Abstract
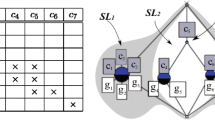
Gene expression data generated by DNA microarray experiments provide a vast resource of medical diagnostic and disease understanding. Unfortunately, the large amount of data makes it hard, sometimes even impossible, to understand the correct behavior of genes. In this work, we develop a possibilistic approach for mining gene microarray data. Our model consists of two steps. In the first step, we use possibilistic clustering to partition the data into groups (or clusters). The optimal number of clusters is evaluated automatically from the data using the Information Entropy as a validity measure. In the second step, we select from each computed cluster the most representative genes and model them as a graph called a proximity graph. This set of graphs (or hyper-graph) will be used to predict the function of new and previously unknown genes. Experimental results using real-world data sets reveal a good performance and a high prediction accuracy of our model.
Similar content being viewed by others
References
Ankerst M, Breunig MM, Kriegel HP, Sander J (1999) OPTICS: ordering points to identify the clustering structure. In: Proc of the ACM SIGMOD int conf on management of data, PH, USA, 1–3 June 1999, pp 49–60
Au W-H, Chan KCC, Wong AKC, Wang Y (2005) Attribute clustering for grouping, selection, and classification of gene expression data. IEEE/ACM Trans Comput Biol Bioinform 2(2):83–101
Baldi P, Hatfield GW (2002) DNA microarrays and gene expression: from experiments to data analysis and modelling. Cambridge University Press, Cambridge
Bezdek JC (1981) Pattern recognition with fuzzy objective function algorithms. Plenum, New York
Bickel DR (2003) Robust cluster analysis of microarray gene expression data with the number of clusters determined biologically. Bioinformatics 19(7):818–824
Blatt M, Wiseman S, Domany E (1996) Super-paramagnetic clustering of data. Phys Rev Lett 76(3251–3254):29–76
Bryan K, Cunningham P, Bolshakova N (2006) Application of simulated annealing to the biclustering of gene expression data. IEEE Trans Inf Technol Biomed 10(3):519–525
Dempster AP, Laird NM, Rubin DB (1977) Maximum likelihood from incomplete data via the EM algorithm. J R Stat Soc B 39(1):1–38
Divina F, Aguilar-Ruiz JS (2006) Biclustering of expression data with evolutionary computation. IEEE Trans Knowl Data Eng 18(5):590–602
Efron B (1982) The jackknife, the bootstrap, and other resampling plans. In: Proc of the CBMS-NSF regional conf series in applied mathematics, vol 38. SIAM, Philadelphia
Eisen MB, Spellman PT, Brown PO, Botstein D (1998) Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA 95(25):14863–14868
Gepas software. http://transcriptome.ens.fr/gepas/
Guthke R, Schmidt-Heck W, Hann D, Pfaff M (2000) Gene expression data mining for functional genomics. In: Proc of the European symp on intel techn, Aachen, Germany, pp 170–177
Hatzis C, Page D (2001). KDD Cup 2001. http://www.cs.wisc.edu/~dpage/kddcup2001
Herrero J, Valencia A, Dopazo J (2001) A hierarchical unsupervised growing neural network for clustering gene expression patterns. Bioinformatics 17(2):126–136
Hinneburg A, Keim DA (1998) An efficient approach to clustering in large multimedia database with noise. In: Proc of the 4th int conf on knowledge discovery and data mining, NY, USA, 27–31 August 1998, pp 58–65
Jiang D, Pei J, Zhang A (2003) DHC: a density-based hierarchical clustering method for time series gene expression data. In: Proc of the 3rd IEEE symp on bio-informatics and bio-engineering, Maryland, USA, 10–12 March 2003, pp 393–400
Jiang D, Tang C, Zhang A (2004) Cluster analysis for gene expression data: a survey. IEEE Trans Knowl Data Eng 16(11):1370–1386
Jinyan L, Huiqing L (2001) Kent Ridge bio-medical data set repository. http://sdmc.lit.org.sg/GEDatasets
Kaufman L, Rousseeuw PJ (1990) Finding groups in data: an introduction to cluster analysis. Wiley, New York
