Summary
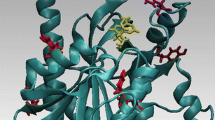
Ricin is an RNA N-glycosidase that hydrolyzes a single adenine base from a conserved loop of 28S ribosomal RNA, thus inactivating protein synthesis. Molecular-dynamics simulation methods are used to analyze the structural interactions and thermodynamics that govern the binding of formycin 5′-monophosphate (FMP) and several of its analogs to the active site of ricin A-chain. Simulations are carried out initiated from the X-ray crystal structure of the ricin-FMP complex with the ligand modeled as a dianion, monoanion and zwitterion. Relative changes in binding free energies are estimated for FMP analogs constructed from amino substitutions at the 2- and 2′-positions, and from hydroxyl substitution at the 2′-position.
Similar content being viewed by others
References
Eiklid, K., Olsnes, S. and Pihl, A., Exp. Cell Res., 126 (1980) 321.
Olsnes, S. and Pihl, A., In Cohen, P. and Van, Heyningen, S. (Eds.) Molecular Action of Toxins and Viruses, Elsevier, New York, NY, 1982, pp. 52–105.
Stilmark, H., Inaugural Lecture, University of Dorpat, Estonia, 1888.
Nicolson, G.L., Lacorbiere, M. and Hunter, T.R., Cancer Res., 35 (1975) 144.
Sperti, S., Montanaro, L. and Stripe, F., Biochem. J., 136 (1973) 813.
Endo, Y. and Tsurugi, K., J. Biol. Chem., 262 (1987) 8128.
Olsnes, S., Fernadez-Puentes, C., Carrasco, L. and Vasquez, D., Eur. J. Biochem., 60 (1975) 281.
Ready, M.P., Kim, Y. and Robertus, J.D., Proteins, 10 (1991) 270.
Endo, Y., Mitsui, K., Motizuki, M. and Tsurugi, K., J. Biol. Chem., 262 (1987) 5908.
Glück, A., Endo, Y. and Wool, I.G., J. Mol. Biol., 226 (1992) 411.
Endo, Y., Tsurugi, K. and Lambert, J.M., Biochem. Biophys. Res. Commun., 150 (1988) 1032.
Collins, E.J., Robertus, J.D., LoPreti, M., Stone, K.L., Williams, K.R., Wu, P., Hwang, K. and Piatak, M., J. Biol. Chem., 265 (1990) 8665.
Habuka, N., Murakami, Y., Noma, M., Kudo, T. and Horikoshi, K., J. Biol. Chem., 264 (1989) 6629.
Ready, M., Wilson, K., Piatak, M. and Robertus, J.D., J. Biol. Chem., 259 (1984) 15252.
Monzingo, A.F., Collins, E.J., Ernst, S.R., Irvin, J.D. and Robertus, J.D., J. Mol. Biol., 233 (1993) 705.
Benatti, L., Saccardo, M., Dani, M., Nitti, G., Sassano, M., Lorenzetti, R., Lappi, D. and Soria, M., Eur. J. Biochem., 183 (1989) 465.
Asano, K., Svensson, B., Svendsen, I., Poulsen, P. and Roepstorff, P., Carlsberg Res. Commun., 51 (1986) 129.
O'Brien, A.D. and Holmes, R.K., Microbiol. Rev., 51 (1987) 206.
Calderwood, S.B., Auclair, F., Donohue-Rolfe, A., Keusch, G.T. and Mekalanos, J.J., Proc. Natl. Acad. Sci. USA, 84 (1987) 4364.
Katzin, B.J., Collins, E.J. and Robertus, J.D., Proteins, 10 (1991) 251.
Rutenber, E., Katzin, B.J., Collins, E.J., Mlsna, D., Ernst, S.E., Ready, M.P. and Robertus, J.D., Proteins, 10 (1991) 240.
Frankel, A., Welsh, P., Richardson, J. and Robertus, J.D., Mol. Cell. Biol., 10 (1990) 6257.
Bradley, J.L. and McGuire, P.M., Int. J. Pept. Protein Res., 35 (1990) 365.
Monzingo, A.F. and Robertus, J.D., J. Mol. Biol., 227 (1992) 1136.
DeWolfJr., W.E., Fullin, F.A. and Schramm, V.L., J. Biol. Chem., 254 (1979) 10868.
Leung, H.B. and Schramm, V.L., Biochemistry, 255 (1980) 10867.
Giranda, V.L., Berman, H.M. and Schramm, V.L., Biochemistry, 27 (1988) 5813.
Beveridge, D.L. and Dicapua, F.M., In Van, Gunsteren, W.F. and Weiner, P.K. (Eds.) Computer Simulation of Biomolecular Systems, Vol. 1, ESCOM, Leiden, 1989, pp. 1–26.
Zwanzig, R.W., J. Chem. Phys., 22 (1954) 1420.
DISCOVER, Version 2.9, Biosym Technologies, Inc., San Diego, CA, 1993.
Dauber-Osguthorpe, P., Roberts, V.A., Osguthorpe, D.J., Wolff, J., Genest, M. and Hagler, A.T., Proteins, 4 (1988) 31.
Stewart, J.J.P., MOPAC 5.0, QCPE Bull., 9 (1989) 123.
Teleman, O., Jönsson, B. and Engström, S., Mol. Phys., 60 (1987) 193.
Watanabe, K., Honjo, E., Tsukamoto, T. and Funatsu, G., FEBS Lett., 304 (1992) 249.
Stote, R.H., States, D.J. and Karplus, M., J. Chem. Phys., 88 (1991) 2419.
Straub, J.E., Lim, C. and Karplus, M., J. Am. Chem. Soc., 116 (1994) 2591.
Szewczak, A.A., Moore, P.B., Chan, Y.-L. and Wool, I.G., Proc. Natl. Acad. Sci. USA, 90 (1993) 9581.
Böhm, H.-J., J. Comput.-Aided Mol. Design, 6 (1992) 61.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Olson, M.A., Scovill, J.P. & Hack, D.C. Simulation analysis of formycin 5′-monophosphate analog substrates in the ricin A-chain active site. J Computer-Aided Mol Des 9, 226–236 (1995). https://doi.org/10.1007/BF00124454
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00124454