Summary
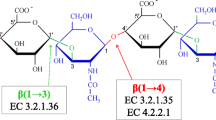
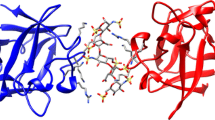
Using the hyaluronic acid (HA) binding region of the receptor for hyaluronan-mediated motility (RHAMM) as a model, a molecular perspective for peptide mimicry of the natural ligand was established by comparing the interaction sites of HA and unnatural peptide–ligands to RHAMM. This was accomplished by obtaining a series of octapeptide–ligands through screening experiments that bound to the HA binding domains of RHAMM (amino acids 517–576) and could be displaced by HA. These molecules were computationally docked onto a three-dimensional NMR based model of RHAMM. The NMR model showed that RHAMM(517–576) was a set of three helices, two of which contained the HA binding domains (HABDs) flanking a central groove. The structure was stabilized by hydrophobic interactions from four pairs of Val and Ile side chains extending into the groove. The presence of solvent exposed, positively charged side chains spaced 11 Å apart matched the spacing of negative charges on HA. Docking experiments using flexible natural and artificial ligands demonstrated that HA and peptide–mimetics preferentially bound to the second helix that contains HABD-2. Three salt bridges between HA carboxylates and Lys548, Lys553 and Lys560 and two hydrophobic interactions involving Val538 and Val559 were predicted to stabilize the RHAMM-HA complex. The high affinity peptides and HA utilized the same charged residues, with additional contacts to other basic residues. However, hydrophobic contacts do not contribute to affinity for peptide ligand-RHAMM complexes. These results offer insight into how selectivity is achieved in the binding of HA to RHAMM, and how peptide competitors may compete for binding with HA on a single hyaladherin.
Similar content being viewed by others
Abbreviations
- CVFF:
-
combined valence force field
- DYANA:
-
dynamics algorithm for NMR applications
- ECM:
-
extracellular matrix
- EDCI:
-
1-[3-(dimethylamino)propyl]-3-ethylcarbodiimide hydrochloride
- Erk:
-
extracellular signal regulated protein kinase
- FAK:
-
focal adhesion kinase
- GAG:
-
glycosaminoglycan
- GCU:
-
glucuronic acid
- GnHCl:
-
guanidinium hydrochloride
- GPI:
-
glycosyl phosphatidyl inositol
- GST:
-
glutathione-S transferase
- HA:
-
hyaluronic acid
- HABD:
-
hyaluronic acid binding domain
- HPLC:
-
high performance liquid chromatography
- MAPK:
-
mitogen activated protein kinase
- NAG:
-
N-acetyl glucosamine
- NOESY:
-
nuclear Overhauser enhanced spectroscopy
- ppm:
-
parts per million
- RHAMM:
-
receptor for hyaluronan mediated motility
- TGF:
-
transforming growth factor
- TOCSY:
-
total correlation spectroscopy
References
J.R. Fraser T.C. Laurent U.B. Laurent (1997) J. Intern. Med. 242 27
J.Y. Lee A.P. Spicer (2000) Curr. Opin. Cell Biol. 12 581
B.P. Toole (2001) Semin. Cell. Dev. Biol. 12 79
J. Entwistle C.L. Hall E.A. Turley (1996) J. Cell. Biochem. 61 569
W.Y. Chen G. Abatangelo (1999) Wound Repair Regen. 7 79
E.A. Turley P.W. Noble L.Y. Bourguignon (2002) J. Biol. Chem. 277 4589
B.D. Lynn X. Li P.A. Cattini E.A. Turley J.I. Nagy (2001) J. Comp. Neurol. 439 315
