Abstract
The H7N9 avian influenza virus is a novel re-assortment from at least four different strains of virus. Neuraminidase, which is a glycoprotein on the surface membrane, has been the target for drug treatment. However, some H7N9 strains that have been isolated from patient after drug treatment have a R292K mutation in neuraminidase. This substitution was found to facilitate drug resistance using protein- and virus- assays, in particular it gave a high resistance to the most commonly used drug, oseltamivir. The aim of this research is to understand the source of oseltamivir resistance using MD simulations and the MM/PB(GB)SA binding free energy approaches. Both methods can predict the reduced susceptibility of oseltamivir in good agreement to the IC 50 binding energy, although MM/GBSA underestimates this prediction compared to the MM/PBSA calculation. Electrostatic interaction is the main contribution for oseltamivir binding in terms of both interaction and solvation. We found that the source of the drug resistance is a decrease in the binding interaction combined with the reduction of the dehydration penalty. The smaller K292 mutated residue has a larger binding pocket cavity compared to the wild-type resulting in the loss of drug carboxylate-K292 hydrogen bonding and an increased accessibility for water molecules around the K292 mutated residue. In addition, oseltamivir does not bind well to the R292K mutant complex as shown by the high degree of fluctuation in ligand RMSD during the simulation and the change in angular distribution of bulky side chain groups.
Similar content being viewed by others
Introduction
The novel H7N9 influenza A virus (IAV) caused an outbreak of human infection in China in 2013. From March 2013 to May 2014, this IAV caused 440 human infections of which ~27 % were fatal [1]. Since then, this virus has additionally had 11 more confirmed human cases resulting in 4 more deaths on the 10 May 2016 (WHO). This new virus occurred by the multiple re-assortment of at least four avian influenza viruses; hemagglutinin subtype 7 (H7), neuraminidase subtype 9 (N9) and other six gene segments from two different groups of avian H9N2 viruses [2, 3]. The cause of poultry-to-human transmission is the mutation or genetic changes in domestic poultry, after which human can be exposed to infected poultry or contaminated environment in live poultry markets [2, 4–7]. Although there is no evidence for human-to-human transmission, the super re-assortment of this novel IVA is unpredictable in its emergence and incites the need for vaccine development [8, 9].
Many researches indicate that cross species transmission from poultry to humans is instigated by several mutations of proteins including the mutations in receptor binding site for hemagglutinin [10]. Such mutations include, the Q226L and G186V substitutions in hemagglutinin that result in stronger binding affinity to the human receptor [3], the mutation of a PB2 gene segment of E627K which is important for RNA replication in mammalian cell and the deletion of five amino acids (residue 69–73) in the stalk region of neuraminidase [2], which causes an enhancement in poultry to human adaptation [11].
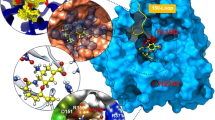
Unlike in the case of the influenza virus where neuraminidase (NA) and the M2 ion-channel can be targeted, here with H7N9 influenza virus due to the S31N mutation only the neuraminidase can be targeted for effective treatment of the virus [2, 9, 12]. The four drugs that act as neuraminidase inhibitors (NAIs) are oseltamivir, zanamivir, peramivir and laninamivir (the latter of which is approved only in Japan). Neuraminidases are classified into two groups based on their primary sequence, except for N10 from bat-origin influenza-like virus and influenza B neuraminidase [13]. Group-1 NAs are N1, N4, N5 and N8 group-2 NAs are N2, N3, N6, N7 and N9 [14]. Whilst N1 and N2 are popular serotypes, generally found in epidemic and pandemic influenza viruses, this is a first time that N9 has infected human [15]. Sequence analysis of neuraminidase from some H7N9 strains have been shown to contain the R292K substitution after oseltamivir treatment such as in A/Shanghai/1/2013 [2, 16]. This R292K substitution is associated with multidrug resistance, particularly for the commonly used oseltamivir [15]. The arginine residue 292 (R292) is one of the arginine triad members (R118 and R371 in N2 numbering) located in the neuraminidase active site and directly interacts with the carboxylate group of sialic acid or with other drugs that bind in a similar fashion to sialic acid (see Fig. 1). In general, the R292K substitution, which causes the resistance of multi NAIs, is mostly found in the N2 and N9 neuraminidase subtypes [17, 18]. The novel H7N9 containing R292K mutation can be treated by NAIs, however, the protein- and virus-based assays clearly indicated drug resistance to oseltamivir and peramivir and a mild resistance to zanamivir and laninamivir, as listed in Table 1. Of particular importance is oseltamivir as it is used frequently in clinical treatment.
