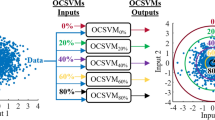
Abstract
Blood pressure measurement based on oscillometry is one of the most popular techniques to check a health condition of individual subjects. This paper proposes a support vector using fusion estimator with a bootstrap technique for oscillometric blood pressure (BP) estimation. However, some inherent problems exist with this approach. First, it is not simple to identify the best support vector regression (SVR) estimator, and worthy information might be omitted when selecting one SVR estimator and discarding others. Additionally, our input feature data, acquired from only five BP measurements per subject, represent a very small sample size. This constitutes a critical limitation when utilizing the SVR technique and can cause overfitting or underfitting, depending on the structure of the algorithm. To overcome these challenges, a fusion with an asymptotic approach (based on combining the bootstrap with the SVR technique) is utilized to generate the pseudo features needed to predict the BP values. This ensemble estimator using the SVR technique can learn to effectively mimic the non-linear relations between the input data acquired from the oscillometry and the nurse’s BPs.
Similar content being viewed by others
Introduction
Blood pressure (BP) is one of the significant vital signs of health. Oscillometric automated BP monitors are now easily and widely available, including in homes, offices and hospitals. The maximum amplitude algorithm (MAA) linearly estimates BP by approximating the mean arterial pressure (ABP) with the cuff pressure at which the maximum amplitude occurs and then using a fixed ratio of empirically derived coefficients [1]. The characteristic fixed ratio of these patients makes it possible to identify when the cuff pressure matches the SBP and DBP, respectively [2]. In fact, SBP is an arterial maximum blood pressure that occurs at the beginning of the cardiac cycle when the heart contracts, and DBP occurs when the heart relaxes between the heartbeat and the minimum blood pressure of the artery at the end of the cardiac cycle [3]. However, the MAA using fixed ratios makes acquiring reliable BP estimation difficult because individual subjects possess their own unique physiological characteristics [4]. To address the problem of fixed ratios in the estimation of BP, a neural network (NN) was performed [3]. However, this scheme did not enough the protocol of standard BP measurements [5]. To follow these recommendations, an advanced machine learning technique is still needed to estimate the BPs without characteristic ratios.
This paper proposes a fusion technique based on a support vector regression (SVR) [6], which can be adopted not only to classify some pattens, but also to estimate target values. Specifically, non-linear functions in SVR are mapped into the derived feature space of high dimensional kernel through linear learning machine, including using the features of the maximum margin algorithm [7]. Therefore, this method can represent the non-linear function between the feature vector collected from the oscillometric waveform (OMW) and the nurse’s BP. Moreover, a general approach is to use the SVR to produce an estimator using a training set with good results. However, there are some disadvantages to this approach. First, it is not simple to determine the best SVR estimator, and worthy information can be omitted when selecting one SVR estimator and discarding the others. Recently, deep networks have become a hot issue in machine learning and signal processing, but this technology requires a lot of time, cost and large data. Additionally, our input feature vectors, obtained from only five measurements per subject constitute a very small sample size, which is a critical weakness while using machine learning techniques, such as the SVR and FFNN. This shortcoming can lead to over-fitting [3]. In order to overcome these problems, we use the bootstrap-aggregation (bagging) technique to generate pseudo-features to estimate the SBP and DBP based on the SVR model [8]. Thus, the number of the pseudo-feature’s sample is drastically increased by employing the bagging technique. The bagging fusion estimator is a strong estimator through assembling a basic estimator that can perform better than random guessing [9], [10], which can effectively learn to mimic the non-linear model between the feature data acquired from the oscillometry and nurse’s BPs. The methodology proposed in the present study is one of the first to use the bagging ensemble regression model using the SVR technique for estimating the BPs. The proposed scheme is to increase accurate BP estimation and mitigate uncertainty BP estimation for a smart healthcare system [11], [12], [13].
This work has the succeeding augmentations and contributions when compared to [3]:
-
This paper introduces a new approach to obtain exact BP estimates using a relatively small sample data from the oscillometry using SVR-based fusion regression estimates.
-
This method can reduce overfitting and unstable estimation, such as large error on the mimic training feature set, through the SVR ensemble regression estimator.
Signal processing and feature extraction from the OMW signals are represented in Section “Signal processing and feature extraction from OMW signals”. The BP estimation based on the fusion technique based on SVR is introduced in Section “BP estimation based on the bootstrap aggregating technique based on SVR”. The results and discussion are presented in Section “Experimental results”. Finally, in Section “Conclusion”, this paper is concluded.
