Abstract
Automated retinal blood vessel segmentation plays an important role in the diagnosis and treatment of various cardiovascular and ophthalmologic diseases. In this paper, an unsupervised algorithm based on denoising and mathematical morphology is proposed to extract blood vessels from color fundus images. Specifically, our method consists of the following steps: (i) green channel extraction; (ii) non-local means denoising; (iii) vessel vasculature enhancement by means of a sum of black top-hat transforms; and (iv) image thresholding for the final segmentation. This method stands out for its simplicity, robustness to parameters change and low computational complexity. Experimental results on the publicly available database DRIVE show our method to be effective in segmenting blood vessels, achieving an accuracy comparable to that of unsupervised state-of-the-art methodologies.
You have full access to this open access chapter, Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
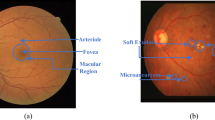
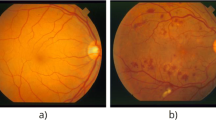
Retinal vessel structure is of great interest in ophthalmology since its characterization in terms of geometric features can provide indicative measures of the presence and severity of different cardiovascular and ophthalmologic diseases such as arteriosclerosis, diabetes, hypertension, and glaucoma [14]. The analysis of the change in morphological features of retinal blood vessels such as length, width, and branching angle, can help in the early detection of these diseases and in the identification of the optimal treatment in order to prevent visual loss [17]. Vessel segmentation is a crucial step for the assessment of these morphological properties. Because of the complexity of the vascular network, manual segmentation is time consuming and difficult: a feasible solution is the automation of the segmentation process. However, the automated segmentation of retinal blood vessels is a challenging task, mainly due to the width variability of vessels and to the low quality of retinal images that are, in general, noisy, unevenly illuminated, and of poor contrast [20]. In recent years, several methods have been proposed for blood vessel segmentation in color fundus images and they can be categorized as either supervised or unsupervised methods.
Supervised methods use hand-crafted or automatically extracted features to train a classifier in order to discriminate vessels and non-vessels pixels. A k-NN approach is used by Niemeijer et al. [23] and Staal et al. [27] to classify the feature vectors that are constructed by a multiscale Gaussian filter and by a ridge detector, respectively. Marín et al. [19] used gray-level and moment invariant-based features to build a 7-D feature vector and utilized a multilayer feed forward neural network for training and classification. Cheng et al. [7] used a random forest to fuse the information encoded in the hybrid context-aware features. Recent works use Deep Convolutional Neural Networks to automatically extract features from fundus images and segment the retinal vasculature [18, 21, 26].
Unsupervised segmentation methods rely on finding inherent properties of retinal blood vessels, which can be applied to distinguish vessel pixels in retinal images. According to the image processing methodologies, these methods can be classified into four sub-categories: matched filtering, vessel tracking, model-based, and mathematical morphology approaches.
In matched filtering techniques the retinal image is convolved with a 2-D kernel, and the matched filter response indicates the presence of the vessel. Matched filter detection was proposed by Chaudhuri et al. [6]. In this method, the authors used 12 rotated versions of a 2-D Gaussian shaped template for searching vessel segments along all possible directions. Azzopardi et al. [2] proposed a novel technique that uses a non-linear trainable filter called B-COSFIRE, designed to detect bar-shaped structures such as blood vessels. Vessels segmentation is achieved by adding up the responses of two B-COSFIRE filters, namely symmetric and asymmetric, followed by thresholding. Tracking methods, also mentioned as exploratory algorithms, start by locating the vessel points used for tracing the vasculature. This procedure can be done either manually or automatically by measuring some local image properties. Most of the methods reported in the literature use Gaussian functions to characterize vessel profiles [8, 13]. This type of approaches can provide highly accurate vessel widths, but they are often unable to detect vessel segments that have no seed points. The model-based techniques apply explicit vessel models that can be regarded as vessel profile [16], parametric deformable [1], or geometric deformable models [28]. The vessel profile models approximate the vessel intensity profiles using mixture of Gaussians or the second-order derivative Gaussian, and other polynomial functions. The parametric and geometric deformable models are based on active contours models and on the theory of curve evolution geometric flows, respectively [11]. Mathematical morphology approaches exploit a priori information on the profile structure of vessels and involves the use of morphological operators. Zana and Klein [31] proposed a combination of morphological filters in order to enhance vessels and cross curvature evaluation to identify linearly coherent structures. Morphological processing for vessel segmentation was also reported in [22, 29]. Fraz et al. [10], proposed a robust technique for the extraction of the retinal vessels map using a combination of centerlines detection and morphological operators. An hybrid technique was proposed by Yank et al. [30], where the blood vessels were enhanced and the background removed with a morphological top-hat operation, then fuzzy clustering was applied to extract the blood vessels.