Molecular dynamics (MD) simulation has been used successfully to elucidate the dynamics behaviors and equilibrium structures of biomolecules; whilst the calculated binding free energy predicts the strength of ligand binding affinity toward the protein target [19–22]. A combination of these two methods could result in better understanding of drug binding mechanism and source of drug resistance, which the obtained information is further useful for designing and developing the more potent inhibitors [23–25]. Although MD simulations of drug-H7N9 neuraminidase complexes have previously been carried out by others and their binding affinities predicted [26, 27], their initial structures were prepared based on homology modeling. Since then the crystal structure of H7N9 neuraminidases have been made available. This work aims to understand oseltamivir binding to the novel H7N9 neuraminidase and provide insight into the role of the R292K mutation at the molecular level by using MD simulations and binding free energy calculations of molecular mechanics combined with Poisson–Boltzmann (PB) or generalized Born (GB) electrostatics solvation and solvent accessible surface area nonpolar solvation (SASA); MM/PB(GB)SA methods. These well-known binding free energy approaches have been successful applied to various ligand binding affinities and drug resistances [28–42].
Materials and methods
Complex preparation
Oseltamivir binds to both wild-type and R292K mutation H7N9 neuraminidase. Complexes were prepared from crystal structure of the unbound neuraminidase Protein Data Bank (PDB) structures, with codes of 4MWJ and 4MWL respectively, with superimposition to their oseltamivir structure of 4MWQ and 4MWW respectively [15]. These systems were then prepared for calculation and carried out using the AMBER 14 software package [46]. The partial charges and parameters of oseltamivir was obtained from previous work [47], and the force fields of the protein were set to the ff12SB model of Amber 14. The ionized states of all side chains of lysine (K), arginine (R), aspartate (D), glutamate (E) and histidine (H) were configured to be at pH 7.0 using PROPKA3.1 [48]. All disulfide bonds between pairs of cysteine (C) residues were defined to stabilize the protein structure. The LEaP module added the missing hydrogen atoms in the oseltamivir-NA complex. A crystal calcium ion was kept, while water molecules were deleted. The placevent algorithm based on the 3D-RISM distribution function was employed to firstly solvate TIP3P water model around solute molecule [49]. The TIP3P waters were then filled with a minimum distance of 10 Å from the solute to the edge of the simulation box (88 × 90 × 81 Å3) and two Cl− ions were added to neutralize the system.
MD simulation and protocol
To optimize the initial structure before performing MD simulations, the complexes were minimized in the following order, hydrogen atoms, water molecules and then the entire system implementing a 1000 steps using the steepest descent algorithm followed by 2000 steps with the conjugated gradient algorithm. The energy minimization process and MD simulations were performed using the pmemd. CUDA module of AMBER. All bonds involving hydrogen atoms were constrained using the SHAKE algorithm [50]. To reduce the computational time of non-bonded interactions, a cutoff distance of 10 Å was used and the particle mesh Ewald method [51] was employed to calculate the long-range electrostatic interaction.
The simulation employed the periodic boundary condition using Langevin dynamics with a collision frequency of 2 ps−1 and a simulation time step of 2 fs. The system was heated up from 10 to 300 K for 100 ps using canonical ensemble (NVT) in the thermalization step. Subsequently, the equilibration step was firstly conducted for 100 ps of NVT followed by NPT ensemble at 1 atm and 300 K until 30 ns. Finally, the system was further simulated for 50 ns. The production phase of the root mean square displacement (RMSD) analysis was assigned from 30 to 50 ns.
MM/PB(GB)SA binding free energy calculations
The binding free energy is the free energy difference of three reactants; protein, ligand and complex, according to Eq. 1. The binding free energy between protein and ligand is the summation of free energy change during ligand binding \((\Delta G_{MM}^{{}} )\) and the solvation free energy \((\Delta \Delta G_{solv}^{{}} )\).
The molecular mechanical energy is the combination of electrostatic and van der Waals interactions as outlined in Eq. 2, similar to the solvation free energy it can be decomposed into electrostatic and nonpolar contributions as showed in Eq. 3. The MM/PBSA and MM/GBSA methods [32, 52–60] compute the polar solvation free energy by Poisson–Boltzmann or generalized Born models, respectively, whereas their nonpolar solvation energy was obtained by solvent accessible surface area (SASA) using a probe radius of 1.4 Å and a surface tension constant (γ) of 0.0072 kcal mol−1 Å2 according to Eq. 4. The binding free energy calculation was averaged over 100 snapshots extracted from the production phase using the MMPBSA.py program [61]. The electrostatic solvation free energies are calculated with a manner of each method. MM/PBSA computes the polar contribution by solving the linear Poisson–Boltzmann equation using the pbsa module implemented in sander. The grid spacing was set to 0.125 Å and a probe radius of water molecule is of 1.4 Å. In contrast for MM/GBSA, the polar solvation free energy is carried out with the Generalized Born (GB) model developed by Onufriev and colleagues (igb = 2) [62] used in this study. The dielectric constants for solute and solvent of 1 and 80 are applied. To reduce the high computer consuming, the structural entropy calculation is estimated over the 25 snapshots of the MD structures by normal mode analysis in the gas phase [63]. Each structure was entirely minimized for 1000 steps until reaching a local minimum by considering energy gradient with less than 0.01 kcal mol−1 Å−1. The Hessian matrix is diagonalized to obtain the vibrational frequencies, while the translational and vibrational entropies are computed by statistical mechanics formula [53, 64].