Signal processing and feature extraction from OMW signals
Data collection and preprocessing
This study was initially confirmed by the study committee and all volunteers agreed to provide information prior to the measurements based on the institutional research ethics committee’s BP research program. The BP measurements was collected from 85 volunteers without a history of cardiovascular disease between the ages of 12 and 80 years. Our BP measurement consists of five measurements for each volunteer, with a one minute break between measurements. The oscillometric BP measurement data of this research was provided by a company [16]. Oscillometric BP signals were obtained by detecting pressure pulsations in a subject’s cuff around the biceps or wrist as represented (in the second box on the Fig. 1). Details on OMW signal and BP measurement processing to estimate BP are provided in [16]. In this study, the OMW signals and envelopes were prepared for feature extraction, as shown in the step 3 on the left of Fig. 1. Afterward, the OMW signal is generated using the peak values of the oscillometric pulse. Next, we use the weighted median (WM) filter for smoothing the signal as shown in the step 4. We then use an asymmetric Gaussian fitting to acquire the smoothed envelope as shown in the step 5.
Features acquired from OMWs
In this subsection, we obtained the original features from the OMW signals and compared the target values utilizing the SVR technique. For more information about the feature vectors, please refer to the reference papers [4], [14] (Table 1).
BP estimation based on the bootstrap aggregating technique based on SVR
As mentioned the first section, we use pseudo-features as explanatory data to predict the reference BPs as dependent variables as shown in the step 8. The block diagram of the proposed bagging regression technique based on SVR is shown Fig. 1. It comprises three parts such as left part (from step 1 to step 8), middle part (from step 9 to step 14), and right part (from step 15 to step 18).

Block diagram for BP prediction utilizing the SVR with the bagging fusion estimator. Left boxes (steps 1–8): Steps of BP measurement, preprocessing, feature extraction, and pseudo features generation. Middle boxes (steps 9–14): SVR training and SVM prediction. Right boxes (steps 15–18): Bagging ensemble to increase accurate and decrease uncertainty, where y(1) denotes the output vector of the first estimator, DT(1) denotes the mean vector of the first estimator, and K is the number of SVR estimator
Support vector regression (SVR) estimator
Assume that one has a training data set {(x1,y1),...,(x l ,y l )}∈x i ×\(\mathbb {R}\), where x i denote the training data and y i denote the target BPs. The SVR is to find \( y = \langle \omega ,\phi (x)\rangle _{\mathcal {H}} + b\), where ω and ϕ(x) denote the vectors obtained from reproducing kernel Hilbert space \(\mathcal {H}\). Therefore, we can minimize the following risks:
where \(\mathcal {L}(y_{i}, x_{i}, f) \) denotes the 𝜖-loss function [6] given by
This problem can be solved by the Lagrangian theory as follows:
where \(\{\hat {\alpha }, \alpha \}\), i = 1,...,l denote the Lagrangian multipliers with respect to the constraints given in Eq. (4), and the solutions are given by
where \(K(x_{i},x_{j}) = \langle \phi (x_{i}), \phi (x_{j}) \rangle _{\mathcal {H}}\) and ζi,j represents the Kronecker symbol. This problem can be resolved using some disassemble methods of the SVR [6]. To summarize, the SVR model can automatically achieve the relationship between the nurse’s BPs (RSBP and RDBP) and pseudo feature data, in the SVR training stage as shown in steps 9 to 11. The SBP and DBP in our SVR model are accurately acquired from the training stage as shown in steps 12 to 14.
Bagging technique
Beyond the bootstrap technique, ensemble-based algorithms using the bagging technique have been developed [8]. In this paper, we use the bagging approach to improve the performance of the SVR model. Specifically, we are interested in estimating the target BP’s function as given by \(f:\mathbb {R}^{\alpha } \rightarrow \mathbb {R}^{\beta }\) based on the data set {(X1,Y1),...,(X N ,Y N )}, where X are explanatory vectors and Y are response vectors. An estimated function \(\hat {f}(\cdot )\) can be built using the given data set. We therefore perform the same process even when the original data set would be changed, using the bagging technique. The fundamental concept of the bagging technique is to use the pseudo data generated from the original data set to obtain multiple estimates as \(\{\hat {f}_{1}(\cdot ),..., \hat {f}_{K}(\cdot )\}\) to address the unstable estimations such as the high variances of mean error or mean square error [8]. Making the ensemble estimators \(\hat {f}_{\varphi }(\cdot ) \) can be given as:
where \(\hat {f}_{k}(\cdot )\) denotes the estimated function acquired from the k th pseudo-data set; and ζ k is a weighted coefficient. In this paper, we can use the bagging technique to increase the reliability of the results and reduce the variance of the mean error in the SVR technique. In addition, the small number of sample problems acquired in each patient and the unstable performance of the SVR model can be solved simultaneously.
