Abstract
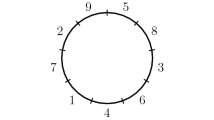
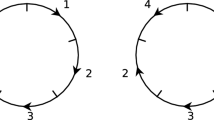
In this paper, we investigate the problem of determining the relationship, represented by similarity of the homologous gene configuration, between paired circular genomes using a regression analysis. We propose a new regression model for studying two circular genomes, where the Möbius transformation naturally arises and is taken as the link function, and propose the least circular distance estimation method, as an appropriate method for analyzing circular variables. The main utility of the new regression model is in identification of a new angular location of one of a homologous gene pair between two circular genomes, for various types of possible gene mutations, given that of the other gene. Furthermore, we demonstrate the utility of our new regression model for grouping of various genomes based on closeness of their relationship. Using angular locations of homologous genes from the five pairs of circular genomes (Horimoto et al. in Bioinformatics 14:789–802, 1998), the new model is compared with the existing models.
Similar content being viewed by others
References
Chakrabarti P, Pal D (2001) The interrelationships of side-chain and main-chain conformations in proteins. Prog Biophys Mol Biol 76:1–102
Downs TD, Mardia KV (2002) Circular regression. Biometrika 89:683–698
Fisher NI, Lee AJ (1992) Regression models for an angular response. Biometrics 48:665–677
Fisher NI (1993) Statistical analysis of circular data. Cambridge University Press, New York
Gould AL (1969) A Regression technique for angular variates. Biometrics 25:683–700
Horimoto K, Suyama M, Toh H, Mori K, Otsuka J (1998) A method for comparing circular genomes from gene locations: application to mitochondrial genomes. Bioinformatics 14:789–802
Jammalamadaka SR, SenGupta A (2001) Topics in circular statistics. World Scientific, New York
Kato S, Shimizu K, Shieh GS (2008) A circular–circular regression model. Stat Sin 18:633–645
Kim S (2009) Inverse circular regression with a possibly asymmetric error distribution. PhD Dissertation. University of California, Riverside
Liu D, Weinberg CR, Peddada SD (2004) A geometric approach to deterine association and coherence of the activation times of cell-cycling genes under different experimental conditions. Bioinformatics 20:2521–2528
Liu D, Peddada SD, Li L, Weinberg CR (2006) Phase analysis of circadian-related genes in two tissues. BMC Bioinf. doi:10.1186/1471-2105-7-87
Presnell B, Morrison SP, Littell RC (1998) Projected multivariate linear models for directional data. J Am Stat Assoc 93:1068–1077
Rivest LP (1997) A decentred predictor for circular–circular regression. Biometrika 84:717–726
SenGupta A, Ugwuowo F (2006) Asymmetric circular–linear multivariate regression models with applications to environmental data. Environ Ecol Stat 13:299–309
Author information
Authors and Affiliations
Corresponding author
Appendix
Appendix
1.1 Proof of Theorem 1
The limiting distribution of \(\zeta =\{a,b\}\) is obtained using an exact first-order Taylor series expansion of the first order condition, for some \(\zeta ^+\) between \(\hat{\zeta }\) and \(\zeta _0\),
where \(Q_n(\zeta )=\frac{1}{n}\sum _{j=1}^n\left[ 1-\cos \{\theta _j-\mu -2\arctan (a+bx_j)\}\right] \). We apply the multivariate CLT for independent random vectors in the following, to obtain an asymptotic multivariate normality of \(\frac{1}{\sqrt{n}}\sum _{j=1}^n \frac{\partial m_j}{\partial \zeta } \sin (\theta _j-m_j)|_{\zeta _{0}}\). Then,
Now, using Slutsky’s theorem, (or Product Limit Normal Rule) we get
Then, the asymptotic distributions are given by
Rights and permissions
About this article
Cite this article
SenGupta, A., Kim, S. Statistical inference for homologous gene pairs between two circular genomes: a new circular–circular regression model. Stat Methods Appl 25, 421–432 (2016). https://doi.org/10.1007/s10260-015-0341-8
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10260-015-0341-8