Results and discussion
-
Stability of complexes
The stabilities of each complex were evaluated using root-mean square displacement (RMSD) to explore the maintained fluctuation of the MD structure compared to their initial structure. In Fig. 2, the RMSD was calculated in terms of backbone atoms, which belong to the neuraminidase protein and the ligand atoms of oseltamivir versus simulation time. For backbone RMSD, both wild-type and R292K complexes show increasing RMSD from ~0.5 to 1.0 Å during the first 10 ns and after which it oscillates at this value until 30 ns. Finally, the fluctuation stabilizes after 30 ns. With regards to the ligand RMSD of oseltamivir the values varied more ranging from 0.5 to 2.0 Å for the entire simulation. It can be clearly seen that oseltamivir structure of R292K system has significantly higher fluctuations compared to the wild-type system. As a result, the MD structures from 30 to 50 ns were adopted as the production phase for further analysis.
-
Binding free energy calculation of drug susceptibility
The aspects of this study are to understand the oseltamivir efficiency towards the H7N9 neuraminidases and to explore the source of reduce susceptibility concerning the R292K mutation. The MM/PBSA and MM/GBSA binding free energies and their components are shown in Table 2 and are also compared to the experimental values.
In the wild-type complex, MM/PBSA method qualitatively predicts the binding free energy of −14.7 kcal mol−1, which is consistent with the experimental values of −12 to −13 kcal mol−1 [15, 43–45], while MM/GBSA approach underestimates the value with a prediction of −28.5 kcal mol−1. The total binding free energy (\(\Delta G_{bind}\)) of these methods is a subtle balance between the negative value of the binding interaction (\(\Delta G_{MM}\)) and the positive value of the solvation free energy (\(\Delta \Delta G_{solv}\)) called the dehydration penalty which involves the loss of solvation around the protein after ligand binding and is clarified in more detail in a previous study [65]. The electrostatic contribution plays a major role for oseltamivir binding as can be seen from the large free energy values of the electrostatic interaction \(\Delta E_{ele}^{MM}\) and the polar solvation free energy \(\Delta \Delta G_{solv}^{polar}\). This is due to the zwitter ion nature of oseltamivir accommodating the highly charged binding pocket of neuraminidase.
The increase in magnitude of binding free energy in the R292K mutated system was calculated for MM/PBSA (−4.5 kcal mol−1) and MM/GBSA (−19.8 kcal mol−1) to be 10.2 and 8.7 kcal mol−1, respectively, compared to those of wild-type. Similar to the wild-type complex, the MM/GBSA method underestimates the prediction of the binding free energy compared to the MM/PBSA approach (experimental values of −5.2 or −7.2 kcal mol-1). The source of reduced oseltamivir susceptibility conferred by the R292K substitution is a reduction in binding interaction (negative value) and solvation free energy (positive energy). Both methodologies show a loss of drug interaction by 18.5 kcal mol−1, and a decrease in dehydration penalty resulting in a change of solvation energy by 8.3 kcal mol−1 for MM/PBSA and 9.8 kcal mol−1 for MM/GBSA. In the following sections we will further elucidate the effect of the R292K substitution towards oseltamivir binding.
-
Change of drug binding interaction
The favorable electrostatic interaction of oseltamivir binding is obtained from strong hydrogen bond (H-bond) formations between polar moieties of the drug (including –COO−, –NHAc and –NH3 +) and the highly charged binding pocket of neuraminidase. In addition, the nonpolar side chain of the pentyl group gives van der Waals contribution stabilizing the hydrophobic pocket formed by the E276 and R224 residues. The H-bond formation is illustrated as a percentage occupation, where occupation is defined as having maximum distance of 3.5 Å between hydrogen donor (D) and acceptor (A) together with the minimum bond angle of 120° between three atoms (D…H…A) depicted in Fig. 3 for the wild-type and the R292K mutant neuraminidases.
Percentage occupation of the hydrogen bond interactions between oseltamivir function groups and the H7N9 neuraminidase binding residues for a wild-type and b R292K strains. Also shown is a close up of oseltamivir in the neuraminidase binding pocket of c wild-type and d R292K strain taken from the last snapshot
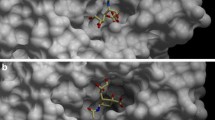
In Fig. 3, oseltamivir forms H-bond interaction with residues of the arginine triad (R118, R292 and R371, according to N2 numbering), R152, E119 and D151 as is generally found to be the case in other neuraminidase subtypes [21, 33, 47, 65]. In the R292K complex, there is a complete loss of the carboxyalte-K292 interaction as previously found in the H1N1 subtype [21], however, this is compensated for by the R118 interaction (Fig. 3b). A smaller side chain of lysine (K) compared to arginine (R) at position 292 cannot form the necessary H-bond with the carboxylate group (distance of around 5 Å), leading to the larger cavity and shifting toward the R118 residue of ligand (Fig. 3d). It is worth noting that the conformation of this K292 mutated residue does not perturb the formation of hydrophobic pocket made by E276 and R244 residues as found in the crystal structure [15, 27].
