Abstract
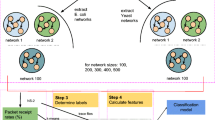
Evolved biological network topologies may resist perturbances to allow for more robust information transport across larger networks in which their network motifs may play a complex role. Although the abundance of individual motifs correlate with some metrics of biological robustness, the extent to which redundant regulatory interactions affect motif connectivity and how this connectivity affects robustness is unknown. To address this problem, we applied machine learning based regression modeling to evaluate how feed-forward loops interlinked by crosstalk altered information transport across a network in terms of packets successfully routed over networks of noisy channels via NS-2 simulation. The sample networks were extracted from the complete transcriptional regulatory networks of two well-studied bacteria: E.coli and Yeast. We developed 233 topological features for the E.coli subnetworks and 842 topological features for the Yeast subnetworks which distinctly account for the opportunities in which two feed-forward loops may exhibit crosstalk. Random forest regression modeling was used to infer significant features from this modest configuration space. The coefficient of determination was used as a primary performance metric to rank features within our regression models. Although only a handful of features were highly ranked, we observed that, in particular, feed-forward loop crosstalk patterns correlated substantially with an improved chance for successful information transmission. Additionally, both E.coli and Yeast subnetworks demonstrate very similar FFL crosstalk patterns that were considered significant in their contribution to information transport robustness in these two organisms. This finding may potentially allude to common design principles in transcriptomic networks from different organisms.










Similar content being viewed by others
Change history
27 November 2017
The original version of this article unfortunately contained a mistake in the author group section. Author Preetam Ghosh’s given name was incorrectly spelled as “Preeetam”.
References
Mayo M, Abdelzaher AF, Perkins E, Ghosh P (2014) Top-level dynamics and the regulated gene response of feed-forward loop transcriptional motifs. Phys Rev E 90(3):032706
Milo R, Shen-Orr S, Itzkovitz S, Kashtan N, Chklovskii D, Alon U (2002) Network motifs: simple building blocks of complex networks. Science 298(5594):824–827
Mangan S, Alon U (2003) Structure and function of the feed-forward loop network motif. Proc Nat Acad Sci 100(21):11980–11985
Kamapantula BK, Mayo M, Perkins E, Ghosh P (2014) Dynamical impacts from structural redundancy of transcriptional motifs in gene-regulatory networks. In: Proceedings of the 8th international conference on bioinspired information and communications technologies (BICT ’14). ICST (Institute for Computer Sciences, Social-Informatics and Telecommunications Engineering), ICST, Brussels, pp 199–206. https://doi.org/10.4108/icst.bict.2014.257928
Kamapantula BK, Mayo M, Perkins E, Abdelzaher AF, Ghosh P (2014) Feature ranking in transcriptional networks: packet receipt as a dynamical metric. In: Proceedings of the 8th international conference on bioinspired information and communications technologies (BICT ’14). . ICST (Institute for Computer Sciences, Social-Informatics and Telecommunications Engineering), ICST, Brussels, pp 1–8. https://doi.org/10.4108/icst.bict.2014.257930
Guo S, Murray RM Prototyping and implementation of a novel feedforward loop in a cell-free transcription-translation system and cells. https://doi.org/10.1101/123190
Barabasi AL, Albert R (1999) Emergence of scaling in random networks. Science 286:509–512
Ghosh S, Ghosh P, Basu K, Das SK (2005) GaMa: an evolutionary algorithmic approach for the design of mesh-based radio access networks. In: Proceedings of the IEEE conference on local computer networks 30th anniversary (LCN ’05). IEEE Computer Society, Washington, pp 374–381. https://doi.org/10.1109/LCN.2005.72
