Abstract
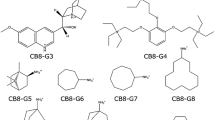
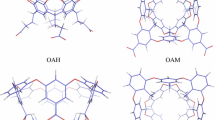
The affinities of two sets of guest–host systems were estimated using the popular end-point methods MM/GBSA (molecular-mechanics with generalised Born and surface-area solvation) and LIE (linear interaction energy). A set of six primary alcohols that bind to α-cyclodextrin (α-CD) and a set of eight guest molecules to cucurbit[8]uril (CB8) were considered. Three different charge schemes were used to obtain charges for the host and guest molecules, viz., AM1-BCC, RESP, and the recently suggested xAvESP (which average ESP charges over a number of molecular dynamics snapshots). Furthermore, both the generalised Born and Poisson–Boltzmann solvation models were used in the MM/GBSA calculations. The two solvation models perform equally well in predicting relative affinities, and hence there is no point in using the more expensive Poisson–Boltzmann model for these systems. Both the LIE and MM/GBSA estimates are shown to be robust with respect to the charge model, and therefore it is recommended to use the cheapest AM1-BCC charges. Using AM1-BCC charges, the MM/GBSA method gave a MADtr (mean absolute deviation after removal of systematic error) of 17 kJ/mol and a correlation coefficient (r 2) of 0.67 for the CB8 complexes, and a MADtr of 10 kJ/mol and an r 2 of 0.96 for the α-CD complexes. The LIE method gave a MADtr of 20 kJ/mol and an r 2 of 0.10 for the CB8 complexes, after optimisation of the non-polar scaling parameter. For the α-CD complexes, no optimisation was necessary and the method gave a MADtr of 2 kJ/mol and a r 2 of 0.96. These results indicate that both MM/GBSA and LIE are able to estimate host–guest affinities accurately.



Similar content being viewed by others
References
Chipot C, Pohorille A (eds) (2007) Free Energy Calculations. Springer, New York
Shirts M, Mobley DL, Chodera JD (2007) Annu Rep Comput Chem 3:41–59
Michel J, Essex JW (2010) J Comput Aided Mol Des 24:639–658
Gohlke H, Klebe G (2002) Angew Chem Int 41:2644–2676
Moghaddam S, Inoue Y, Gilson MK (2009) J Am Chem Soc 131:4012–4021
Mikulskis P, Genheden S, Sandberg L, Olsen L, Ryde U (2011) J Comput Aided Mol Des, submitted
Srinivasan J, Cheatham TE III, Cieplak P, Kollman PA, Case DA (1998) J Am Chem Soc 37:9401–9409
Kollman PA, Massova I, Reyes I, Kuhn B, Huo S, Chong L, Lee M, Lee T, Duan Y, Wang W, Donini O, Cieplak P, Srinivasan J, Case DA, CheathamIII TE (2000) Acc Chem Res 33:889–897
Åqvist J, Medina C, Samuelsson J-E (1994) Protein Eng 7:385–391
Hansson T, Marelius J, Åqvist J (1998) J Comput-Aided Mol Design 12:27–35
Cornell WD, Cieplak P, Bayly CI, Gould IR, Merz KM, Ferguson DM, Spellmeyer DC, Fox T, Caldwell JW, Kollman PA (1995) J Am Chem Soc 117:5179–5197
MacKerell AD Jr, Bashford D, Bellott M, Dunbrack RL Jr, Evanseck JD, Field MJ, Fischer S, Gao J, Guo H, Ha S, Joseph-McCarthy D, Kuchnir L, Kuczera K, Lau FTK, Mattos C, Michnick S, Ngo T, Nguyen DT, Prodhom B, Reiher WE III, Roux B, Schlenkrich M, Smith JC, Stote R, Straub J, Watanabe M, Wiórkiewicz-Kuczera J, Yin D, Karplus M (1998) J Phys Chem B 102:3586–3616
Jorgensen WL, Maxwell DS, Tirado-Rives J (1996) J Am Chem Soc 118:11225–11236
Oostenbrink C, Villa A, Mark AE, van Gunsteren WF (2004) J Comput Chem 25:1656–1676
Wang JM, Wolf RM, Caldwell KW, Kollman PA, Case DA (2004) J Comput Chem 25:1157–1174
Vanommeslaeghe K, Hatcher E, Acharya C, Kundu S, Zhong S, Shim J, Darian E, Guvench O, Lopes P, Vorobyov I, Mackerell AD Jr (2009) J Comput Chem 31:671–690
Bayly CI, Cieplak P, Cornell WD, Kollman PA (1997) J Phys Chem 97:10269–10280
Jakalian A, Jack DB, Bayly CI (2002) J Comput Chem 23:1623–1641
Söderhjelm P, Ryde U (2009) J Comput Chem 30:750–760
Genheden S, Söderhjelm P, Ryde U (2011) Int J Quant Chem. doi:10.1002/qua.22967
Wall ID, Leach AR, Salt DW (1999) J Med Chem 42:5142–5152