In this paper we present an unsupervised method for retinal blood vessel segmentation in color fundus images based on denoising and mathematical morphology. The denoising step is required to prepare images for the further vessel enhancement phase. We use the non-local means denoising method that preserves image details and contrast properties and allows to obtain well-contrasted images with a smoothed background that are suited for the application of mathematical morphology. The use of morphological operators for vessel enhancement builds on the observation that a vessel can be defined as a dark pattern with Gaussian-shape cross-section profile, piece-wise connected, and locally linear [9]. Based on this assumption, in our method a sum of black top-hat transforms is used to highlight vessels with respect to these morphological features.
2 Dataset
The proposed method has been tested on images of the publicly available database DRIVE. It contains 40 color images of the retina, with \(565\times 584\) pixels and 8 bits per color channel, captured from a Canon CR5 non-mydriatic 3CCD camera at \(45^{\circ }\) field of view, and saved in JPEG-format. Besides the color images, the database includes masks with the delimitation of a field of view (FOV) of approximately 540 pixels in diameter for each image, and binary images with the results of manual vessel segmentation. The 40 images were divided into a training set and a test set by the authors of the database. The results of the manual segmentation are available for all the images of the two sets. We used the training images in the vessel enhancement step to verify the robustness of our approach to different parameters configurations, and the test images to evaluate the performance measures.
3 Method
The proposed approach consists of four phases. First, the green channel is extracted from RGB images, because it shows better vessels-to-background contrast [11, 20]. Hence, from now on, we deal with grayscale images in which retinal vessels appear darker than the background. Then, the non-local means denoising algorithm is employed to remove the noise due to the image acquisition process [31]. Afterwards, a sum of black top-hat transforms is applied to emphasize the entire vasculature. A final thresholding is performed on the enhanced vessel map to obtain the desired segmentation. In the following section we describe in detail the last three steps of our approach. A diagram of the proposed approach is reported in Fig. 1.
3.1 Denoising: Non-local Means Algorithm
The success of an automatic retinal vessel segmentation algorithm is influenced by the presence of noise mainly caused by the digitization process [31]. Traditional denoising methods, like local smoothing and frequency domain filters, provide noise reduction and reconstruction of the main geometrical patterns of the image. These kind of filters are designed to remove only the high frequency noise of the image, but they often do not preserve fine details and structures that present the same frequency values as noise. To prevent this loss of information, in our work we apply the non-local means denoising. The main idea behind this denoising approach is that images contain a high level of redundancy, which means that it is possible to find similar intensity profiles either in adjacent or non-adjacent pixels. Thus, given a discrete noisy image f, for the pixel i, the denoised value NL(f(i)) is computed as a weighted average of all the pixels in the image:
where the family of weights \(\left\{ w(i,j)\right\} _{j}\) depends on the similarity between the pixels i and j, \(0\le w(i,j) \le 1\) and \(\sum _{j}w(i,j)=1\). Let \(N_{i}\) be a square neighborhood of size \(n\times n\) centered on i, the restriction of f to a neighborhood \(N_{i}\) is defined by \(f(N_{i})\):
The measure of the similarity between the intensity gray level vectors \(f(N_{i})\) and \(f(N_{j})\) is obtained by means of a Gaussian weighted Euclidean distance \(\left| \left| f(N_{i})-f(N_{j})\right| \right| ^{2}_{2,\sigma }\), which is a \(L_{2}\) norm convolved with a Gaussian kernel of standard deviation \(\sigma \). The weights are defined according to the following equation:
where Z(i) is a normalizing constant \(Z(i)=\sum _{j}e^{{-\frac{\left| \left| f(N_{i})-f(N_{j})\right| \right| ^{2}_{2,\sigma }}{h^{2}}}}\). This results in giving larger weights to those neighborhoods that are similar in terms of gray levels. The variable h in Eq. 3 is a parameter which controls the decay of the exponential function and consequently the decay of the weights. In order to reduce the computational cost, it shall be defined a \(m\times m\) search window M on the image f, in which the similarity of the neighborhoods is evaluated. In this way, the denoised value NL(f(i)) is computed as a weighted average of all the pixels of the search window. In our application, we set all the parameters to the values suggested in the default implementation of the algorithm [5]. Therefore \(h=3\), \(n=3\) and \(m=21\).
