Abstract
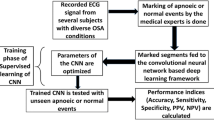
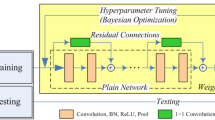
In this study, we propose a method for the automated detection of obstructive sleep apnea (OSA) from a single-lead electrocardiogram (ECG) using a convolutional neural network (CNN). A CNN model was designed with six optimized convolution layers including activation, pooling, and dropout layers. One-dimensional (1D) convolution, rectified linear units (ReLU), and max pooling were applied to the convolution, activation, and pooling layers, respectively. For training and evaluation of the CNN model, a single-lead ECG dataset was collected from 82 subjects with OSA and was divided into training (including data from 63 patients with 34,281 events) and testing (including data from 19 patients with 8571 events) datasets. Using this CNN model, a precision of 0.99%, a recall of 0.99%, and an F1-score of 0.99% were attained with the training dataset; these values were all 0.96% when the CNN was applied to the testing dataset. These results show that the proposed CNN model can be used to detect OSA accurately on the basis of a single-lead ECG. Ultimately, this CNN model may be used as a screening tool for those suspected to suffer from OSA.




Similar content being viewed by others
References
Young, T., Evans, L., Finn, L., and Palta, M., Estimation of the clinically diagnosed proportion of sleep apnea syndrome in middle-aged men and women. Sleep 20(9):705–706, 1997.
Berry, R. B., Brooks, R., Gamaldo, C. E., Harding, S. M., Marcus, C., and Vaughn, B., AASM manual for the scoring of sleep and associated events. Rules, terminology and technical specifications. Darien, IL: AASM, 2012.
Engleman, H. M., and Douglas, N. J., Sleep 4: Sleepiness, cognitive function, and quality of life in obstructive sleep apnoea/hypopnea syndrome. Thorax 59(7):618–622, 2004. https://doi.org/10.1136/thx.2003.015867.
Vgontzas, A. N., Papanicolaou, D. A., Bixler, E. O., Hopper, K., Lotsikas, A., Lin, H.-M., Kales, A., and Chrousos, G. P., Sleep apnea and daytime sleepiness and fatigue: Relation to visceral obesity, insulin resistance, and hypercytokinemia. J. Clin. Endocrinol. Metab. 85(3):1151–1158, 2000.
Barbé, F., Pericas, J., Munoz, A., Findley, L., Anto, J. M., and Agusti, A. G., Automobile accidents in patients with sleep apnea syndrome: An epidemiological and mechanistic study. A J. Res. Crit. Care Med. 158(1):18–22, 1998.
Bhattacharjee, R., Kheirandish-Gozal, L., Pillar, G., and Gozal, D., Cardiovascular complications of obstructive sleep apnea syndrome: Evidence from children. Prog. Cardiovasc. Dis. 51(5):416–433, 2009.
Lopez-Jimenez, F., Sert Kuniyoshi, F. H., Gami, A., and Somers, V. K., Obstructive sleep apnea: Implications for cardiac and vascular disease. Chest 133(3):793–804, 2008. https://doi.org/10.1378/chest.07-0800.
Penzel, T., The apnoea-ECG database. Comput. Cardiol. 27:255–258, 2000.
Penzel, T., McNames, J., de Chazal, P., Raymond, B., Murray, A., and Moody, G., Systematic comparison of different algorithms for apnoea detection based on ECG recordings. Med. Biol. Eng. Comput. 40(4):402–407, 2002.
Yildiz, A., Akın, M., and Poyraz, M., An expert system for automated recognition of patients with obstructive sleep apnea using electrocardiogram recordings. Expert Syst. Appl. 38(10):12880–12890, 2011.
Marcos, J. V., Hornero, R., Alvarez, D., Aboy, M., and Del Campo, F., Automated prediction of the apnea-hypopnea index from nocturnal oximetry recordings. IEEE Trans. Biomed. Eng. 59(1):141–149, 2012. https://doi.org/10.1109/TBME.2011.2167971.
Park, J. U., Lee, H. K., Lee, J., Urtnasan, E., Kim, H., and Lee, K. J., Automatic classification of apnea/hypopnea events through sleep/wake states and severity of SDB from a pulse oximeter. Physiol. Meas. 36(9):2009–2025, 2015.
Koley, B. L., and Dey, D., Automatic detection of sleep apnea and hypopnea events from single channel measurement of respiration signal employing ensemble binary SVM classifiers. Measurement 46(7):2082–2092, 2013.
Solà-Soler, J., Fiz, J. A., Morera, J., and Jané, R., Multiclass classification of subjects with sleep apnoea–hypopnoea syndrome through snoring analysis. Med. Eng. Phys. 34(9):1213–1220, 2012.
Erdenebayar, U., Park, J. U., Jeong, P., and Lee, K. J., Obstructive sleep apnea screening using a piezo-electric sensor. J. Korean Med. Sci. 32(6):893–899, 2017.
Khandoker, A. H., Gubbi, J., and Palaniswami, M., Automated scoring of obstructive sleep apnea and hypopnea events using short-term electrocardiogram recordings. EEE Trans Inf Technol Biomed 13(6):1057–1067, 2009. https://doi.org/10.1109/TITB.2009.2031639.
