Abstract
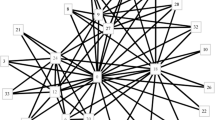
In this paper, we study the following Chromatic kernel (CK) problem: given an \(n\)-partite graph (called a chromatic correlation graph) \(G=(V,E)\) with \(V=V_{1}\bigcup \cdots \bigcup V_{n}\) and each partite set \(V_{i}\) containing a constant number \(\lambda \) of vertices, compute a subgraph \(G[V_{CK}]\) of \(G\) with exactly one vertex from each partite set and the maximum number of edges or the maximum total edge weight, if \(G\) is edge-weighted (among all such subgraphs). CK is a new problem motivated by several applications and no existing algorithm directly solves it. In this paper, we first show that CK is NP-hard even if \(\lambda =2\), and cannot be approximated within a factor of \(16/17\) unless P = NP. Then, we present a random-sampling-based PTAS for dense CK. As its application, we show that CK can be used to determine the pattern of chromosome associations in the nucleus for a population of cells. We test our approach by using random and real biological data; experimental results suggest that our approach yields near optimal solutions, and significantly outperforms existing approaches.



Similar content being viewed by others
Notes
The Jaccard similarity of two unweighted graph \(G_{1}=(V_{1},E_{1})\) and \(G_{2}=(V_{2},E_{2})\) is \(\frac{|E_{1}\bigcap E_{2}|}{|E_{1}\bigcup E_{2}|}\). If \(G_{1}\) and \(G_{2}\) are weighted complete graphs (missing edges can be viewed as edges with \(0\) weight), their Jaccard similarity is \(\frac{\sum _{1\le i\le |E_{1}\cup E_{2}|}\min \{w_{i}, w'_{i}\}}{\sum _{1\le i\le |E_{1}\cup E_{2}|}\max \{w_{i}, w'_{i}\}}.\)
Given a set of sets, Jaccard median is the set (which might be a new set, and not from the given sets) with minimum total Jaccard distances to the given sets. Here, each labeled graph can be viewed as a set of edges.
References
Alon N, Spencer JH (1992) The probabilistic method. Wiley, New Jersey
Andersen R (2010) A local algorithm for finding dense subgraphs. ACM Trans Algoritm 6(4):60
Arora S, Karger DR, Karpinski M (1995) Polynomial time approximation schemes for dense instances of NP-hard problems. STOC 284–293
Arora S, Barak B (2009) Computational complexity: a modern approach. Cambridge University Press, Cambridge
Asahiro Y, Iwama K, Tamaki H, Tokuyama T (1996) Greedily finding a dense subgraph. SWAT’96, pp 136–148
Bansal N, Blum A, Chawla S (2004) Correlation clustering. Mach Learn 56(1–3):89–113
Berezney R (2002) Regulating the mammalian genome: the role of nuclear architecture. Adv Enzym Regul 42:39–52
Charikar M (2000) Greedy approximation algorithms for finding dense components in a graph. APPROX, pp 84–95
Chazelle B, Kingsford C, Singh M (2004) A semidefinite programming approach to side chain positioning with new rounding strategies. INFORMS J Comput 16(4):380–392
Chierichetti F, Kumar R, Pandey S, Vassilvitskii S (2010) Finding the Jaccard median. SODA 293–311
Cremer T (2001) Chromosome territories, nuclear architecture and gene regulation in mamalian cells. Nat Rev Genet 2:292–301
Ding H, Xu J (2011) Solving the chromatic cone clustering problem via mini- mum spanning sphere, ICALP’11, 4–8, Switzerland
Feige U, Kortsarz G, Peleg D (1997) The dense k-subgraph problem. Algorithmica 29:410–421
Goldberg AV (1984) Finding a maximum density subgraph. Tech Rep
Hastad J (2001) Some optimal inapproximability results. J ACM 48(4):798–859
Khuller S, Saha B (2009) On finding dense subgraphs. ICALP 1:597–608
Kingsford C, Chazelle B, Singh M (2005) Solving and analyzing side-chain positioning problems using linear and integer programming. Bioinformatics 21(7):1028–1039
Mukherjee L, Singh V, Peng J, Xu J, Zeitz M, Berezney R (2009) Generalized median graphs: theory and applications. JCO 17(1):21–44
Stojkovic B, Zhu Y, Xu J, Fritz A, Zeitz MJ, Vecerova J, Berezney R (2010) Computing maximum association graph in microscopic nucleus images. MICCAI 2:530–537
Trevisan L, Sorkin G, Sudan M, Williamson D (2000) Gadgets, approximation, and linear programming. SIAM J Comput 29(6):2074–2097
Acknowledgments
This research was partially supported by NSF through Grants IIS-1115220, IIS-1422591, and CCF-1422324.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ding, H., Stojkovic, B., Chen, Z. et al. Chromatic kernel and its applications. J Comb Optim 31, 1298–1315 (2016). https://doi.org/10.1007/s10878-014-9824-z
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10878-014-9824-z