3.2 Mathematical Morphology: Black Top-Hat
It can be assumed that retinal vessels are defined by a dark pattern with morphological properties like local linearity, connectivity, width, and a Gaussian-like cross-section profile [31]. These observations allow the use of mathematical morphology for retinal vessel enhancement. In particular, in this work, we apply a series of morphological black top-hat transforms. In general, let f be a grayscale image and b a structuring element (SE), the black top-hat transform of f by b at the ith pixel is defined as follows:
where \(f\bullet {b}\) denotes the closing of f by b. We employ this operation because it enhances objects smaller than b and darker than their surroundings. Given the linearity property of vessels, we use a linear SE \(b_{L}\), slightly longer than the width L of the largest vessels, to obtain the enhancement of all the vessels orthogonal to \(b_{L}\). To target the orientation of a specific vessel, we use multiple \(BTH(f,b_{L}^{\alpha })\) for D different angles \(\alpha \) equally spaced in the range \(\left[ 0,\pi \right) \). The enhancement of the complete vessel tree is achieved with the sum of top-hats (STH)
In this case, the length of the SE \(b_{L}\) and the number of angles used are set respectively to \(L=16\) and \(D=12\). The choice of \(L=16\) is based on the visual inspection of the images by considering the width of primary large vessels, whereas D is chosen according to [6] where 12 different templates are applied to search for vessel segments along all possible directions. We also note that varying the parameter L in the range \(\left[ 12,20\right] \) and D in the range \(\left[ 10,16\right] \) did not affect the performance on the training images significantly, suggesting that the proposed method does not need a fine tuning of its parameters.
3.3 Image Thresholding
The enhanced image is then thresholded in order to obtain a binary image in which only vessel pixels are extracted, providing the final vessel segmentation. Given a threshold \(\theta \), all the pixels with an intensity gray level greater than \(\theta \) are set to 255, whereas the others are set to 0. The value of \(\theta \) is selected according to the desired accuracy level, i.e., the value that maximizes the segmentation accuracy on the training images of DRIVE dataset.
4 Experiments
To evaluate the performance of our algorithm we compared each resulting binary image from the test set with the corresponding ground truth, by computing the number of correctly classified pixels and misclassified ones. According to these values we calculated the classification accuracy as the number of correctly classified pixels with respect to the number of pixels in the image FOV. Furthermore, sensitivity and specificity were calculated. Experimental results in terms of accuracy (Acc), sensitivity (Se) and specificity (Sp) are reported in Table 1. We compared the effectiveness of the proposed method with other existing vessel segmentation methodologies, whose performances are also shown in Table 1. Despite the proposed method is unsupervised, we compared it with both supervised and unsupervised techniques. Table 1 shows that the achieved accuracy \(Acc=94.40\) is higher or comparable than that of other unsupervised methods, and even comparable to that obtained by some reported supervised methods [19, 23, 27]. A visual step-by-step representation of the proposed framework applied to retinal images of DRIVE dataset is depicted in Fig. 2.
5 Conclusions
In this work we proposed an automated retinal vessel segmentation algorithm that provides for the use of a denoising technique and of mathematical morphology. One of the main advantage of this method is that it presents a low number of parameters to set up, and moreover it proves to be robust to their choice. It should also be stressed that the proposed algorithm is easy-to-implement and very time-efficient (<1 s per image). The performance was measured in terms of accuracy on the test set of the DRIVE dataset. The results shown in Table 1 demonstrate that we obtain an accuracy directly comparable with more complex methods for vessel segmentation, either supervised or unsupervised. These observations suggest that the performance could be improved in different ways: (i) by adding a post-processing method that aims at filling the gaps in the detected blood vessels and at removing small isolated areas misclassified as blood vessels; (ii) by constructing a feature vector from the unthresholded black top-hats as in [25] in order to apply methods designed to face the high imbalance between the two classes [3, 4, 12, 15]. In addition, since denoising revealed to be crucial for our method, future directions also include experimenting other existing denoising techniques, such as the Bilateral Filter [24].
