Abstract
S-Adenosylmethionine (AdoMet) is involved in many biological processes as cofactor in enzymes transferring its sulfonium methyl group to various substrates. Additionally, it is used as drug and nutritional supplement to reduce the pain in osteoarthritis and against depression. Due to the biological relevance of AdoMet it has been part of various computational simulation studies and will also be in the future. However, to our knowledge no rigorous force field parameter development for its simulation in biological systems has been reported. Here, we use electronic structure calculations combined with molecular dynamics simulations in explicit solvent to develop force field parameters compatible with the AMBER99 force field. Additionally, we propose new dynamic Hirshfeld-I atomic charges which are derived from the polarized electron density of AdoMet in aqueous solution to describe its electrostatic interactions in biological systems. The validation of the force field parameters and the atomic charges is performed against experimental interproton NOE distances of AdoMet in aqueous solution and crystal structures of AdoMet in the cavity of three representative proteins.
Graphical Abstract
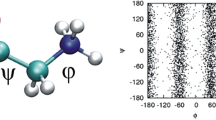
A S-Adenosylmethionine force field was developed together with Dynamic Hirshfeld-I charges (shown color coded in figure) and validated against various experimental data.








Similar content being viewed by others
References
Cantoni GL (1975) Annu Rev Biochem 44:435
Ravanel S, Gakière B, Job D, Douce R (1998) Proc Natl Acad Sci USA 95:7805
Jones PA, Takai D (2001) Science 293:1068
Stock JB, Surette MG, McCleary WR, Stock AM (1992) J Biol Chem 267:19753
Berger SL (2001) Science 292:64
Frey PA, Magnusson OT et al (2003) Chem Rev 103:2129–2148
Frey PA (2001) Annu Rev Biochem 70:121
Cantoni GL (1952) J Am Chem Soc 74:2942
Parks LW, Schlenk F et al (1958) J Biol Chem 230:295
Borchardt RT (1979) J Am Chem Soc 101:458
Follmann H, Kuntz I, Zacharias W (1975) Eur J Biochem 58:31
Follmann H, Gremels G (1974) Eur J Biochem 47:187
Klee WA, Mudd SH (1967) Biochemistry 6:988
Stolowitz ML, Minch MJ (1981) J Am Chem Soc 103:6017
Markham GD, Norrby P-O, Bock CW (2002) Biochemistry 41:7636
Hu P, Wang S, Zhang Y (2008) J Am Chem Soc 130:3806
Stacklies W, Xia F, Gräter F (2009) PLoS Comput Biol 5:e1000574
Huang W, Kim J, Jha S, Aboul-ela F (2009) Nucleic acids res 37:6528
Whitford PC, Schug A, Saunders J, Hennelly SP, Onuchic JN, Sanbonmatsu KY (2009) Biophys J 96:L7
Wang J, Cieplak P, Kollman PA (2000) J Comput Chem 21:1049
Wang J, Wolf RM, Caldwell JW, Kollman PA, Case DA (2004) J Comput Chem 25:1157
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Montgomery J, Vreven T, Kudin KN, Burant JC et al (2004) Gaussian 03, Revision E. 01. Gaussian, Inc., Wallingford
Bayly CI, Cieplak P, Cornell W, Kollman PA (1993) J Phys Chem 97:10269
Dupradeau F-Y, Pigache A, Zaffran T, Savineau C, Lelong R, Grivel N, Lelong D, Rosanski W, Cieplak P (2010) Phys chem chem phys 12:7821
Bultinck P, Van Alsenoy C, Ayers PW, Carbó-Dorca R (2007) J Chem Phys 126:144111
Hirshfeld FL (1977) Theoretical chemistry accounts: theory, computation, and modeling. Theor Chim Acta 44:129
Van Damme S, Bultinck P, Fias S (2009) J Chem Theory Comput 5:334
Verstraelen T HORTON 1.2.1
Vöhringer-Martinez E, Verstraelen T, Ayers PW (2014) J Phys Chem B 118:9871
Perone CS (2009) ACM SIGEVOlution 4:12
Hess B, Kutzner C, van der Spoel D, Lindahl E (2008) J ChemTheory Comput 4:435
Bussi G, Donadio D, Parrinello M (2007) J Chem Phys 126:014101
Hess B, Bekker H, Berendsen H, Fraaije J (1997) J Comput Chem 18:1463
Berendsen H, Grigera JR, Straatsma TP (1987) J Phys Chem 91:6269
Tsai ML, Cronin N, Djordjevic S (2011) Acta Crystallogr Sect D 67:14
Kwon T, Chang JH, Kwak E, Lee CW, Joachimiak A, Kim YC, Lee J, Cho Y (2003) EMBO J. 22:292
Lai C-W, Chen H-L, Lin K-Y, Liu F-C, Chong K-Y, Cheng WTK, Chen C-M (2014) PLoS ONE 9:e90818
Daura X, Antes I, van Gunsteren WF (1999) Proteins: Structure
Burgi R, Pitera J, van Gunsteren WF (2001) J Biomol NMR 19:305
Zagrovic B, van Gunsteren WF (2006) Proteins 63:210
Yildirim I, Stern HA, Kennedy SD, Tubbs JD, Turner DH (2010) J Chem Theory Comput 6:1520
Acknowledgments
EVM is thankful for funding support provided by FONDECYT 11121179 and Grant ICM No 120082. DS thanks CONICYT for the graduate scholarship 21130517.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Saez, D.A., Vöhringer-Martinez, E. A consistent S-Adenosylmethionine force field improved by dynamic Hirshfeld-I atomic charges for biomolecular simulation. J Comput Aided Mol Des 29, 951–961 (2015). https://doi.org/10.1007/s10822-015-9864-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-015-9864-1