Mayo M, Abdelzaher AF, Perkins EJ, Ghosh P (2012) Motif Participation by Genes in E. coli transcriptional networks. Front Physiol 3:357. https://doi.org/10.3389/fphys.2012.00357
Kamapantula BK, Abdelzaher A, Ghosh P, Mayo M, Perkins E, Das SK (2012) Performance of wireless sensor topologies inspired by E. coli genetic networks. In: 2012 IEEE international conference on pervasive computing and communications workshops (PERCOM Workshops). IEEE, pp 302–307. https://doi.org/10.1109/PerComW.2012.6197500
Ghosh P, Mayo M, Chaitankar V, Habib T, Perkins E, Das SK (2011) Principles of genomi crobustness inspire fault-tolerant wsn topologies: a network science based case study. In: 2011 IEEE international conference on pervasive computing and communications workshops (PERCOM Workshops). IEEE, pp 160–165. https://doi.org/10.1109/PERCOMW.2011.5766861
Kamapantula BK, Abdelzaher A, Ghosh P et al (2014) Leveraging the robustness of genetic networks: a case study on bio-inspired wireless sensor network topologies. J Ambient Intell Human Comput 5:323. https://doi.org/10.1007/s12652-013-0180-0
Nazi A, Raj M, Di Francesco M, Ghosh P, Das SK (2016) Efficient communications in wireless sensor networks based on biological robustness. In: Proceedings of the 2016 international conference on distributed computing in sensor systems (DCOSS), pp 161–168
Nazi A, Raj M, Di Francesco M, Ghosh P, Das SK (2015) Exploiting gene regulatory networks for robust wireless sensor networking. In: Proceedings of the 2015 IEEE global communications conference (GLOBECOM), pp 1–7. https://doi.org/10.1109/GLOCOM.2015.7416957
Nazi A, Raj M, Di Francesco M, Ghosh P, Das SK (2013) Robust deployment of wireless sensor networks using gene regulatory networks. In: Proceedings of the 2013 international conference on distributed computing and networking, pp 192–207
Nazi A, Raj M, Di Francesco M, Ghosh P, Das SK (2014) Deployment of robust wireless sensor networks using gene regulatory networks: an isomorphism-based approach. Pervas Mob Comput 13:246–257
Chan H, Akoglu L, Tong H (2014) Make it or break it: manipulating robustness in large networks. In: Proceedings of the 2014 SIAM data mining conference. SIAM, pp 325–333
de la Peña JA, Gutman I, Rada J (2007) Estimating the Estrada index. Linear Algebra Appl 427:70–76
Kamapantula BK, Abdelzaher AF, Mayo M, Perkins E, Das SK, Ghosh P (2017) Quantifying robustness in biological networks using NS-2. Model Methodol Tools Molecular Nano-scale Commun 9:273–290
Schaffter T, Marbach D, Floreano D (2011) GeneNetWeaver: in silico benchmark generation and performance profiling of network inference methods. Bioinformatics 27:2263–2270
Rowland MA, Abdelzaher AF, Ghosh P, Mayo ML (2017) Crosstalk and the dynamical modularity of feed-forward loops in transcriptional regulatory networks. Biophys J 112:1539–1550
Pedregosa F, Varoquaux G, Gramfort A, Michel V, Thirion B, Grisel O, Blondel M, Prettenhofer P, Weiss R, Dubourg V et al (2011) Scikit-learn: machine learning in Python. J Mach Learn Res 12:2825–2830
Schult DA, Swart PJ (2008) Exploring network structure, dynamics, and function using NetworkX. Proc 7th Python Sci Conf (SciPy 2008(2008):11–16
Breiman L (2001) Random forests. Mach Learn 45:5–32
Acknowledgments
Funding was provided by the US Army’s Environmental Quality and Installations 6.1 Basic Research program. Opinions, interpretations, conclusions, and recommendations are those of the author(s) and are not necessarily endorsed by the US Army.
Author information
Authors and Affiliations
Corresponding author
Additional information
The original version of this article was revised: There was a mistake in the author group section. Author Preetam Ghosh’s given name was incorrectly spelled as “Preeetam”.
Rights and permissions
About this article
Cite this article
Syed, K., Abdelzaher, A., Mayo, M. et al. Similar Feed-forward Loop Crosstalk Patterns may Impact Robust Information Transport Across E. coli and S. Cerevisiae Transcriptional Networks. Mobile Netw Appl 25, 1970–1982 (2020). https://doi.org/10.1007/s11036-017-0944-4
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11036-017-0944-4