Abstract
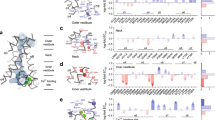
Calcium-activated chloride channels (CaCCs) play vital roles in a variety of physiological processes. Transmembrane protein 16A (TMEM16A) has been confirmed as the molecular counterpart of CaCCs which greatly pushes the molecular insights of CaCCs forward. However, the detailed mechanism of Ca2+ binding and activating the channel is still obscure. Here, we utilized a combination of computational and electrophysiological approaches to discern the molecular mechanism by which Ca2+ regulates the gating of TMEM16A channels. The simulation results show that the first intracellular loop serves as a Ca2+ binding site including D439, E444 and E447. The experimental results indicate that a novel residue, E447, plays key role in Ca2+ binding. Compared with WT TMEM16A, E447Y produces a 30-fold increase in EC50 of Ca2+ activation and leads to a 100-fold increase in Ca2+ concentrations that is needed to fully activate the channel. The following steered molecular dynamic (SMD) simulation data suggests that the mutations at 447 reduce the Ca2+ dissociation energy. Our results indicated that both the electrical property and the size of the side-chain at residue 447 have significant effects on Ca2+ dependent gating of TMEM16A.








Similar content being viewed by others
References
Hartzell C, Putzier I, Arreola J (2005) Calcium-activated chloride channels. Annu Rev Physiol 67:719–758
Jentsch TJ, Stein V, Weinreich F, Zdebik AA (2002) Molecular structure and physiological function of chloride channels. Physiol Rev 82:503–568
Caputo A, Caci E, Ferrera L, Pedemonte N, Barsanti C, Sondo E, Pfeffer U, Ravazzolo R, Zegarra-Moran O, Galietta LJ (2008) TMEM16A, a membrane protein associated with calcium-dependent chloride channel activity. Science 322:590–594
Schroeder BC, Cheng T, Jan YN, Jan LY (2008) Expression cloning of TMEM16A as a calcium-activated chloride channel subunit. Cell 134:1019–1029
Yang YD, Cho H, Koo JY, Tak MH, Cho Y, Shim WS, Park SP, Lee J, Lee B, Kim BM, Raouf R, Shin YK, Oh U (2008) TMEM16A confers receptor-activated calcium-dependent chloride conductance. Nature 455:1210–1215
Sontz PA, Song WJ, Tezcan FA (2014) Interfacial metal coordination in engineered protein and peptide assemblies. Curr Opin Chem Biol 19:42–49
Huang F, Wong X, Jan LY (2012) International Union of Basic and Clinical Pharmacology. LXXXV: calcium-activated chloride channels. Pharmacol Rev 64:1–15
Tian Y, Kongsuphol P, Hug M, Ousingsawat J, Witzgall R, Schreiber R, Kunzelmann K (2011) Calmodulin-dependent activation of the epithelial calcium-dependent chloride channel TMEM16A. FASEB J 25:1058–1068
Jung J, Nam JH, Park HW, Oh U, Yoon JH, Lee MG (2013) Dynamic modulation of ANO1/TMEM16A HCO3(–) permeability by Ca2+/calmodulin. Proc Natl Acad Sci USA 110:360–365
Brunner JD, Lim NK, Schenck S, Duerst A, Dutzler R (2014) X-ray structure of a calcium-activated TMEM16 lipid scramblase. Nature 516:207–212
Yu K, Duran C, Qu Z, Cui YY, Hartzell HC (2012) Explaining calcium-dependent gating of anoctamin-1 chloride channels requires a revised topology. Circ Res 110:990–999
Lee J, Jung J, Tak MH, Wee J, Lee B, Jang Y, Chun H, Yang DJ, Yang YD, Park SH, Han BW, Hyun S, Yu J, Cho H, Hartzell HC, Oh U (2015) Two helices in the third intracellular loop determine anoctamin 1 (TMEM16A) activation by calcium. Pflugers Arch 467:1677–1687
Tien J, Peters CJ, Wong XM, Cheng T, Jan YN, Jan LY, Yang H (2014) A comprehensive search for calcium binding sites critical for TMEM16A calcium-activated chloride channel activity. eLife 3:e02772. doi:10.7554/eLife.02772
Yang H, Zhang G, Cui J (2015) BK channels: multiple sensors, one activation gate. Front Physiol 6:29
