Abstract
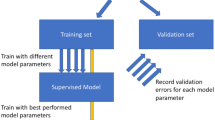
This study aimed to propose a sample selection strategy termed SPXYE (sample set partitioning based on joint X–Y–E distances) for data partition in multivariate modeling, where training and validation sets are required. This method was applied to choose the training set according to X (the independent variables), Y (the dependent variables), and E (the error of the preliminarily calculated results with the dependent variables) spaces. This selection strategy provided a valuable tool for multivariate calibration. The proposed technique SPXYE was applied to three household chemical molecular databases to obtain training and validation sets for partial least squares (PLS) modeling. For comparison, the training and validation sets were also generated using random sampling, Kennard–Stone, and sample set partitioning based on joint X–Y distances methods. The predictions of all associated PLS regression models were performed upon the same testing set, which was different from either the training set or the validation set. The results indicated that the proposed SPXYE strategy might serve as an alternative partition strategy.


Similar content being viewed by others
References
Allegrini, F., Olivier, A.C.: An integrated approach to the simultaneous selection of variables, mathematical pre-processing and calibration samples in partial least-squares multivariate calibration. Talanta 115, 755–760 (2013)
Lorber, A., Kowalski, B.R.: The effect of interferences and calbiration design on accuracy: implications for sensor and sample selection. J. Chemom. 2(1), 67–79 (1988)
Kocjančič, R., Zupan, J.: Modelling of the river flowrate: the influence of the training set selection. Chemom. Intell. Lab. Syst. 54(1), 21–34 (2000)
Jia, R.D., Mao, Z.Z., Chang, Y.Q., Zhang, S.-N.: Kernel partial robust M-regression as a flexible robust nonlinear modeling technique. Chemom. Intell. Lab. Syst. 100(2), 91–98 (2010)
Westad, F., Marini, F.: Validation of chemometric models—a tutorial. Anal. Chim. Acta 893, 14–24 (2015)
Ferre, J., Rius, F.X.: Selection of the best calibration sample subset for multivariate regression. Anal. Chem. 68(9), 1565–1571 (1996)
Hu, Y., Peng, S., Bi, Y., Tang, L.: Calibration transfer based on maximum margin criterion for qualitative analysis using Fourier transform infrared spectroscopy. Analyst 137(24), 5913–5918 (2012)
Filho, H.A.D., Galvão, R.K.H., Araújo, M.C.U., et al.: A strategy for selecting calibration samples for multivariate modelling. Chemom. Intell. Lab. Syst. 72(1), 83–91 (2004)
Capitán-Vallvey, L.F., Navas, N., Del Olmo, M., Consonni, V., Todeschini, R.: Resolution of mixtures of three nonsteroidal anti-inflammatory drugs by fluorescence using partial least squares multivariate calibration with previous wavelength selection by Kohonen artificial neural networks. Talanta 52(6), 1069–1079 (2000)
Rajer-Kanduč, K., Zupan, J., Majcen, N.: Separation of data on the training and test set for modelling: a case study for modelling of five colour properties of a white pigment. Chemom. Intell. Lab. Syst. 65(2), 221–229 (2003)
Kennard, R.W., Stone, L.A.: computer aided design of experiments. Technometrics 11(1), 137 (1969)
Galvão, R.K.H., Araujo, M.C.U., José, G.E., et al.: A method for calibration and validation subset partitioning. Talanta 67(4), 736–740 (2005)
Wu, W., Walczak, B., Massart, D.L., et al.: Artificial neural networks in classification of NIR spectral data: design of the training set. Chemom. Intell. Lab. Syst. 33(1), 35–46 (1996)
Groot, P.J., Postma, G.J., Melssen, W.J., Buydens, L.M.C.: Selecting a representative training set for the classification of demolition waste using remote NIR sensing. Anal. Chim. Acta 392(1), 67–75 (1999)
Galvão, R.K.H., José, G.E.D., Filho, H.A.D., et al.: Optimal wavelet filter construction using X and Y data. Chemom. Intell. Lab. Syst. 70(1), 1–10 (2004)
Shamsipur, M., Zare-Shahabadi, V., Hemmateenejad, B., Akhond, M.: Ant colony optimisation: a powerful tool for wavelength selection. J. Chemom. 20(3–4), 146–157 (2006)
