Abstract
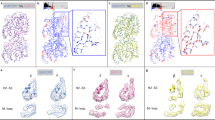
The dynamics of microtubules is essential for many microtubule-dependent cellular functions such as the mitosis. It has been recognized for a long time that GTP hydrolysis in αβ-tubulin polymers plays a critical role in this dynamics. However, the effects of the changes in the nature of the guanosine nucleotide at the E-site in β-tubulin on microtubule structure and stability are still not well understood. In the present work, we performed all-atom molecular dynamics simulations of a αβα-tubulin heterotrimer harboring a guanosine nucleotide in three different states at the E-site: GTP, GDP-Pi and GDP. We found that changes in the nucleotide state is associated with significant conformational variations at the α-tubulin N- and β-tubulin M-loops which impact the interactions between tubulin protofilaments. The results also show that GTP hydrolysis reduces αβ-tubulin interdimer contacts in favor of intradimer interface. From an atomistic point view, we propose a role for α-tubulin glutamate residue 254 in catalytic magnesium coordination and identified a water molecule in the nucleotide binding pocket which is most probably required for nucleotide hydrolysis. Finally, the results are discussed with reference to the role of taxol in microtubule stability and the recent tubulin-sT2R crystal structures.






Similar content being viewed by others
Abbreviations
- GTP:
-
Guanosine triphosphate
- GDP:
-
Guanosine diphosphate
- Pi:
-
Inorganic phosphate
- MT:
-
Microtubule
References
Desai A, Mitchison TJ (1997) Annu Rev Cell Dev Biol 13(1):83–117
Nogales E (2001) Annu Rev Biophys Biomol Struct 30(1):397–420
Amos L, Klug A (1974) J Cell Sci 14(3):523–549
Walker RA, O’Brien ET, Pryer NK, Soboeiro MF, Voter WA, Erickson HP, Salmon ED (1988) J Cell Biol 107(4):1437–1448
Allen C, Borisy GG (1974) J Mol Biol 90(2):381–402
Burns RG (1991) Cell Motil Cytoskelet 20(3):181–189
Nogales E, Wolf SG, Downing KH (1998) Nature 391(6663):199–203
Lowe J, Li H, Downing KH, Nogales E (2001) J Mol Biol 313(5):1045–1057
Ravelli RB, Gigant B, Curmi PA, Jourdain I, Lachkar S, Sobel A, Knossow M (2004) Nature 428(6979):198–202
Serrano L, de la Torre J, Maccioni RB, Avila J (1984) Proc Natl Acad Sci USA 81(19):5989–5993
Spiegelman BM, Penningroth SM, Kirschner MW (1977) Cell 12(3):587–600
David-Pfeuty T, Erickson HP, Pantaloni D (1977) Proc Natl Acad Sci USA 74(12):5372–5376
Carlier MF, Didry D, Pantaloni D (1997) Biophys J 73(1):418–427
Correia JJ, Beth AH, Williams RC (1988) J Biol Chem 263(22):10681
Grover S, Hamel E (1994) Eur J Biochem 222(1):163–172
Caplow M, Shanks J (1996) Mol Biol Cell 7(4):663–675
Carlier MF, Didry D, Pantaloni D (1987) Biochemistry 26(14):4428–4437
O’Brien ET, Voter WA, Erickson HP (1987) Biochemistry 26(13):4148–4156
Panda D, Miller HP, Wilson L (2002) Biochemistry 41(5):1609–1617
Schek HT 3rd, Gardner MK, Cheng J, Odde DJ, Hunt AJ (2007) Curr Biol 17(17):1445–1455
Caplow M, Ruhlen RL (1994) J Cell Biol 127(3):779–788
Nogales E, Wang HW (2006) Curr Opin Struct Biol 16(2):221–229
Wang HW, Nogales E (2005) Nature 435(7044):911–915