The conformational change of the pentyl group was evaluated by the distribution of torsion angles (Fig. 4). Both wild-type and R292K systems have different angular distributions, particularly for τz. The torsion distribution of τz in the mutant system dramatically changes from the normal ~60° of the wild-type, while τx and τy show the same pattern but different distributions. This suggests that the drug in the R292K complex is oscillating. This oscillation can clearly be seen from the high fluctuation of ligand RMSD during simulation (in Fig. 2). The twist of pentyl group was found to disrupt the hydrophobic binding in a previous study [26].
To summarize, the unsuitable accommodation of oseltamivir in R292K neuraminidase results in a decrease of drug binding interaction by 13.8 kcal mol−1 for electrostatic interaction and 5.1 kcal mol−1 for van der Waals interaction compared to the wild-type complex.
-
Change of solvation conferring R292K substitution
The latter source of the high-level oseltamivir resistance toward R292K substitution is due to the decrease of the positive contribution of the solvation free energy resulting in a smaller dehydration penalty compared to the wild-type. As mentioned in previous sections, the shorter side chain of K292 causes poor binding with the drug carboxylate group and a larger cavity. This results in greater accessibility for water molecules towards the neuraminidase binding pocket as evident by the radial distribution function (RDF) around oseltamivir heteroatoms and residue 292 (lables shown in Fig. 3c, d) depicted in Fig. 5.
In Fig. 5a, the first peak of water distribution around oseltamivir heteroatoms occurs at 3.5 Å and is mostly sharp, which suggests that the accessible water molecules strongly interact with oseltamivir. The nonzero value at the terminal of the first peak as found in the O1, O2 and N1 graphs indicate a high degree of transfer of water molecules in this shell. Switching of water molecules around the oseltamivir molecule was found to occur between the O1 and O2 atoms at the oseltamivir carboxylate group (see illustration in Fig. 3c, d), while the rest of the heteroatoms are conserved for both wild-type and R292K systems. Additionally two water molecules were found to bind around the K292 residue, compared to the one molecule of the wild-type (Fig. 5b). This mutated residue was strongly solvated by three water molecules as can be seen by the approximately close to zero values at the fist minimum, which is in good agreement with the crystal structures of H7N9 and H11N9 [15, 66]. Thus, the R292K mutation of neuraminidase can affect the accessibility of water in the binding pocket, which causes a decreased dehydration penalty.
Conclusion
In this study, oseltamivir binding towards the novel H7N9 influenza neuraminidase and the high resistance R292K substitution was investigated using MD simulations and MM/PB(GB)SA binding free energy calculations. MM/PB(GB)SA methods were able to predict the susceptibility of oseltamivir toward both the wild-type and the R292K strain and is in good agreement with the energy converted from the IC 50 data, of which the MM/PBSA approach was shown to give a more reliable prediction. A structural entropy calculation was necessary to improve the qualitative of the predicted binding free energy. Similar to the previous study of influenza B neuraminidase, the main factor for oseltamivir binding toward H7N9 neuraminidase is a subtle balance of the large negative value of binding interaction and the large positive contribution of solvation conferring the dehydration penalty during drug binding. The major interaction of oseltamivir binding is the electrostatic contribution, as is generally found for all neuraminidase subtypes, caused by the opposite charges of oseltamivir and neuraminidase binding residues. A dramatic increase in magnitude of binding free energy was found in the R292K system compared to the wild-type. The role of R292K mutation in H7N9 neuraminidase is to decrease the binding interaction as was shown by the high fluctuation of drug RMSD in the mutant pocket and the reduction of the dehydration penalty resulting in an increased accessibility for water molecules around the K292 mutated residue and the carboxylate group of drug. The results from this work suggest that a hydrophilic bulky group as found in zanamivir and laninamivir, instead of the pentyl group of oseltamivir, promises to be a potent neuraminidase inhibitor by maintaining a strong interaction toward residue 292 of both arginine and lysine amino acids.