Kaymak U (2000) A unified approach for practical applications of fuzzy clustering. In: Proc of 20th Belgium-Netherlands conf on artif intel, Efteling, Kaatsheuvel, 1–2 November 2000, pp 101–108
Kaymak U, Babuska R (1995) Compatible cluster merging for fuzzy modeling. In: Proc of the 4th IEEE international conf on fuz syst, vol 2, Yokohama, Japan, March 1995, pp 897–904
Keedwell E, Narayanan A (2005) Discovering gene networks with a neural-genetic hybrid. IEEE/ACM Trans Comput Biol Bioinform 2(3):231–242
Krishnapuram R, Keller M (1993) A possibilistic approach to clustering. IEEE Trans Fuzzy Syst 1(2):98–110
Kumar M, Stoll R, Stoll N (2006) A min-max approach to fuzzy clustering, estimation, and identification. IEEE Trans Fuzzy Syst 14(2):248–262
Looney CG (2002) Interactive clustering and merging with a new fuzzy expected value. Patern Recogn 35(11):2413–2423
Ma PCH, Chan KCC, Yao X, Chiu DKY (2006) An evolutionary clustering algorithm for gene expression microarray data analysis. IEEE Trans Evol Comput 10(3):296–314
Madeira SC, Oliveira AL (2004) Biclustering algorithms for biological data analysis: a survey. IEEE/ACM Trans Comput Biol Bioinform 1(1):24–45
McLachlan GJ, Bean RW, Peel D (2002) A mixture model-based approach to the clustering of microarray expression data. Bioinformatics 18(3):413–422
Pal NR, Bezdek JC (1995) On cluster validity for the fuzzy c-means model. IEEE Trans Fuzzy Syst 3(2):370–379
Pal NR, Keller JM, Bezdek JC (2005) A possibilistic fuzzy C-means clustering algorithm. IEEE Trans Fuzzy Syst 13(4):517–530
Ralf-Herwig PA, Muller C, Bull C, Lehrach H, O’Brien J (1999) Large-scale clustering of cDNA-fingerprinting data. Genome Res 9(11):1093–1105
Romdhane LB, Ayeb B, Wang S (2002) On computing the fuzzifier in ↓FLVQ: a data driven approach. Int J Neurol Sys 12(2):149–157
Rose K (1998) Deterministic annealing for clustering, compression, classification, regression, and related optimization problems. Proc IEEE 86(11):2210–2239
Shamir R, Sharan R (2000) CLICK: A clustering algorithm for gene expression analysis. In: Proc 8th int conf on intelligent systems for molecular biology, CA, USA, 19–23 August 2000, pp 307–316
Shannon CE (1948) A mathematical theory of communication. Bell Syst Tech J 27(379–423):623–656
Smet FD, Mathys J, Marchal K, Thijs G, Moor BD, Moreau Y (2002) Adaptive quality-based clustering of gene expression profiles. Bioinformatics 18(5):735–746
Sun H, Wang S, Jiang Q (2004) Fcm-based model selection algorithms for determining the number of clusters. Patern Recogn 37(10):2027–2037
Tomida S, Hanai T, Honda H, Kobayashi T (2002) Analysis of expression profile using fuzzy adaptive resonance theory. Bioinformatics 18(8):1073–1083
Tou JT, Gonzalez R (1974) Pattern recognition principles. Addison-Wesley, Reading
Tsai H-K, Yang J-M, Tsai Y-F, Kao C-Y (2004) An evolutionary approach for gene expression patterns. IEEE Trans Inf Technol Biomed 8(2):69–78
Wu S, Liew A, Yan H, Yang M (2004) Cluster analysis of gene expression data based on self-splitting and merging competitive learning. IEEE Trans Inf Technol Biomed 8(1):5–15
Yang J, Wang W, Wang H, Yu P (2002) δ-Clusters: capturing subspace correlation in a large data set. In: Proc 18th IEEE int conf on data engineering. IEEE Computer Society, Los Alamitos, pp 517–528
Yeung KY, Fraley C, Murua A, Raftery AE, Ruzz WL (2001) Model-based clustering and data transformations for gene expression data. Bioinformatics 17(10):977–987
Ying X, Vijay VR, Praveen D, Xiaoquan Z (2005) A new fuzzy clustering algorithm for optimally finding granular prototypes. Int J Approx Reason 40:109–124
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Romdhane, L.B., Shili, H. & Ayeb, B. Mining microarray gene expression data with unsupervised possibilistic clustering and proximity graphs. Appl Intell 33, 220–231 (2010). https://doi.org/10.1007/s10489-009-0161-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10489-009-0161-3