Abstract
Comparative molecular field analysis (CoMFA) and quantum chemical calculations were performed on cycloguanil (Cyc) derivatives of the wild type and the quadruple mutant (Asn51Ile, Cys59Arg, Ser108Asn, Ile164Leu) of Plasmodium falciparum dihydrofolate reductase (PfDHFR). The represented CoMFA models of wild type (\( r_{\text{cv}}^{2} = 0.727 \) and r 2 = 0.985) and mutant type (\( r_{\text{cv}}^{2} = 0.786 \) and r 2 = 0.979) can describe the differences of the Cyc structural requirements for the two types of PfDHFR enzymes and can be useful to guide the design of new inhibitors. Moreover, the obtained particular interaction energies between the Cyc and the surrounding residues in the binding pocket indicated that Asn108 of mutant enzyme was the cause of Cyc resistance by producing steric clash with p-Cl of Cyc. Consequently, comparing the energy contributions with the potent flexible WR99210 inhibitor, it was found that the key mutant residue, Asn108, demonstrates attractive interaction with this inhibitor and some residues, Leu46, Ile112, Pro113, Phe116, and Leu119, seem to perform as second binding site with WR99210. Therefore, quantum chemical calculations can be useful for investigating residue interactions to clarify the cause of drug resistance.
Graphical Abstract
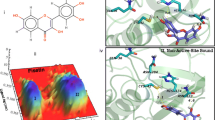
CoMFA steric contour maps of Cyc derivatives against the quadruple mutant PfDHFR.

Electrostatic van der Waals surface interaction of Cyc and the key resistance residue Asn108.








Similar content being viewed by others
References
Prapunwattana P, Sirawaraporn W, Yuthavong Y, Santi DV (1996) Mol Biochem Parasitol 83:93
Dasgupta T, Anderson KS (2008) Biochemistry 47(5):1336
Yuthavong Y, Kamchonwongpaisan S, Leartsakulpanich U, Chitnumsub P (2006) Future Microb 1(1):113
Yuthavong Y, Yuvaniyama J, Chitnumsub P, Vanichtanankul J, Chusacultanachai S, Tarnchompoo B, Vilaivan T, Kamchonwongpaisan S (2005) Parasitology 130:249
Yuvaniyama J, Chitnumsub P, Kamchonwongpaisan S, Vanichtanankul J, Sirawaraporn W, Taylor P, Walkinshaw MD, Yuthavong Y (2003) Nat Struct Biol 10:357
Yuthavong Y (2002) Microbes Infect 4:175
Nzila A (2006) Drug Discov Today 11(19–20):936
Cody V, Schwalbe CH (2006) Crystallog Rev 12(4):301
Martin S (2007) ChemMedChem 2(7):944
Warhurst DC (2002) Sci Prog 85:89
Hastings IM, Donnelly MJ (2005) Drug Resist Updates 8:43
Nzila A (2006) J Antimicrob Chemother 57:1043
Gregson A, Plowe CV (2005) Pharmacol Rev 57:117
Uhlemann AC, Yuthavong Y, Fidock DA (2005) Mol Approaches Malar. ASM Press, Washington DC, pp 429–461
Rastelli G, Sirawaraporn W, Sompornpisut P, Vilaivan T, Kamchonwongpaisan S, Quarrell R, Lowe G, Thebtaranonth Y, Yuthavong Y (2000) Bioorg Med Chem 8:1117
Peterson DS, Milhous WK, Wellems TE (1990) Proc Natl Acad Sci USA 87:3018
Sirawaraporn W, Yongkiettrakul S, Sirawaraporn R, Yuthavong Y, Santi DV (1997) Exp Parasitol 87:245
Kamchonwongpaisan S, Quarrell R, Charoensetakul N, Ponsinet R, Vilaivan T, Vanichtanankul J, Tarnchompoo B, Sirawaraporn W, Lowe G, Yuthavong Y (2004) J Med Chem 47:673
Maitarad P, Saparpakorn P, Hannongbua S, Kamchonwongpaisan S, Tarnchompoo B, Yuthavong Y (2008) Particular interaction between pyrimethamine derivatives and quadruple mutant type dihydrofolate reductase of Plasmodium falciparum: CoMFA and quantum chemical calculations studies. J Enzym Inhib Med Chem. doi:JEIMC07/1339
Kubinyi H (1993) QSAR: hansch analysis and related approaches. VCH, Weinheim
Hansch C, Leo A (1995) Exploring QSAR. Fundamentals and applications in chemistry and biology. American Chemical Society, Washington, DC
Cramer RDIII, Patterson DE, Bunce JD (1988) J Am Chem Soc 110:5959
Kubinyi H, Flokers G, Martin YC (1998) Perspect Drug Discov Des 12–14:v–vii
Cramer RDIII, Patterson DE, Bunce JD (1989) Prog Clin Biol Res 291:161