References
Watanabe T, Watanabe S, Maher EA, Neumann G, Kawaoka Y (2014) Pandemic potential of H7N9 influenza viruses. Trends Microbiol 22:623–631
Gao R, Cao B, Hu Y, Feng Z, Wang D, Hu W, Chen J, Jie Z, Qiu H, Xu K, Xu X, Lu H, Zhu W, Gao Z, Xiang N, Shen Y, He Z, Gu Y, Zhang Z, Yang Y, Zhao X, Zhou L, Li X, Zou S, Zhang Y, Li X, Yang L, Guo J, Dong J, Li Q, Dong L, Zhu Y, Bai T, Wang S, Hao P, Yang W, Zhang Y, Han J, Yu H, Li D, Gao GF, Wu G, Wang Y, Yuan Z, Shu Y (2013) Human infection with a novel avian-origin influenza A (H7N9) virus. N Engl J Med 368:1888–1897
Liu D, Shi W, Shi Y, Wang D, Xiao H, Li W, Bi Y, Wu Y, Li X, Yan J, Liu W, Zhao G, Yang W, Wang Y, Ma J, Shu Y, Lei F, Gao GF (2013) Origin and diversity of novel avian influenza A H7N9 viruses causing human infection: phylogenetic, structural, and coalescent analyses. The Lancet 381:1926–1932
Pantin-Jackwood MJ, Miller PJ, Spackman E, Swayne DE, Susta L, Costa-Hurtado M, Suarez DL (2014) Role of poultry in spread of novel H7N9 influenza virus in China. J Virol 88:5381–5390
Richard M, Schrauwen EJA, de Graaf M, Bestebroer TM, Spronken MIJ, van Boheemen S, de Meulder D, Lexmond P, Linster M, Herfst S, Smith DJ, van den Brand JM, Burke DF, Kuiken T, Rimmelzwaan GF, Osterhaus ADME, Fouchier RAM (2013) Limited airborne transmission of H7N9 influenza A virus between ferrets. Nature 501:560–563
Shi J, Xie J, He Z, Hu Y, He Y, Huang Q, Leng B, He W, Sheng Y, Li F, Song Y, Bai C, Gu Y, Jie Z (2013) A detailed epidemiological and clinical description of 6 human cases of avian-origin influenza A (H7N9) virus infection in Shanghai. PLoS One 8:e77651
van Riel D, Leijten LME, de Graaf M, Siegers JY, Short KR, Spronken MIJ, Schrauwen EJA, Fouchier RAM, Osterhaus ADME, Kuiken T (2013) Novel avian-origin influenza A (H7N9) virus attaches to epithelium in both upper and lower respiratory tract of humans. Am J Pathol 183:1137–1143
Tan K-X, Jacob SA, Chan K-G, Lee L-H (2015) An overview of the characteristics of the novel avian influenza A H7N9 virus in humans. Front Microbiol 6:140
Qi Y, Fan H, Qi X, Zhu Z, Guo X, Chen Y, Ge Y, Zhao K, Wu T, Li Y, Shan Y, Zhou M, Shi Z, Wang H, Cui L (2014) A novel pyrosequencing assay for the detection of neuraminidase inhibitor resistance-conferring mutations among clinical isolates of avian H7N9 influenza virus. Virus Res 179:119–124
Shi Y, Zhang W, Wang F, Qi J, Wu Y, Song H, Gao F, Bi Y, Zhang Y, Fan Z, Qin C, Sun H, Liu J, Haywood J, Liu W, Gong W, Wang D, Shu Y, Wang Y, Yan J, Gao GF (2013) Structures and receptor binding of hemagglutinins from human-infecting H7N9 influenza viruses. Science 342:243–247
Matrosovich M, Zhou N, Kawaoka Y, Webster R (1999) The surface glycoproteins of H5 influenza viruses isolated from humans, chickens, and wild aquatic birds have distinguishable properties. J Virol 73:1146–1155
Li Q, Zhou L, Zhou M, Chen Z, Li F, Wu H, Xiang N, Chen E, Tang F, Wang D, Meng L, Hong Z, Tu W, Cao Y, Li L, Ding F, Liu B, Wang M, Xie R, Gao R, Li X, Bai T, Zou S, He J, Hu J, Xu Y, Chai C, Wang S, Gao Y, Jin L, Zhang Y, Luo H, Yu H, He J, Li Q, Wang X, Gao L, Pang X, Liu G, Yan Y, Yuan H, Shu Y, Yang W, Wang Y, Wu F, Uyeki TM, Feng Z (2014) Epidemiology of human infections with avian influenza A(H7N9) virus in China. N Engl J Med 370:520–532
Li Q, Sun X, Li Z, Liu Y, Vavricka CJ, Qi J, Gao GF (2012) Structural and functional characterization of neuraminidase-like molecule N10 derived from bat influenza A virus. Proc Natl Acad Sci USA 109:18897–18902
Russell RJ, Haire LF, Stevens DJ, Collins PJ, Lin YP, Blackburn GM, Hay AJ, Gamblin SJ, Skehel JJ (2006) The structure of H5N1 avian influenza neuraminidase suggests new opportunities for drug design. Nature 443:45–49
Wu Y, Bi Y, Vavricka CJ, Sun X, Zhang Y, Gao F, Zhao M, Xiao H, Qin C, He J, Liu W, Yan J, Qi J, Gao GF (2013) Characterization of two distinct neuraminidases from avian-origin human-infecting H7N9 influenza viruses. Cell Res 23:1347–1355
Hu Y, Lu S, Song Z, Wang W, Hao P, Li J, Zhang X, Yen H-L, Shi B, Li T, Guan W, Xu L, Liu Y, Wang S, Zhang X, Tian D, Zhu Z, He J, Huang K, Chen H, Zheng L, Li X, Ping J, Kang B, Xi X, Zha L, Li Y, Zhang Z, Peiris M, Yuan Z (2013) Association between adverse clinical outcome in human disease caused by novel influenza A H7N9 virus and sustained viral shedding and emergence of antiviral resistance. The Lancet 381:2273–2279