The target expectation is given by \(\mathbb {E}[{Y}|{X}]\), considering the proposed SVR model. Suppose that X = {x1,...,x N } and Y = {y1,...,y N } are random samples of the \({\mathbb {F}}\) distribution whose parameters are unknown {x,σ2}, respectively.
Therefore, we use the \(\hat {\mathbb {F}}\) employing sample parameters \((\bar {x}, \sigma ^{2}| {X})\), where the mean and variance are obtained from \(\mathbb {E}(x|{X}) \simeq \bar {x}= \frac {1}{n} {\sum }_{i = 1}^{n} x_{i}\) and \(\mathbb {E}(\sigma ^{2}|{X} ) \simeq \hat \sigma ^{2} = {\frac {1}{n-1} {\sum }_{i = 1}^{n}(x_{i} - \bar {x})^{2} }\), where \(\hat {\mathbb {F}}\simeq \mathcal {N}(\bar {x}, \sigma ^{2})\) is a normally distributed, referred to as the parametric bootstrap [15]. The parametric bootstrap is employed to generate the pseudo-feature vectors acquired from the mean and variance of the original for each subject. An identical process is utilized to generate the pseudo-features from Y = {y1,...,y N } to use the target BPs. Therefore, sufficient samples are prepared to predict the nurse’s BPs (SBP and DBP) as a training data set, and new features that do not participate in the training phase are utilized as test data.
Experimental results
BP monitors can pass standard protocols if the ME is less than 5 mmHg and the SDE is less than 8 mmHg [5]. The MEs of the proposed methodologies were readily computed by \((e_{i} = {t}_{i} - \hat {t}_{i})\). The ME and MAE were given as \((n^{-1} {\sum }_{i = 1 }^{n } e_{i}), (n^{-1} {\sum }_{i = 1 }^{n } \mid e_{i} \mid )\), respectively. The SDE values of ME and MAE were also easily computed using statistical equations.
Statistical analysis
The parameters of the proposed SVR ensemble algorithm are summarized in Table 2. Based on this layout, we performed a test to assess the performance of the SVR fusion technique when using equal conditions with a number of bootstrap replications (B(= 100)) for the pseudo-features. We confirmed that the SVR fusion technique achieved consistently good results. We showed that the SVR fusion estimator produced superior results compared to the FFNN, as demonstrated in Table 3. We evaluated the ME of SBP and DBP acquired employing the SVR fusion estimator with that of MAA[9] and FFNN [3].
Discussion
The SDE values obtained by FFNN were achieved to be 7.61 and 6.79 mmHg, respectively, as presented in Table 3. The SDEs for the proposed SVR fusion model were improved by 2.18 and 1.58 mmHg compared to those of the MAA, as represented in Table 4. We confirmed differences of 0.26 and 0.32 mmHg in the SDE values, respectively, between the SVR fusion algorithm and the SVR single algorithm. Additionally, we compared to conventional FFNN algorithm. According to the BHS protocol, the SVR fusion methodology acquired grades of A and A for the SBP and DBP, respectively. The results of the SVR fusion estimator were 66.82 % (≤ 5 mmHg), 88.00 % (≤ 10 mmHg), and 95.53 % (≤ 15 mmHg) for the SBP in the given experimental test, and 74.82 % (≤ 5 mmHg), 94.12 % (≤ 10 mmHg), and 98.82 % (≤ 15 mmHg) for the DBP in the experimental test, as shown in Table 4. Conclusively, we confirmed that the SVR fusion estimator outperformed the conventional models, as indicated in Tables 3 and 4. This implies that enough pseudo-features are a critical factor for enhancing the generalization ability of the SVR fusion estimator. The results indicated that the estimated blood pressure values obtained by the SVR ensemble estimator were close to the reference blood pressures (SBP and DBP). The probabilities of the BHS standard protocol [17,18,19,20,21] represented that the proposed SVR fusion algorithm offered accurate BP estimates compared with the FFNN estimator as shown Figs. 2 and 3.
The Bland-Altman plots [5]
The Bland-Altman plots [5]
Conclusion
In conclusion, the proposed technique incorporates an SVR fusion estimator and offers a method that can mitigate uncertainty for oscillometric BP measurements using the bootstrap ensemble technique. The results demonstrate that this methodology improves the accuracy of BP estimation. The proposed strategy can be used for various kinds of oscillation phenomena such as a speech, a photoplethysmography, and an electrocardiography signals.
References
Lee, S., Chang, J.-H., Nam, S.W., Lim, C., Rajan, S., Dajani, H., and Groza, V., Oscillometric blood pressure estimation based on maximum amplitude algorithm employing Gaussin mixture regression. IEEE Trans. Instumen. Meas. 62(12):3387–3389, 2013.