V.B. Lokeshwar M.G. Selzer (2000) J. Biol. Chem. 275 27641
J. Greiner M. Ringhoffer M. Taniguchi A. Schmitt D. Kirchner G. Krahn V. Heilmann J. Gschwend L. Bergmann H. Dohner M. Schmitt (2002) Exp. Hematol. 30 1029
A.J. Day (1999) Biochem. Soc. Trans. 27 115
D. Kohda C.J. Morton A.A. Parkar H. Hatanaka F.M. Inagaki I.D. Campbell A.J. Day (1996) Cell 86 767 Occurrence Handle1:CAS:528:DyaK28XlslShtrg%3D Occurrence Handle8797823
J. Lesley N.M. English I. Gal K. Mikecz A.J. Day R. Hyman (2002) J. Biol. Chem. 277 26600
J. Bajorath B. Greenfield S.B. Munro A.J. Day A. Aruffo (1998) J. Biol. Chem. 273 338
C.D. Blundell D.J. Mahoney A. Almond P.L. DeAngelis J.D. Kahmann P. Teriete A.R. Pickford I.D. Campbell A.J. Day (2003) J. Biol. Chem. 11 11
B. Yang B.L. Yang R.C. Savani E.A. Turley (1994) Embo J. 13 286
A.J. Day G.D. Prestwich (2002) J. Biol. Chem. 277 4585
M.R. Ziebell Z.G. Zhao B. Luo Y. Luo E.A. Turley G.D. Prestwich (2001) Chem. Biol. 8 1081
C.L. Hall B. Yang X. Yang S. Zhang M. Turley S. Samuel L.A. Lange C. Wang G.D. Curpen R.C. Savani et al. (1995) Cell 82 19
J. Sambrook E.F. Fritsch T. Maniatis (1989) Molecular Cloning A Laboratory Manual EditionNumber2 Cold Spring Harbor Laboratory Press Cold Spring Harbor, NY
C. Wang J. Entwistle G. Hou Q. Li E.A. Turley (1996) Gene 174 299 Occurrence Handle10.1016/0378-1119(96)00080-7
K. Nordstrand H. Ponstingl A. Holmgren G. Otting (1996) Eur. Biophys. J. 24 179
C. Bartels T.H. Xia M. Billeter P. Guntert K. Wüthrich (1995) J. Biomol. NMR 6 1
G.M. Clore J.G. Omichinski K. Sakaguchi N. Zambrano H. Sakamoto E. Appella A.M. Gronenborn (1994) Science 265 386
K. Wüthrich M. Billeter W. Braun (1984) J. Mol. Biol. 180 715
T.F. Havel K. Wüthrich (1985) J. Mol. Biol. 182 281
P. Guntert C. Mumenthaler K. Wüthrich (1997) J. Mol. Biol. 273 283
L.A. Kelley R.M. MacCallum M.J. Sternberg (2000) J. Mol. Biol. 299 499
P. Mallick K.E. Goodwill S. Fitz-Gibbon J.H. Miller D. Eisenberg (2000) Proc. Natl. Acad. Sci. USA, 97 2450
T.J. Ewing S. Makino A.G. Skillman I.D. Kuntz (2001) J. Comput.-Aided. Mol. Des. 15 411
T.J. Ewing I.D. Kuntz (1997) J. Comput. Chem. 18 1175
J. Wang P.A. Kollman I.D. Kuntz (1999) Proteins 36 1
M.L. Connolly (1986) J. Mol. Graphics 4 3
M.L. Connolly (1983) Science 221 709
Hendrix, D.K. and Kuntz, I.D., Pac. Symp. Biocomput. (1998) 317.
R.M. Knegtel M. Wagener (1999) Proteins 37 334
X. Fradera R.M. Knegtel J. Mestres (2000) Proteins 40 623
D.J. Mahoney C.D. Blundell A.J. Day (2001) J. Biol. Chem. 276 22764
J.E. Scott (1989) Ciba Found. Symp. 143 6
B.A. Bray (2001) J. Theor. Biol. 210 121
S. Li M.J. Jedrzejas (2001) J. Biol. Chem. 276 41407
M. Nukui K.B. Taylor D.T. McPherson M.K. Shigenaga M.J. Jedrzejas (2003) J. Biol. Chem. 278 3079
B. Yang C.L. Hall B.L. Yang R.C. Savani E.A. Turley (1994) J. Cell. Biochem. 56 455
S. Cai J.L. Dufner-Beattie G.D. Prestwich (2004) Anal. Biochem. 326 33
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ziebell, M.R., Prestwich, G.D. Interactions of peptide mimics of hyaluronic acid with the receptor for hyaluronan mediated motility (RHAMM). J Comput Aided Mol Des 18, 597–614 (2004). https://doi.org/10.1007/s10822-004-5433-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-004-5433-8