Quantitative Imaging Feature Extraction
Motivated by [9, 13], three groups of cellular features were extracted using subtype cell detection results. These features cover cell-level information (e.g., appearance and shapes) of individual subtype cells and also texture properties of background tissue regions.
Group 1: Geometry Features. Geometry properties are calculated for each segmented subtype cell, including area, perimeter, circularity, major-minor axis ratio. Zernike moments were also applied on each type of cells. When combined with different tiles, we calculated mean, median and std. of each feature with a total of 564 features.
Group 2: Texture Features. This group of features contains Gabor “wavelet” features, co-occurrence matrix and granularity to measure texture properties of objects (e.g., cells and tissues), resulting in 1685 texture features.
Group 3: Holistic Statistics. The four holistic statistics include overall information like the total area, perimeter, number and the corresponding ratio of each subtype cells.
2.3 Imaging Biomarkers Discovery
The objective of this step is to find important imaging biomarkers since not all features were highly correlated with patients’ survival outcomes. Different from traditional applications, selecting features in survival analysis is a censoring problem (subjects are censored if they are not followed up or the study ends before they die). In this study, we built the predictive models using two well-established types of methods: (1) the multivariate Cox proportional hazards model with L1 penalized log partial likelihood (Lasso) [7] or component-wise likelihood based boosting (CoxBoost) [3] for feature selection, and (2) random survival forest (RSF) [4]. Because of the high dimension of the image features, we first applied univariate Cox regression and kept those with Wald test p value less than 0.05. Then we conducted the feature selection on a small candidates set for survival model to improve the speed.
3 Experimental Results
3.1 Materials
We focused on two widely used lung cancer dataset NLST (National Lung Screening Trial)Footnote 3 and TCGA Data PortalFootnote 4. Both dataset contain complete patients’ pathology images with survival and clinical information while TCGA cohorts can provide additional molecular profiling data. In NLST, we collected 144 ADC and 113 SCC patients. In TCGA, we focused on SCC case and collected 106 patients with four types of molecular data including: Copy number variation (CNV), mRNA, microRNA and protein expression (RPPA). To examine whether imaging biomarkers from the proposed framework can achieve better predictions than traditional imaging biomarkers and molecular profiling data (biomarkers), we evaluated with two state-of-the-arts framework in lung cancer [9, 15].
3.2 Imaging Biomarker Discovery for Survival Analysis
ADC vs SCC samples. In this experiment, we followed the framework in [9] and investigated differences in imaging biomarkers selecting from the set of ADC and SCC markers, and combining ADC and SCC markers together. To ensure the robustness of selection, we resampled the whole dataset with replacements and performed the boosting feature selection procedure [3] and calculated the frequency of choosing a variable. Figure 4 shows that key features (high frequencies shown in the green rectangle) chosen from the combination set are very different from those of ADC and SCC, respectively. These differences convinced us the prognosis models for ADC and SCC should be developed separately. This discovery verified the evidence in lung cancer pathology, that lung cancer subtypes are highly heterogeneous and cannot be combined together.
For ADC and SCC, selected features include information about suptype cell distributions, cell shape and granularity. Among them, subtype cell distributions and granularity have been confirmed to be associated with survival outcomes [8, 14]. To test these imaging biomarkers, we built multivariate Cox regression using the top 50 selected features on testing sets (47 for ADC and 37 for SCC). Figure 5 presents the predictive power on a partitioning into two groups on testing set (a–b for ADC and c–d for SCC). A significant difference (Wald-Test) in survival times can be seen in Fig. 5(a),(c). It demonstrates that discovered imaging biomarkers which cover subtype cell distributions and granularity are more often associated with survival outcomes than traditional imaging biomarkers.
Kaplan-Meier survival curves of two groups on testing set. The x axis is the time in days and the y axis denotes the probability of overall survival. (a,c) are from the framework developed in this research, while (b,d) are using features from [9].
Then we randomly divided the whole set to 50 splits (2/3 for training, 1/3 for testing). Each feature selection method performed 10-fold cross validation for parameter optimization. Figure 6 shows the concordance index (C-index) results of the two methods on ADC and SCC set.
The C-index is a nonparametric measurement to quantify the discriminatory power of a predictive model: 1 indicates perfect prediction accuracy, and a C-index of 0.5 is as good as a random guess. From Fig. 6, it can see the higher median C-index of the discovered imaging markers in both cases with different survival models. This illustrates the robustness of the proposed method since the discovered imaging biomarkers are highly associated with tumor growth and survival outcomes.
3.3 Comparison of Survival Model with Imaging and Molecular Data
To examine whether the proposed imaging biomarkers can provide better prediction power than traditional molecular data, we conducted experiments on TCGA LUSC cohort following the recent study [15]. We applied 50 random splits and assessed the C-index of a model built from the individual imaging and molecular data sets alone. Figure 7A presents the highest median C-index value of survival models built on the discovered imaging biomarkers. When each type of data integrates with clinical variables (“+” means the integration), all prediction accuracies increase while the proposed method still has the best results (Fig. 7B). It verified the discovered imaging biomarkers can better describe tumor morphology which enabled the proposed framework to have the best predictions for survival analysis.
4 Conclusions
In this paper, we investigated subtype cell information and found that they have useful patterns for predicting patients survival. These results are consistent with recent study in lung cancer pathology [10]. Extensive experiments have been conducted to demonstrate that imaging biomarkers from subtype cell information can better describe tumor morphology and provide more accurate prediction than state-of-the-art method using imaging and molecular profilers. In the future, we will try to find more quantitative measurements to better describe tumor morphology and further improve the prediction performances.
