Abstract
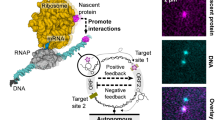
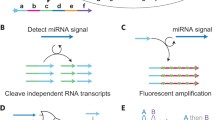
In recent years, various DNA nanomachines driven by DNA hybridizations have been developed as a remarkable application of DNA computers for nanotechnology. Here, we propose an oscillatory reaction system as a nano-sized nucleic acid engine to control the nanomachines. It utilizes DNA/RNA and their molecular reactions, and is modeled after the circadian rhythm in life systems. The molecular reactions consist of nucleic acid hybridization, RNA transcription, DNA extension, RNA degradation, and uracil-containing DNA degradation. Numerical analyses of rate equations for the reactions demonstrate that oscillatory conditions of the reaction system are determined by the balance between RNA influx into the system and RNA degradation out of the system. The analytical results will provide important information when the oscillator is constructed in in vitro experiments.
Similar content being viewed by others
References
Alberts, B., Johnson, A., Lewis, J., Raff, M. and Roberts, K., Molecular Biology of the Cell, Garland Science, 2008.
Atkinson, M. R., Savageau, M. A., Myers, J. T. and Ninfa, A. J., “Development of Genetic Circuitry Exhibiting Toggle Switch or Oscillatory Behavior in Escherichia coli,” Cell, 113, pp. 597–607, 2003.
Chen, Y. and Mao, C., “Putting a Brake on an Autonomous DNA Nanomotor,” J. Am. Chem. Soc., 126, pp. 8626–8627, 2004.
Dittmer, W. U., Kempter, S., Radler, J. O. and Simmel, F. C., “Using Gene Regulation to Program DNA-Based Molecular Devices,” Small, 1, pp. 709–712, 2005.
Dittmer, W. U. and Simmel, F. C., “Transcriptional Control of DNA-Based Nanomachines,” Nano Lett., 4, pp. 689–691, 2004.
Elowitz, M. B. and Leibler, S., “A synthetic oscillatory network of transcriptional regulators,” Nature, 403, pp. 335–338, 2000.
Fall, C. P., Marland, E. S., Wagner, J. M. and Tyson, J. J., Computational Cell Biology, Springer, 2002.
Field, R. J. and Noyes, R. M., “Oscillations in chemical systems. IV. Limit cycle behavior in a model of a real chemical reaction,” J. Chem. Phys., 60, pp. 1877–1884, 1974.
Fung, E., Wong, W. W., Suen, J. K., Bulter, T., Lee, S.-g. and Liao, J. C., “A synthetic gene-metabolic oscillator,” Nature, 435, pp. 118–122, 2005.
Hirano, N., Haruki, M., Morikawa, M. and Kanaya, S., “Enhancement of the Enzymatic Activity of Ribonuclease HI from Thermus thermophilus HB8 with a Suppressor Mutation Method,” Biochemistry, 39, pp. 13285–13294, 2000.
Mao, C., Sun, W., Shen, Z. and Seeman, N. C., “A nanomechanical device based on the B-Z transition of DNA,” Nature, 397, pp. 144–146, 1999.
Martint, C. T. and Coleman, J. E., “Kinetic Analysis of T7 RNA Polymerase-Promoter Interactions with Small Synthetic Promoters,” Biochemistry, 26, pp. 2690–2696, 1987.
Murray, J. D., Mathematical Biology I: An Introduction, 3rd ed., Springer, 2002.
Nicolis, G. and Prigogine, I., Self-Organization in Nonequilibrium Systems -From Dissipative Structures to Order through Fluctuations, John Wiley & Sons, 1977.
Seelig, G., Yurke, B. and Winfree, E., “Catalyzed Relaxation of a Metastable DNA Fuel,” J. Am. Chem. Soc., 128, pp. 12211–12220, 2006.
Sherman, W. B. and Seeman, N. C., “A Precisely Controlled DNA Biped Walking Device,” Nano Lett., 4, pp. 1203–1207, 2004.
Shin, J.-S. and Pierce, N. A., “A Synthetic DNA Walker for Molecular Transport,” J. Am. Chem. Soc., 126, pp. 10834–10835, 2004.
Simmel, F. C. and Dittmer, W. U., “DNA Nanodevices,” Small, 1, pp. 284–299, 2005.
Strogatz, S. H., Nonlinear Dynamics and Chaos: With Applications to Physics, Biology, Chemistry, and Engineering, Westview Pr, 2001.
Takinoue, M., Kiga, D., Shohda, K.-i. and Suyama, A., “Design and numerical analysis of RNA oscillator,” Proceedings in Information and Communications Technology: Natural Computing, 1, pp. 201–212, 2008.
Takinoue, M., Kiga, D., Shohda, K.-i. and Suyama, A., “Experiments and simulation models of a basic computation element of an autonomous molecular computing system,” Phys. Rev. E., 78, article no. 041921, 2008.
Tian, Y., He, Y., Chen, Y., Yin, P. and Mao, C., “A DNAzyme That Walks Processively and Autonomously along a One-Dimensional Track,” Angew. Chem. Int. Ed., 44, pp. 2–5, 2005.
Tian, Y. and Mao, C., “Molecular Gears: A Pair of DNA Circles Continuously Rolls against Each,” J. Am. Chem. Soc., 126, pp. 11410–11411, 2004.
Turberfield, A. J., Mitchell, J. C., Yurke, B., Mills Jr., A. P., Blakey, M. I. and Simmel, F. C., “DNA Fuel for Free-Running Nanomachines”, Phys. Rev. Lett., 90, article no. 118102, 2003.
Varshney, U. and Sande, J. H. v. d., “Specificities and Kinetics of Uracil Excision from Uracil-Containing DNA Oligomers by Escherichia coli Uracil DNA Glycosylase,” Biochemistry, 30, pp. 4055–4061, 1991.
Yan, H., Zhang, X., Shen, Z. and Seeman, N. C., “A robust DNA mechanical device controlled by hybridization topology,” Nature, 415, pp. 62–65, 2002.
Yin, P., Yan, H., Daniell, X. G., Turberfield, A. J. and Reif, J. H., “A Unidirectional DNAWalker That Moves Autonomously along a Track,” Angew. Chem. Int. Ed., 43, pp. 4906–4911, 2004.
Yurke, B., Turberfield, A. J., Mills Jr., A. P., Simmel, F. C. and Neumann, J. L., “A DNA-fuelled molecular machine made of DNA,” Nature, 406, pp. 605–608, 2000.
Author information
Authors and Affiliations
Corresponding author
About this article
Cite this article
Takinoue, M., Kiga, D., Shohda, Ki. et al. RNA Oscillator: Limit Cycle Oscillations based on Artificial Biomolecular Reactions. New Gener. Comput. 27, 107–127 (2009). https://doi.org/10.1007/s00354-008-0057-5
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00354-008-0057-5