References
Al-Diri, B., Hunter, A., Steel, D.: An active contour model for segmenting and measuring retinal vessels. IEEE Trans. Med. Imaging 28(9), 1488–1497 (2009)
Azzopardi, G., Strisciuglio, N., Vento, M., Petkov, N.: Trainable cosfire filters for vessel delineation with application to retinal images. Med. Image Anal. 19(1), 46–57 (2015)
Bria, A., Marrocco, C., Molinara, M., Tortorella, F.: An effective learning strategy for cascaded object detection. Inf. Sci. 340, 17–26 (2016)
Bria, A., Marrocco, C., Molinara, M., Tortorella, F.: A ranking-based cascade approach for unbalanced data. In: 2012 21st International Conference on Pattern Recognition (ICPR), pp. 3439–3442. IEEE (2012)
Buades, A., Coll, B., Morel, J.M.: Non-local means denoising. Image Process. Line 1, 208–212 (2011)
Chaudhuri, S., Chatterjee, S., Katz, N., Nelson, M., Goldbaum, M.: Detection of blood vessels in retinal images using two-dimensional matched filters. IEEE Trans. Med. Imaging 8(3), 263–269 (1989)
Cheng, E., Du, L., Wu, Y., Zhu, Y.J., Megalooikonomou, V., Ling, H.: Discriminative vessel segmentation in retinal images by fusing context-aware hybrid features. Mach. Vis. Appl. 25(7), 1779–1792 (2014)
Chutatape, O., Zheng, L., Krishnan, S.M.: Retinal blood vessel detection and tracking by matched Gaussian and Kalman filters. In: Proceedings of the 20th Annual International Conference of the IEEE Engineering in Medicine and Biology Society, vol. 6, pp. 3144–3149. IEEE (1998)
Fang, B., Hsu, W., Lee, M.L.: Reconstruction of vascular structures in retinal images. In: 2003 International Conference on Image Processing, ICIP 2003, Proceedings, vol. 2, pp. II–157. IEEE (2003)
Fraz, M.M., Barman, S., Remagnino, P., Hoppe, A., Basit, A., Uyyanonvara, B., Rudnicka, A.R., Owen, C.G.: An approach to localize the retinal blood vessels using bit planes and centerline detection. Comput. Methods Programs Biomed. 108(2), 600–616 (2012)
Fraz, M.M., Remagnino, P., Hoppe, A., Uyyanonvara, B., Rudnicka, A.R., Owen, C.G., Barman, S.A.: Blood vessel segmentation methodologies in retinal images-a survey. Comput. Methods Programs Biomed. 108(1), 407–433 (2012)
Galar, M., Fernandez, A., Barrenechea, E., Bustince, H., Herrera, F.: A review on ensembles for the class imbalance problem: bagging-, boosting-, and hybrid-based approaches. IEEE Trans. Syst. Man Cybern. Part C (Appl. Rev.) 42(4), 463–484 (2012)
Gao, X., Bharath, A., Stanton, A., Hughes, A., Chapman, N., Thom, S.: A method of vessel tracking for vessel diameter measurement on retinal images. In: 2001 International Conference on Image Processing, Proceedings, vol. 2, pp. 881–884. IEEE (2001)
Kanski, J.J., Bowling, B.: Clinical Ophthalmology: A Systematic Approach. Elsevier Health Sciences, Amsterdam (2011)
Lahiri, A., Roy, A.G., Sheet, D., Biswas, P.K.: Deep neural ensemble for retinal vessel segmentation in fundus images towards achieving label-free angiography. In: 2016 IEEE 38th Annual International Conference of the Engineering in Medicine and Biology Society (EMBC), pp. 1340–1343. IEEE (2016)
Lam, B.S., Gao, Y., Liew, A.W.C.: General retinal vessel segmentation using regularization-based multiconcavity modeling. IEEE Trans. Med. Imaging 29(7), 1369–1381 (2010)
Leontidis, G., Al-Diri, B., Wigdahl, J., Hunter, A.: Evaluation of geometric features as biomarkers of diabetic retinopathy for characterizing the retinal vascular changes during the progression of diabetes. In: 2015 37th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), pp. 5255–5259. IEEE (2015)