References
Barker, J., Hoogi, A., Depeursinge, A., Rubin, D.L.: Automated classification of brain tumor type in whole-slide digital pathology images using local representative tiles. Med. Image Anal. 30, 60–71 (2016)
Beck, A.H., Sangoi, A.R., Leung, S., Marinelli, R.J., Nielsen, T.O., van de Vijver, M.J., West, R.B., van de Rijn, M., Koller, D.: Systematic analysis of breast cancer morphology uncovers stromal features associated with survival. Sci. Transl. Med. 3(108), 108ra113 (2011)
Binder, H., Schumacher, M.: Allowing for mandatory covariates in boosting estimation of sparse high-dimensional survival models. BMC Bioinform. 9(1), 1–10 (2008)
Ishwaran, H., Kogalur, U.B., Blackstone, E.H., Lauer, M.S.: Random survival forests. Ann. Appl. Stat. 2(3), 841–860 (2008)
Li, H., Zhao, R., Wang, X.: Highly efficient forward and backward propagation of convolutional neural networks for pixelwise classification. arXiv preprint arXiv:1412.4526 (2014)
Shedden, K., Taylor, J.M., Enkemann, S.A., Tsao, M.S., Yeatman, T.J., Gerald, W.L., Eschrich, S., Jurisica, I., Giordano, T.J., Misek, D.E., et al.: Gene expression-based survival prediction in lung adenocarcinoma: a multi-site, blinded validation study. Nat. Med. 14(8), 822–827 (2008)
Tibshirani, R.: Regression shrinkage and selection via the lasso. J. Roy. Stat. Soc.: Ser. B (Methodol.) 58, 267–288 (1996)
Travis, W.D., Harris, C.: Pathology and genetics of tumours of the lung, pleura, thymus and heart (2004)
Wang, H., Xing, F., Su, H., Stromberg, A., Yang, L.: Novel image markers for non-small cell lung cancer classification and survival prediction. BMC Bioinform. 15(1), 310 (2014)
Warth, A., Muley, T., Meister, M., Stenzinger, A., Thomas, M., Schirmacher, P., Schnabel, P.A., Budczies, J., Hoffmann, H., Weichert, W.: The novel histologic international association for the study of lung cancer/american thoracic society/european respiratory society classification system of lung adenocarcinoma is a stage-independent predictor of survival. J. Clin. Oncol. 30(13), 1438–1446 (2012)
Xie, Y., Xing, F., Kong, X., Su, H., Yang, L.: Beyond classification: structured regression for robust cell detection using convolutional neural network. In: Navab, N., Hornegger, J., Wells, W.M., Frangi, A.F. (eds.) MICCAI 2015. LNCS, vol. 9351, pp. 358–365. Springer, Heidelberg (2015). doi:10.1007/978-3-319-24574-4_43
Xu, Z., Huang, J.: Efficient lung cancer cell detection with deep convolution neural network. In: Wu, G., Coupé, P., Zhan, Y., Munsell, B., Rueckert, D. (eds.) Patch-MI 2015. LNCS, vol. 9467, pp. 79–86. Springer, Heidelberg (2015). doi:10.1007/978-3-319-28194-0_10
Yao, J., Ganti, D., Luo, X., Xiao, G., Xie, Y., Yan, S., Huang, J.: Computer-assisted diagnosis of lung cancer using quantitative topology features. In: Zhou, L., Wang, L., Wang, Q., Shi, Y. (eds.) MLMI 2015. LNCS, vol. 9352, pp. 288–295. Springer, Heidelberg (2015). doi:10.1007/978-3-319-24888-2_35
Yuan, Y., Failmezger, H., Rueda, O.M., Ali, H.R., Gräf, S., Chin, S.F., Schwarz, R.F., Curtis, C., Dunning, M.J., Bardwell, H., et al.: Quantitative image analysis of cellular heterogeneity in breast tumors complements genomic profiling. Sci. Transl. Med. 4(157), 157ra143 (2012)
Yuan, Y., Van Allen, E.M., Omberg, L., Wagle, N., Amin-Mansour, A., Sokolov, A., Byers, L.A., Xu, Y., Hess, K.R., Diao, L., et al.: Assessing the clinical utility of cancer genomic and proteomic data across tumor types. Nat. Biotechnol. 32(7), 644–652 (2014)
Zhu, X., Yao, J., Luo, X., Xiao, G., Xie, Y., Gazdar, A., Huang, J.: Lung cancer survival prediction from pathological images and genetic data - an integration study. In: 2016 IEEE 13th International Symposium on Biomedical Imaging (ISBI), pp. 1173–1176 (2016)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing AG
About this paper
Cite this paper
Yao, J., Wang, S., Zhu, X., Huang, J. (2016). Imaging Biomarker Discovery for Lung Cancer Survival Prediction. In: Ourselin, S., Joskowicz, L., Sabuncu, M., Unal, G., Wells, W. (eds) Medical Image Computing and Computer-Assisted Intervention – MICCAI 2016. MICCAI 2016. Lecture Notes in Computer Science(), vol 9901. Springer, Cham. https://doi.org/10.1007/978-3-319-46723-8_75
Download citation
DOI: https://doi.org/10.1007/978-3-319-46723-8_75
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-46722-1
Online ISBN: 978-3-319-46723-8
eBook Packages: Computer ScienceComputer Science (R0)