Xiao Q, Yu K, Perez-Cornejo P, Cui Y, Arreola J, Hartzell HC (2011) Voltage- and calcium-dependent gating of TMEM16A/Ano1 chloride channels are physically coupled by the first intracellular loop. Proc Natl Acad Sci USA 108:8891–8896
Xiao Q, Cui Y (2014) Acidic amino acids in the first intracellular loop contribute to voltage- and calcium- dependent gating of anoctamin1/TMEM16A. PLoS ONE 9:e99376
Roberts E, Eargle J, Wright D, Luthey-Schulten Z (2006) MultiSeq: unifying sequence and structure data for evolutionary analysis. BMC Bioinform 7:382
Eswar N, Webb B, Marti-Renom MA, Madhusudhan MS, Eramian D, Shen MY, Pieper U, Sali A (2006) Comparative protein structure modeling using Modeller. Curr Protoc Bioinform Chapter 5, Unit 5 6
Laskowski RA, Rullmannn JA, MacArthur MW, Kaptein R, Thornton JM (1996) AQUA and PROCHECK-NMR: programs for checking the quality of protein structures solved by NMR. J Biomol NMR 8:477–486
Bashford D (1997) An object-oriented programming suite for electrostatic effects in biological molecules An experience report on the MEAD project. Sci Comput Object Oriented Parallel Environ Lect Notes Comput Sci 1343:233–240
Bashford D, Gerwert K (1992) Electrostatic calculations of the pKa values of ionizable groups in bacteriorhodopsin. J Mol Biol 224:473–486
Phillips JC, Braun R, Wang W, Gumbart J, Tajkhorshid E, Villa E, Chipot C, Skeel RD, Kale L, Schulten K (2005) Scalable molecular dynamics with NAMD. J Comput Chem 26:1781–1802
Kalé L, Skeel R, Bhandarkar M, Brunner R, Gursoy A, Krawetz N, Phillips J, Shinozaki A, Varadarajan K, Schulten K (1999) NAMD2: greater scalability for parallel molecular dynamics. J Comput Phys 151:283–312
MacKerell AD, Bashford D, Bellott D, Dunbrack RL (1998) All-atom empirical potential for molecular modeling and dynamics studies of proteins. J Phys Chem B 102:3586–3616
Park S, Schulten K (2004) Calculating potentials of mean force from steered molecular dynamics simulations. J Chem Phys 120:5946–5961
Rathore N, Yan Q, de Pablo JJ (2004) Molecular simulation of the reversible mechanical unfolding of proteins. J Chem Phys 120:5781–5788
Park S, Khalili-Araghi F, Tajkhorshid E, Schulten K (2003) Free energy calculation from steered molecular dynamics simulations using Jarzynski’s equality. J Chem Phys 119:8
Pang C, Cao T, Li J, Jia M, Zhang S, Ren S, An H, Zhan Y (2013) Combining fragment homology modeling with molecular dynamics aims at prediction of Ca binding sites in CaBPs. J Comput Aided Mol Des 27:697–705
Ferrera L, Caputo A, Ubby I, Bussani E, Zegarra-Moran O, Ravazzolo R, Pagani F, Galietta LJ (2009) Regulation of TMEM16A chloride channel properties by alternative splicing. J Biol Chem 284:33360–33368
Zhang JL, Zheng QC, Li ZQ, Zhang HX (2012) Molecular dynamics simulations suggest ligand’s binding to nicotinamidase/pyrazinamidase. PLoS ONE 7:e39546
Park S, Schulten K (2009) Calculating potentials of mean force from steered molecular dynamics simulations. J Chem Phys 120:5946–5961
Falke JJ, Drake SK, Hazard AL, Peersen OB (1994) Molecular tuning of ion binding to calcium signaling proteins. Q Rev Biophys 27:219–290
Hartzell HC, Yu K, Xiao Q, Chien LT, Qu Z (2009) Anoctamin/TMEM16 family members are Ca2+-activated Cl- channels. J Physiol 587:2127–2139
Acknowledgments
The work is supported by National Nature Science Foundation of China Grants 11175055 to YZ, 113471215 to SZ.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Pang, CL., Yuan, HB., Cao, TG. et al. Molecular simulation assisted identification of Ca2+ binding residues in TMEM16A. J Comput Aided Mol Des 29, 1035–1043 (2015). https://doi.org/10.1007/s10822-015-9876-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-015-9876-x