McKimm-Breschkin JL (2000) Resistance of influenza viruses to neuraminidase inhibitors—a review. Antiviral Res 47:1–17
Yen H-L, Hoffmann E, Taylor G, Scholtissek C, Monto AS, Webster RG, Govorkova EA (2006) Importance of neuraminidase active-site residues to the neuraminidase inhibitor resistance of influenza viruses. J Virol 80:8787–8795
Rungrotmongkol T, Mulholland AJ, Hannongbua S (2007) Active site dynamics and combined quantum mechanics/molecular mechanics (QM/MM) modelling of a HIV-1 reverse transcriptase/DNA/dTTP complex. J Mol Graph Model 26:1–13
Decha P, Rungrotmongkol T, Intharathep P, Malaisree M, Aruksakunwong O, Laohpongspaisan C, Parasuk V, Sompornpisut P, Pianwanit S, Kokpol S, Hannongbua S (2008) Source of high pathogenicity of an avian influenza virus H5N1: why H5 is better cleaved by furin. Biophys J 95:128–134
Rungrotmongkol T, Malaisree M, Nunthaboot N, Sompornpisut P, Hannongbua S (2010) Molecular prediction of oseltamivir efficiency against probable influenza A (H1N1-2009) mutants: molecular modeling approach. Amino Acids 39:393–398
Khuntawee W, Rungrotmongkol T, Hannongbua S (2011) Molecular dynamic behavior and binding affinity of flavonoid analogues to the cyclin dependent kinase 6/cyclin D complex. J Chem Inf Model 52:76–83
Rungrotmongkol T, Frecer V, De-Eknamkul W, Hannongbua S, Miertus S (2009) Design of oseltamivir analogs inhibiting neuraminidase of avian influenza virus H5N1. Antiviral Res 82:51–58
Rungrotmongkol T, Udommaneethanakit T, Frecer V, Miertus S (2010) Combinatorial design of avian influenza neuraminidase inhibitors containing pyrrolidine core with a reduced susceptibility to viral drug resistance. Comb Chem High Throughput Screening 13:268–277
Kongkamnerd J, Cappelletti L, Prandi A, Seneci P, Rungrotmongkol T, Jongaroonngamsang N, Rojsitthisak P, Frecer V, Milani A, Cattoli G, Terregino C, Capua I, Beneduce L, Gallotta A, Pengo P, Fassina G, Miertus S, De-Eknamkul W (2012) Synthesis and in vitro study of novel neuraminidase inhibitors against avian influenza virus. Biorgan Med Chem 20:2152–2157
Woods CJ, Malaisree M, Long B, McIntosh-Smith S, Mulholland AJ (2013) Computational assay of H7N9 influenza neuraminidase reveals R292K mutation reduces drug binding affinity. Sci Rep 3:3561
Tran-To SuC, Ouyang X, Zheng J, Kwoh C-K (2013) Structural analysis of the novel influenza A (H7N9) viral neuraminidase interactions with current approved neuraminidase inhibitors oseltamivir, zanamivir, and peramivir in the presence of mutation R289K. BMC Bioinformatics 14:S7
Hou T, Yu R (2007) Molecular dynamics and free energy studies on the wild-type and double mutant HIV-1 protease complexed with amprenavir and two amprenavir-related inhibitors: mechanism for binding and drug resistance. J Med Chem 50:1177–1188
Hou T, McLaughlin WA, Wang W (2008) Evaluating the potency of HIV-1 protease drugs to combat resistance. Proteins Struct Funct Bioinform 71:1163–1174
Rungrotmongkol T, Nunthaboot N, Malaisree M, Kaiyawet N, Yotmanee P, Meeprasert A, Hannongbua S (2010) Molecular insight into the specific binding of ADP-ribose to the nsP3 macro domains of chikungunya and venezuelan equine encephalitis viruses: molecular dynamics simulations and free energy calculations. J Mol Graph Model 29:347–353
Liu H, Yao X, Wang C, Han J (2010) In silico identification of the potential drug resistance sites over 2009 influenza A (H1N1) virus neuraminidase. Mol Pharm 7:894–904
Hou T, Wang J, Li Y, Wang W (2011) Assessing the performance of the MM/PBSA and MM/GBSA methods. 1. The accuracy of binding free energy calculations based on molecular dynamics simulations. J Chem Inf Model 51:69–82
Kar P, Knecht V (2012) Mutation-induced loop opening and energetics for binding of tamiflu to influenza N8 neuraminidase. J Phys Chem B 116:6137–6149
Li L, Li Y, Zhang L, Hou T (2012) Theoretical studies on the susceptibility of oseltamivir against variants of 2009 A/H1N1 influenza neuraminidase. J Chem Inf Model 52:2715–2729