Almlöf M, Brandsdal BO, Åqvist J (2004) J Comput Chem 25:1242–1254
Lee MC, Yang R, Duan Y (2005) J Mol Model 12:101–110
Genheden S, Luchko T, Gusarov S, Kovalenko A, Ryde U (2010) J Phys Chem B 114:8505–8516
Liu S, Ruspic C, Mukhopadhyay P, Chakrabarti S, Zavalij PY, Isaacs L (2005) J Am Chem Soc 127:15959–15967
Puliti R, Mattia CA, Paduano L (1998) Carbohydr Res 310:1–8
Linder K, Saenger W (1982) Carbohydr Res 99:103–115
Jeon WS, Kim H-J, Lee C, Kim K (2002) Chem Commun 1828–1929
Matsui Y, Mochida K (1979) Bull Chem Soc Jpn 52:2808–2814
Choi Y, Jung S (2004) Carbohydr Res 339:1961–1966
El-Barghouthi MI, Assaf KI, Rawashdeh AMM (2010) J Chem Theory Comput 6:984–992
El-Barghouthi MI, Jaime C, Al-Sakhen NA, Issa AA, Abdoh AA, Al Omari MM, Badwan AA, Zughul MB (2008) J Mol Struct: Theochem 853:45–52
ChemAxon, Marvin version 5.6 (2011)
Case DA, Darden TA, Cheatham TE III, Simmerling CL, Wang J, Duke RE, Luo R, Crowley M, Walker RC, Zhang W, Merz KM, Wang B, Hayik S, Roitberg A, Seabra G, Kolossvary I, Wong KF, Paesani F, Vanicek J, Wu X, Brozell SR, Steinbrecher T, Gohlke H, Yang L, Tan C, Mongan J, Hornak V, Cui G, Mathews DH, Seetin MG, Sagui C, Babin V (2008) Kollman, P A Amber 10. University of California, San Francisco
Besler BH, Merz KM, Kollman PA (1990) J Comput Chem 11:431–439
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Montgomery JA Jr, Vreven T, Kudin KN, Burant JC, Millam JM, Iyengar SS, Tomasi J, Barone V, Mennucci B, Cossi M, Scalmani G, Rega N, Petersson GA, Nakatsuji H, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Klene M, Li X, Knox JE, Hratchian HP, Cross JB, Bakken V, Adamo C, Jaramillo J, Gomperts R, Stratmann RE, Yazyev O, Austin AJ, Cammi R, Pomelli C, Ochterski JW, Ayala PY, Morokuma K, Voth GA, Salvador P, Dannenberg JJ, Zakrzewski VG, Dapprich S, Daniels AD, Strain MC, Farkas O, Malick DK, Rabuck AD, Raghavachari K, Foresman JB, Ortiz JV, Cui Q, Baboul AG, Clifford S, Cioslowski J, Stefanov BB, Liu G, Liashenko A, Piskorz P, Komaromi I, Martin RL, Fox DJ, Keith T, Al-Laham MA, Peng CY, Nanayakkara A, Challacombe M, Gill PMW, Johnson B, Chen W, Wong MW, Gonzalez C, Pople JA (2004) Gaussian, 03 Revision. Gaussian Inc, Wallingford
Jorgensen WL, Chandrasekhar J, Madura JD, Impley RW, Klein ML (1983) J Chem Phys 79:926–935
Ryckaert JP, Ciccotti G, Berendsen HJC (1977) J Comput Phys 23:327–341
Darden T, York D, Pedersen L (1993) J Chem Phys 98:10089–10092
Wu X, Brooks BR (2003) Chem Phys Lett 381:512–518
Berendsen HJC, Postma JPM, Van Gunsteren WF, Dinola A, Haak JR (1984) J Chem Phys 81:3684–3690
Genheden S, Ryde U (2011) J Comput Chem 32:187–195
Onufriev A, Bashford D, Case DA (2004) Proteins 55:383–394
Kuhn B, Kollman PA (2000) J Med Chem 43:3786–3791
Åqvist J, Hansson T (1996) J Phys Chem 100:9512–9521
Pearlman DA, Chareifson PS (2001) J Med Chem 44:3417–3423
Genheden S, Ryde U (2010) J Comput Chem 31:837–846
Singh N, Warshel A (2010) Proteins 78:1705–1723
Hayes JM, Skamnaki VT, Archontis G, Lamprakis C, Sarrou J, Bischler N, Skaltsounis A-L, Zographos SE, Oikonomakos NH (2011) Proteins 79:704–719
Hou T, Wang J, Li Y, Wang W (2011) J Chem Inf Model 51:69–82
Shivakumar D, Deng Y, Roux B (2009) J Chem Theory Comput 5:919–930
Acknowledgments
This investigation has been supported by grants from the Research school in pharmaceutical science. It has also been supported by computer resources of LUNARC at Lund University (project SNIC001-10-225), NSC at Linköping University and HPC2 N at Umeå University (project SNIC014-10-24). Prof. Ulf Ryde and Svante Hedström are acknowledged for discussions and critical proofreading.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Genheden, S. MM/GBSA and LIE estimates of host–guest affinities: dependence on charges and solvation model. J Comput Aided Mol Des 25, 1085–1093 (2011). https://doi.org/10.1007/s10822-011-9486-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-011-9486-1