Liu, W., Zhao, Z., Yuan, H., et al.: An optimal selection method of samples of calibration set and validation set for spectral multivariate analysis. Spectrosc. Spectr. Anal. 34(4), 947–951 (2014)
He, Z., Li, M., Ma, Z.: Design of a reference value-based sample-selection method and evaluation of its prediction capability. Chemom. Intell. Lab. Syst. 148, 72–76 (2015)
Gani, W., Limam, M.: A kernel distance-based representative subset selection method. J. Stat. Comput. Simul. 86(1), 135–148 (2016)
Chen, W.R., Yun, Y.H., Wen, M., et al.: Representative subset selection and outlier detection via isolation forest. Anal. Methods 8(39), 7225–7231 (2016)
Shao, X.G., Bian, X.H., Cai, W.S.: An improved boosting partial least squares method for near-infrared spectroscopic quantitative. Anal. Chim. Acta 666, 32–37 (2010)
Li, Y.K., Jing, J.: A consensus PLS method based on diverse wavelength variables models for analysis of near-infrared spectra. Chemom. Intell. Lab. 130, 45–49 (2014)
Gao, T., Shi, L.L., Li, H.B., et al.: Improving the accuracy of low level quantum chemical calculation for absorption energies: the genetic algorithm and neural network approach. Phys. Chem. Chem. Phys. 11(25), 5124–5129 (2009)
Gao, T., Sun, S.L., Shi, L.L., et al.: An accurate density functional theory calculation for electronic excitation energies: the least-squares support vector machine. J. Chem. Phys. 130(18), 184104 (2009)
Gao, T., Li, H., Li, W., Li, L., Fang, C., Li, H., et al.: A machine learning correction for DFT non-covalent interactions based on the S22 S66 and X40 benchmark databases. J. Cheminform. 8, 24 (2016)
Li, H.Z., Tao, W., Gao, T., et al.: Improving the Accuracy of Density Functional theory (DFT) calculation for homolysis bond dissociation energies of Y-NO bond: generalized regression neural network based on grey relational analysis and principal component analysis. Int. J. Mol. Sci. 12(4), 2242–2261 (2011)
Jurecka, P., Sponer, J., Cerny, J., Hobza, P.: Benchmark database of accurate (MP2 and CCSD(T) complete basis set limit) interaction energies of small model complexes, DNA base pairs, and amino acid pairs. Phys. Chem. Chem. Phys. 8(17), 1985–1993 (2006)
Rezac, J., Riley, K.E., Hobza, P.: S66: a well-balanced database of benchmark interaction energies relevant to biomolecular structures. J. Chem. Theory Comput. 7(8), 2427–2438 (2011)
Rezac, J., Riley, K.E., Hobza, P.: Benchmark calculations of noncovalent interactions of halogenated molecules. J. Chem. Theory Comput. 8(11), 4285–4292 (2012)
Acknowledgements
The authors gratefully acknowledge financial support from NSFC (21473025 and 21131001), the Science and Technology Development Planning of Jilin Province (20150204041GX and 20130522109JH), and the Education Projects of Jilin Province (2015552, 2014B045, 2015553 and 2015556).
Author information
Authors and Affiliations
Contributions
TG designed the study. LNH and ZZJ executed the jobs on a computer. TNX, CF and HZL provided help with the study design. TG and LHH drafted the manuscript. YHL and HL supervised the study. All authors read and approved the final manuscript.
Corresponding authors
Appendix: Matlab implementation of the proposed SPXYE algorithm
Appendix: Matlab implementation of the proposed SPXYE algorithm
In this Matlab function, X, y and e are the parameter matrix (independent variables), the experiment value matrix (dependent variable) and the error matrix (the errors between the experiment values and calculated values), respectively. Ncal is the number of samples to be selected for the training set. The indexes of the selected samples are returned in vector m.


Rights and permissions
About this article
Cite this article
Gao, T., Hu, L., Jia, Z. et al. SPXYE: an improved method for partitioning training and validation sets. Cluster Comput 22 (Suppl 2), 3069–3078 (2019). https://doi.org/10.1007/s10586-018-1877-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10586-018-1877-9