Pan P, Li L, Li Y, Li D, Hou T (2013) Insights into susceptibility of antiviral drugs against the E119G mutant of 2009 influenza A (H1N1) neuraminidase by molecular dynamics simulations and free energy calculations. Antiviral Res 100:356–364
Barakat KH, Law J, Prunotto A, Magee WC, Evans DH, Tyrrell DL, Tuszynski J, Houghton M (2013) Detailed computational study of the active site of the hepatitis C viral RNA polymerase to aid novel drug design. J Chem Inf Model 53:3031–3043
Tzoupis H, Leonis G, Mavromoustakos T, Papadopoulos MG (2013) A comparative molecular dynamics, MM–PBSA and thermodynamic integration study of saquinavir complexes with wild-type HIV-1 PR and L10I, G48V, L63P, A71V, G73S, V82A and I84V single mutants. J Chem Theory Comput 9:1754–1764
Guan Y, Sun H, Li Y, Pan P, Li D, Hou T (2014) The competitive binding between inhibitors and substrates of HCV NS3/4A protease: a general mechanism of drug resistance. Antiviral Res 103:60–70
Meeprasert A, Hannongbua S, Rungrotmongkol T (2014) Key binding and susceptibility of NS3/4A serine protease inhibitors against hepatitis C virus. J Chem Inf Model 54:1208–1217
Sun H, Li Y, Tian S, Wang J, Hou T (2014) P-loop conformation governed crizotinib resistance in G2032R-mutated ROS1 tyrosine kinase: clues from free energy landscape. PLoS Comput Biol 10:e1003729
Tzoupis H, Leonis G, Avramopoulos A, Mavromoustakos T, Papadopoulos MG (2014) Systematic molecular dynamics, MM-PBSA, and ab initio approaches to the saquinavir resistance mechanism in HIV-1 PR due to 11 double and multiple mutations. J Phys Chem B 118:9538–9552
Li N, Ainsworth RI, Ding B, Hou T, Wang W (2015) Using hierarchical virtual screening to combat drug resistance of the HIV-1 protease. J Chem Inf Model 55:1400–1412
Yen H-L, McKimm-Breschkin JL, Choy K-T, Wong DDY, Cheung PPH, Zhou J, Ng IH, Zhu H, Webby RJ, Guan Y, Webster RG, Peiris JSM (2013) Resistance to neuraminidase inhibitors conferred by an R292K mutation in a human influenza virus H7N9 isolate can be masked by a mixed R/K viral population. MBio 4:1–8
Sleeman K, Guo Z, Barnes J, Shaw M, Stevens J, Gubareva LV (2013) R292K substitution and drug susceptibility of influenza A(H7N9) viruses. Emerg Infect Dis 19:1521–1524
Hai R, Schmolke M, Leyva-Grado VH, Thangavel RR, Margine I, Jaffe EL, Krammer F, Solorzano A, Garcia-Sastre A, Palese P, Bouvier NM (2013) Influenza A(H7N9) virus gains neuraminidase inhibitor resistance without loss of in vivo virulence or transmissibility. Nat Commun 4:2854
Case DA, Babin V, Berryman J, Betz RM, Cai Q, Cerutti DS, Cheatham III TE, Darden TA, Duke RE, Gohlke H, Goetz AW, Gusarov S, Homeyer N, Janowski P, Kaus J, Kolossváry I, Kovalenko A, Lee TS, LeGrand S, Luchko T, Luo R, Madej B, Merz KM, Paesani F, Roe DR, Roitberg A, Sagui C, Salomon-Ferrer R, Seabra G, Simmerling CL, Smith W, Swails J, Walker RC, Wang J, Wolf RM, Wu X, Kollman PA (2014) AMBER 14
Malaisree M, Rungrotmongkol T, Decha P, Intharathep P, Aruksakunwong O, Hannongbua S (2008) Understanding of known drug-target interactions in the catalytic pocket of neuraminidase subtype N1. Proteins Struct Funct Bioinform 71:1908–1918
Olsson MH, Sondergaard CR, Rostkowski M, Jensen JH (2011) PROPKA3: consistent treatment of internal and surface residues in empirical pKa predictions. J Chem Theory Comput 7:525–537
Sindhikara DJ, Yoshida N, Hirata F (2012) Placevent: an algorithm for prediction of explicit solvent atom distribution-application to HIV-1 protease and F-ATP synthase. J Comput Chem 33:1536–1543
Ryckaert J-P, Ciccotti G, Berendsen HJC (1977) Numerical integration of the cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. J Comput Phys 23:327–341
York DM, Darden TA, Pedersen LG (1993) The effect of long-range electrostatic interactions in simulations of macromolecular crystals: a comparison of the Ewald and truncated list methods. J Chem Phys 99:8345–8348
Srinivasan J, Cheatham TE, Cieplak P, Kollman PA, Case DA (1998) Continuum solvent studies of the stability of DNA, RNA, and Phosphoramidate-DNA helices. J Am Chem Soc 120:9401–9409