SYBYL 7.0, Tripos Associates Inc., 1699 South Hanley Road, Suite 303, St Louis, Missouri 63144, USA
Golbraikh A, Tropsha A (2002) J Mol Graph Mod 20:269
Nilsson J, De Jong S, Smilde AK (1997) J Chemom 11:511
Hannongbua S, Nivesanond K, Lawtrakul L, Pungpo P, Wolschann P (2001) J Chem Inf Comput Sci 41:848
Cornell WD, Cieplak P, Bayly CI, Kollman PA (1993) J Am Chem Soc 115:9620
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Montgomery JA Jr, Vreven T, Kudin KN, Burant JC, Millam JM, Iyengar SS, Tomasi J, Barone V, Mennucci B, Cossi M, Scalmani G, Rega N, Petersson GA, Nakatsuji H, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Klene M, Li X, Knox JE, Hratchian HP, Cross JB, Adamo C, Jaramillo J, Gomperts R, Stratmann RE, Yazyev O, Austin AJ, Cammi R, Pomelli C, Ochterski JW, Ayala PY, Morokuma K, Voth GA, Salvador P, Dannenberg JJ, Zakrzewski VG, Dapprich S, Daniels AD, Strain MC, Farkas O, Malick DK, Rabuck AD, Raghavachari K, Foresman JB, Ortiz JV, Cui Q, Baboul AG, Clifford S, Cioslowski J, Stefanov BB, Liu G, Liashenko A, Piskorz P, Komaromi I, Martin RL, Fox DJ, Keith T, Al-Laham MA, Peng CY, Nanayakkara A, Challacombe M, Gill PMW, Johnson B, Chen W, Wong MW, Gonzalez C, Pople JA (2003) Gaussian 03, revision B.05. Gaussian, Inc., Pittsburgh
AMBER parameter database. http://www.pharmacy.manchester.ac.uk/bryce/amber/
Cummins PL, Ramnarayan K, Singh UC, Gready JE (1991) J Am Chem Soc 113(22):8247
Case DA, Darden TA, Cheatham III TE, Simmerling CL, Wang J, Duke RE, Luo R, Merz KM, Pearlman DA, Crowley M, Walker RC, Zhang W, Wang B, Hayik S, Roitberg A, Seabra G, Wong KF, Paesani F, Wu X, Brozell S, Tsui V, Gohlke H, Yang L, Tan C, Mongan J, Hornak V, Cui G, Beroza P, Mathews DH, Schafmeister C, Ross WS, Kollman PA (2006) AMBER9.0
Boys SF, Bernardi F (1970) Mol Phys 19:553
Saen-oon S, Kuno M, Hannongbua S (2005) Proteins 61:859
Kuno M, Hongkrengkai R, Hannongbua S (2006) Chem Phys Lett 424:172
Saen-oon S, Aruksakunwong O, Wittayanarakul K, Sompornpisut P, Hannongbua S (2007) J Mol Graph Model 26:720
Acknowledgements
This work was supported by the Thai Graduate Institute of Science and Technology (TGIST) Funds (TG-22-11-845D) under the National Science and Technology Development Agency, Thailand. S.H. is grateful to the Thailand Research Fund (RTA5080005). The Postgraduate Education and Research Program in Petroleum Petrochemical Technology and Advanced Materials, the Commission on Higher Education (CHE), the Kasetsart University Research and Development Institute (KURDI), and the Graduate School Kasetsart University Scholarship are gratefully acknowledged for financial support. The authors would like to thank the Large-Scale Research Laboratory of the National Electronics and Computer Technology (NECTEC), the National Center for Genetic Engineering and Biotechnology (BIOTEC), LCAC, and the computing center of KU for computing research facilities. This work has been partially supported by the National Nanotechnology Center (NANOTEC), NSTDA, Ministry of Science and Technology, Thailand, through its Center of Excellence Network program. Medicines for Malaria Venture (MMV), Wellcome Trust, and UNICEF/UNDP/World Bank/WHO Special Programme for Research and Training in Tropical Diseases (TDR) for Y.Y. and S.K. S.K. is an international research scholar of Howard Hughes Medical Institute (HHMI), USA. Finally, P.M. would like to thank Dr. Chak Sangma and Dr. Witcha Treesuwan for their AMBER supports.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Maitarad, P., Kamchonwongpaisan, S., Vanichtanankul, J. et al. Interactions between cycloguanil derivatives and wild type and resistance-associated mutant Plasmodium falciparum dihydrofolate reductases. J Comput Aided Mol Des 23, 241–252 (2009). https://doi.org/10.1007/s10822-008-9254-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-008-9254-z