Liskowski, P., Krawiec, K.: Segmenting retinal blood vessels with deep neural networks. IEEE Trans. Med. Imaging 35(11), 2369–2380 (2016)
Marín, D., Aquino, A., Gegúndez-Arias, M.E., Bravo, J.M.: A new supervised method for blood vessel segmentation in retinal images by using gray-level and moment invariants-based features. IEEE Trans. Med. Imaging 30(1), 146–158 (2011)
Mary, M.C.V.S., Rajsingh, E.B., Naik, G.R.: Retinal fundus image analysis for diagnosis of glaucoma: a comprehensive survey. IEEE Access 4, 4327–4354 (2016)
Melinščak, M., Prentašić, P., Lončarić, S.: Retinal vessel segmentation using deep neural networks. In: VISAPP 2015, 10th International Conference on Computer Vision Theory and Applications (2015)
Mendonca, A.M., Campilho, A.: Segmentation of retinal blood vessels by combining the detection of centerlines and morphological reconstruction. IEEE Trans. Med. Imaging 25(9), 1200–1213 (2006)
Niemeijer, M., Staal, J., van Ginneken, B., Loog, M., Abramoff, M.D.: Comparative study of retinal vessel segmentation methods on a new publicly available database. In: SPIE Medical Imaging, vol. 5370, pp. 648–656. SPIE (2004)
Paris, S., Kornprobst, P., Tumblin, J., Durand, F., et al.: Bilateral filtering: theory and applications. Found. Trends® Comput. Graph. Vis. 4(1), 1–73 (2009)
Ricci, E., Perfetti, R.: Retinal blood vessel segmentation using line operators and support vector classification. IEEE Trans. Med. Imaging 26(10), 1357–1365 (2007)
Savelli, B., Bria, A., Galdran, A., Marrocco, C., Molinara, M., Campilho, A., Tortorella, F.: Illumination correction by dehazing for retinal vessel segmentation. In: 2017 IEEE 30th International Symposium on Computer-Based Medical Systems (CBMS). IEEE (2017)
Staal, J., Abràmoff, M.D., Niemeijer, M., Viergever, M.A., Van Ginneken, B.: Ridge-based vessel segmentation in color images of the retina. IEEE Trans. Med. Imaging 23(4), 501–509 (2004)
Sum, K., Cheung, P.Y.: Vessel extraction under non-uniform illumination: a level set approach. IEEE Trans. Biomed. Eng. 55(1), 358–360 (2008)
Xiang, Y., Gao, X., Zou, B., Zhu, C., Qiu, C., Li, X.: Segmentation of retinal blood vessels based on divergence and bot-hat transform. In: 2014 International Conference on Progress in Informatics and Computing (PIC), pp. 316–320. IEEE (2014)
Yang, Y., Huang, S., Rao, N.: An automatic hybrid method for retinal blood vessel extraction. Int. J. Appl. Math. Comput. Sci. 18(3), 399–407 (2008)
Zana, F., Klein, J.C.: Segmentation of vessel-like patterns using mathematical morphology and curvature evaluation. IEEE Trans. Image Process. 10(7), 1010–1019 (2001)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this paper
Cite this paper
Savelli, B., Marchesi, A., Bria, A., Marrocco, C., Molinara, M., Tortorella, F. (2017). Retinal Vessel Segmentation Through Denoising and Mathematical Morphology. In: Battiato, S., Gallo, G., Schettini, R., Stanco, F. (eds) Image Analysis and Processing - ICIAP 2017 . ICIAP 2017. Lecture Notes in Computer Science(), vol 10485. Springer, Cham. https://doi.org/10.1007/978-3-319-68548-9_25
Download citation
DOI: https://doi.org/10.1007/978-3-319-68548-9_25
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-68547-2
Online ISBN: 978-3-319-68548-9
eBook Packages: Computer ScienceComputer Science (R0)