Álvarez-Estévez, D., and Moret-Bonillo, V., Fuzzy reasoning used to detect apneic events in the sleep apnea-hypopnea syndrome. Expert Syst. Appl. 36(4):7778–7785, 2009. https://doi.org/10.1016/j.eswa.2008.11.043.
Mendez, M. O., Corthout, J., Van Huffel, S., Matteucci, M., Penzel, T., Cerutti, S., and Bianchi, A. M., Automatic screening of obstructive sleep apnea from the ECG based on empirical mode decomposition and wavelet analysis. Physiol. Meas. 31(3):273–289, 2010. https://doi.org/10.1088/0967-3334/31/3/001.
Al-Angari, H. M., and Sahakian, A. V., Automated recognition of obstructive sleep apnea syndrome using support vector machine classifier. IEEE Trans. Inf. Technol. Biomed. 16(3):463–468, 2012. https://doi.org/10.1109/TITB.2012.2185809.
Angermueller, C., Pärnamaa, T., Parts, L., and Stegle, O., Deep learning for computational biology. Mol. Syst. Biol. 12(7):878, 2016.
LeCun, Y., Bengio, Y., and Hinton, G., Deep learning. Nature 521(7553):436–444, 2015. https://doi.org/10.1038/nature14539.
Krizhevsky, A., Sutskever, I., Hinton, G. E., Imagenet classification with deep convolutional neural networks. Adv. Neural Inf. Process Syst. 1:1097–1105, 2012
Hinton, G., Deng, L., Yu, D., Dahl, G. E., Mohamed, A. R., Jaitly, N., and Kingsbury, B., Deep neural networks for acoustic modeling in speech recognition: The shared views of four research groups. IEEE Signal Process. Mag. 29(6):82–97, 2012.
Sutskever, I., Vinyals, O., Le, Q. V., Sequence to sequence learning with neural networks. Adv. Neural Inf. Process Syst. 2:3104–3112, 2014
Ravi, D., Wong, C., Deligianni, F., Berthelot, M., Perez, J. A., Lo, B., and Yang, G. Z., Deep learning for health informatics. IEEE J. Biomed. Health Inform. 21(1):4–21, 2016. https://doi.org/10.1109/JBHI.2016.2636665.
Guo, Y., Liu, Y., Oerlemans, A., Lao, S., Wu, S., and Lew, M. S., Deep learning for visual understanding: A review. Neurocomputing 187:27–48, 2016. https://doi.org/10.1016/j.neucom.2015.09.116.
Kiranyaz, S., Ince, T., and Gabbouj, M., Real-time patient-specific ECG classification by 1-D convolutional neural networks. IEEE Trans. Biomed. Eng. 63(3):664–675, 2016.
Ioffe, S., Szegedy, C., Batch normalization: accelerating deep network training by reducing internal covariate shift. International Conference on Machine Learning 448–456, 2015.
Rosasco, L., Vito, E. D., Caponnetto, A., Piana, M., and Verri, A., Are loss functions all the same? Neural Comput. 16(5):1063–1076, 2004.
Srivastava, N., Hinton, G., Krizhevsky, A., Sutskever, I., and Salakhutdinov, R., Dropout: A simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 15:1929–1958, 2014.
Yu, X., Efe, M. O., and Kaynak, O., A general backpropagation algorithm for feedforward neural networks learning. IEEE Trans. Neural Netw. 13(1):251–254, 2002.
Chollet, F, Keras, 2015. http://keras.io/.
Abadi, M., Barham, P., Chen, J., Chen, Z., Davis, A., Dean, J., and Kudlur, M., TensorFlow: A system for large-scale machine learning. OSDI 16:265–283, 2016.
Kingma, D. P., Ba, J., Adam: A method for stochastic optimization. arXiv preprint arXiv:1412.6980, 2014.
Chien, C., Batch size selection for the batch means method. Proceedings of the 26th Conference on Winter Simulation, Society for Computer Simulation International, 345–352, 1994.
Xie, B., and Minn, H., Real-time sleep apnea detection by classifier combination. IEEE Trans. Inf. Technol. Biomed. 16(3):469–477, 2012. https://doi.org/10.1109/TITB.2012.2188299.
Jafari, A., Sleep apnoea detection from ECG using features extracted from reconstructed phase space and frequency domain. Biomed Signal Process Control 8(6):551–558, 2013.
Chen, L., Zhang, X., and Song, C., An automatic screening approach for obstructive sleep apnea diagnosis based on single-lead electrocardiogram. IEEE Trans. Autom. Sci. Eng. 2:106–115, 2015.
Acknowledgments
This work was supported by a National Research Foundation of Korea (NRF) grant funded by the Ministry of Education. It is a product of the Local Innovative Creative Human Resource Training Project (NRF-2014H1C1A1063845).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declares that he or she has no conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
This article is part of the Topical Collection on Mobile & Wireless Health
Rights and permissions
About this article
Cite this article
Urtnasan, E., Park, JU., Joo, EY. et al. Automated Detection of Obstructive Sleep Apnea Events from a Single-Lead Electrocardiogram Using a Convolutional Neural Network. J Med Syst 42, 104 (2018). https://doi.org/10.1007/s10916-018-0963-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10916-018-0963-0