Kollman PA, Massova I, Reyes C, Kuhn B, Huo S, Chong L, Lee M, Lee T, Duan Y, Wang W, Donini O, Cieplak P, Srinivasan J, Case DA, Cheatham TE (2000) Calculating structures and free energies of complex molecules: combining molecular mechanics and continuum models. Acc Chem Res 33:889–897
Hou T, Wang J, Li Y, Wang W (2011) Assessing the performance of the molecular mechanics/Poisson Boltzmann surface area and molecular mechanics/generalized Born surface area methods. II. The accuracy of ranking poses generated from docking. J Comput Chem 32:866–877
Homeyer N, Gohlke H (2012) Free energy calculations by the molecular mechanics Poisson–Boltzmann surface area method. Mol Inform 31:114–122
Hou T, Li N, Li Y, Wang W (2012) Characterization of domain–peptide interaction interface: prediction of SH3 domain-mediated protein–protein interaction network in yeast by generic structure-based models. J Proteome Res 11:2982–2995
Xu L, Sun H, Li Y, Wang J, Hou T (2013) Assessing the performance of MM/PBSA and MM/GBSA methods. 3. The impact of force fields and ligand charge models. J Phys Chem B 117:8408–8421
Sun H, Li Y, Tian S, Xu L, Hou T (2014) Assessing the performance of MM/PBSA and MM/GBSA methods. 4. Accuracies of MM/PBSA and MM/GBSA methodologies evaluated by various simulation protocols using PDBbind data set. Phys Chem Chem Phys 16:16719–16729
Sun H, Li Y, Shen M, Tian S, Xu L, Pan P, Guan Y, Hou T (2014) Assessing the performance of MM/PBSA and MM/GBSA methods. 5. Improved docking performance using high solute dielectric constant MM/GBSA and MM/PBSA rescoring. Phys Chem Chem Phys 16:22035–22045
Chen F, Liu H, Sun H, Pan P, Li Y, Li D, Hou T (2016) Assessing the performance of the MM/PBSA and MM/GBSA methods. 6. Capability to predict protein-protein binding free energies and re-rank binding poses generated by protein–protein docking. Phys Chem Chem Phys 18:22129–22139
Miller BR, McGee TD, Swails JM, Homeyer N, Gohlke H, Roitberg AE (2012) MMPBSA.py: an efficient program for end-state free energy calculations. J Chem Theory Comput 8:3314–3321
Onufriev A, Bashford D, Case DA (2004) Exploring protein native states and large-scale conformational changes with a modified generalized born model. Proteins Struct Funct Bioinform 55:383–394
Genheden S, Kuhn O, Mikulskis P, Hoffmann D, Ryde U (2012) The normal-mode entropy in the MM/GBSA method: effect of system truncation, buffer region, and dielectric constant. J Chem Inf Model 52:2079–2088
Jensen F (2006) Introduction to computational chemistry. John Wiley & Sons, England
Phanich J, Rungrotmongkol T, Sindhikara D, Phongphanphanee S, Yoshida N, Hirata F, Kungwan N, Hannongbua S (2016) A 3D-RISM/RISM study of the oseltamivir binding efficiency with the wild-type and resistance-associated mutant forms of the viral influenza B neuraminidase. Protein Sci 25:147–158
Varghese JN, Smith PW, Sollis SL, Blick TJ, Sahasrabudhe A, McKimm-Breschkin JL, Colman PM (1998) Drug design against a shifting target: a structural basis for resistance to inhibitors in a variant of influenza virus neuraminidase. Structure 6:735–746
Acknowledgments
The authors would like to thank Dr. Sunan Kitjaruwankul for useful discussion and Dr. Andrew King for kind proof reading.
Funding
This research is supported by the Rachadaphiseksomphot Endowment Fund Part of the “Strengthen Chulalongkorn University Researcher’s Project”. We would like to thank the Research Chair Grant, the National Science and Technology Development Agency (NSTDA), Thailand. JP is thankful for Science Achievement Scholarship of Thailand (SAST) and the 90th Anniversary of Chulalongkorn University, Rachadaphiseksomphot Endowment Fund for scholarship. TR would like to thank the Structural and Computational Biology Research Group, Special Task Force for Activating Research (STAR), Faculty of Science, Chulalongkorn University, and the Thailand Research Fund (IRG5780008).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Phanich, J., Rungrotmongkol, T., Kungwan, N. et al. Role of R292K mutation in influenza H7N9 neuraminidase toward oseltamivir susceptibility: MD and MM/PB(GB)SA study. J Comput Aided Mol Des 30, 917–926 (2016). https://doi.org/10.1007/s10822-016-9981-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-016-9981-5