Buey RM, Diaz JF, Andreu JM (2006) Biochemistry 45(19):5933–5938
Rice LM, Montabana EA, Agard DA (2008) Proc Natl Acad Sci USA 105(14):5378–5383
Nawrotek A, Knossow M, Gigant B (2011) J Mol Biol 412(1):35–42
Gebremichael Y, Chu JW, Voth GA (2008) Biophys J 95(5):2487–2499
Bennett MJ, Chik JK, Slysz GW, Luchko T, Tuszynski J, Sackett DL, Schriemer DC (2009) Biochemistry 48(22):4858–4870
Keskin O, Durell SR, Bahar I, Jernigan RL (2002) Biophys J 83(2):663–680
Mitra A, Sept D (2008) Biophys J 95(7):3252–3258
Grafmüller A, Voth GA (2011) Structure 19(3):409–417
Fiser A, Sali A (2003) Meth Enzymol 374:461–491
Laskowski RA, MacArthur MW, Moss DS, Thornton JM (1993) J App Crystallogr 26(2):283–291
The PyMOL molecular Graphics System, version 1.3.1 Schrödinger, LLC. New York
Cornell WD, Cieplak P, Bayly CI, Gould IR, Merz KM, Ferguson DM, Spellmeyer DC, Fox T, Caldwell JW, Kollman PA (1995) J Am Chem Soc 117(19):5179–5197
Duan Y, Wu C, Chowdhury S, Lee MC, Xiong G, Zhang W, Yang R, Cieplak P, Luo R, Lee T, Caldwell J, Wang J, Kollman P (2003) J Comput Chem 24(16):1999–2012
Case DA, Darden TA, Cheatham Iii TE, Simmerling CL, Wang J, Duke RE, Luo R, Walker RC, Zhang W, Merz KM, Roberts B, Wang B, Hayik S, Roitberg A, Seabra G, Kolossvai I, Wong KF, Paesani F, Vanicek J, Liu J, Wu X, Brozell SR, Steinbrecher T, Gohlke H, Cai Q, Ye X, Hsieh MJ, Cui G, Roe DR, Mathews DH, Seetin MG, Sagui C, Babin V, Luchko T, Gusarov S, Kovalenko A, Kollman PA (2010) AMBER 11. University of California, San Francisco, CA
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML (1983) J Chem Phys 79(2):926
Meagher KL, Redman LT, Carlson HA (2003) J Comput Chem 24(9):1016–1025
Sousa Da Silva AW, Wranken WF, Laue ED (To be submitted)
Van Der Spoel D, Lindahl E, Hess B, Groenhof G, Mark AE, Berendsen HJ (2005) J Comput Chem 26(16):1701–1718
Darden TA, York D, Pedersen L (1993) J Chem Phys 98(12):10089
Berendsen HJC, Postma JPM, van Gunsteren WF, DiNola A, Haak JR (1984) J Chem Phys 81(8):3684
Bussi G, Donadio D, Parrinello M (2007) J Chem Phys 126(1):014101
Humphrey W, Dalke A, Schulten K (1996) J Mol Graph 14(1):33–38, 27–38
Golovin A, Dimitropoulos D, Oldfield T, Rachedi A, Henrick K (2005) Proteins 58(1):190–199
Golovin A, Henrick K (2008) BMC Bioinf 9:312
Golovin A, Henrick K (2009) J Chem Inf Model 49(1):22–27
Dougherty CA, Sage CR, Davis A, Farrell KW (2001) Biochemistry 40(51):15725–15732
Nogales E, Downing KH, Amos LA, Lowe J (1998) Nat Struct Biol 5(6):451–458
Friedman ZY, Devary Y (2005) Proteins 59(3):528–533
Caplow M, Shanks J (1998) Biochemistry 37(37):12994–13002
Giannakakou P, Gussio R, Nogales E, Downing KH, Zaharevitz D, Bollbuck B, Poy G, Sackett D, Nicolaou KC, Fojo T (2000) Proc Natl Acad Sci USA 97(6):2904–2909
Morrissette NS, Mitra A, Sept D, Sibley LD (2004) Mol Biol Cell 15(4):1960–1968
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
André, J.R., Clément, MJ., Adjadj, E. et al. The state of the guanosine nucleotide allosterically affects the interfaces of tubulin in protofilament. J Comput Aided Mol Des 26, 397–407 (2012). https://doi.org/10.1007/s10822-012-9566-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-012-9566-x