Lee, S., Bolic, M., Groza, V., Dajani, H., and Rajan, S., Confidence interval estimation for oscillometric blood pressure measurements using bootstrap approach. IEEE Trans. Instumen. Meas. 60(10):3405–3415, 2011.
Forouzanfar, M., Dajani, H., Groza, V., Bolic, M., and Rajan, S., Feature-based neural network approach for oscillometric blood pressure estimation. IEEE Trans. Instumen. Meas. 60(8):2786–2796, 2011.
Lee, S. et al., Improved Gaussian mixture regression based on pseudo feature generation using bootstrap in blood pressure measurement. IEEE Trans. Ind. Informat. 12(6):2269–2280 , 2016.
Association for the advancement of medical instrumentation (AAMI), American national standard manual, electronic or automated sphygmonanometers. AASI/AAMI SP 10:2002, 2003.
Rakotomamonjy, A., Analysis of SVM regression bound for variable ranking. Neurocomputing 70:1489–1491, 2007.
Theodoridis, S., Machine learning. London: Academic Press, 2015.
Buhlmann, P., and Yu, B., Analyzing bagging. ANN. STAT. 30(4):927–961, 2002.
Lee, S., and Chang, J.-H., Deep belief networks ensemble for blood pressure estimation. IEEE ACCESS 5:9962–9972, 2017.
Lee, S., and Chang, J.-H., Deep learning ensemble with asymptotic techniques based on bootstrap for oscillometric blood pressure estimation. Comput. Methods Prog. Biomed. 151:1–13, 2017.
Sangaiah, A.K., Samuel, O.W., Li, X., Abdel-Basset, M., and Wang, H.: Towards an efficient risk assessment in software projects–Fuzzy reinforcement paradigm. Computers & Electrical Engineering. in press, 2017
Aborokbah, M.M., Al-Mutairi, S., Sangaiah, A.K., and Samuel, O.W.: Adaptive context aware decision computing paradigm for intensive health care delivery in smart cities—A case analysis, sustainable cities and society,in press, 2017
Wu, F., Li, X., Sangaiah, A.K., Xu, L., Kumari, S., Wu, L., and Shen, J.: A lightweight and robust two-factor authentication scheme for personalized healthcare systems using wireless medical sensor networks, Futur. Gener. Comput. Syst. in press, 2017
Ahmad, S., Bolic, M., Dajani, H., Groza, V., Batkin, I., and Rajan, S., Measurement of heart rate variability using an oscillometric blood pressure monitor. IEEE Trans. Instumen. Meas. 59(10):2575–2590, 2010.
Efron, B., and Tibshirani, R., Bootstrap methods for standard errors, confidence interval, and other measures of statistical accuracy. Stat. Sci. 1(1):54–77, 1986.
Lee, S., Rajan, S., Park, C.H., Chang, J.-H., Dajani, H., and Groza, V., Estimated confidence interval from single blood pressure measurement based on algorithm fusion. Comput. Biol. Med. 62:154–163, 2015.
O’Brien, E. et al., European society of hypertension recommendations for conventional, ambulatory and home blood pressure measurement. J. of Hypertension 21(5):821–848, 2003.
Sangaiah, A.K., Thangavelu, A.K., Gao, X.Z., Anbazhagan, N., and Durai, M.S., An ANFIS approach for evaluation of team-level service climate in GSD projects using Taguchi-genetic learning algorithm. Appl. Soft Comput. 30:628–635, 2015.
Medhane, D. V., and Sangaiah, A. K., ESCAPE: Effective Scalable Clustering Approach For Parallel Execution of continuous position-based queries in position monitoring applications. IEEE Transactions on Sustainable Computing 2(2):49–61 , 2017.
Qiu, T., Zhang, Y., Qiao, D., Zhang, X., Wymore, M.L., and Sangaiah, A.K.: A robust time synchronization scheme for industrial internet of things, IEEE Trans. Ind. Inf., to appear
Qiu, T., Qiao, R., Han, M., Sangaiah, A. K., and Lee, I., A Lifetime-Enhanced data collecting scheme for the internet of things. IEEE Communications Magazine 55(11):132–137 , 2017.
Acknowledgments
This work was supported by the NRF Grant funded by the Korean Government 2016R1D1A1B03932925 and 2015R1D1A1A01058171.
Author information
Authors and Affiliations
Corresponding authors
Additional information
This article is part of the Topical Collection on Image & Signal Processing
Rights and permissions
About this article
Cite this article
Lee, S., Ahmad, A. & Jeon, G. Combining Bootstrap Aggregation with Support Vector Regression for Small Blood Pressure Measurement. J Med Syst 42, 63 (2018). https://doi.org/10.1007/s10916-018-0913-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10916-018-0913-x