Abstract
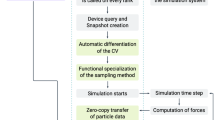
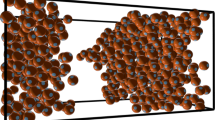
By their structure and operation, biomolecules have resolved fundamental problems as a distributed computational system that we are just beginning to unveil. One advantageous approach to gain a good understanding of the processes and algorithms involved is simulation on conventional computers. Simulations allow better understanding of the capabilities of molecules because they can occur at the level of reliability, efficiency, and programmability that are standard in conventional computation and are desirable for experiments in vitro. Here, we describe in some detail the architecture of a general-purpose simulation environment in silico, EdnaCo, establish its soundness and reliability, and benchmark its performance. The system can be described as an emulation of the events in a real test tube. We describe the major pieces of its architecture, namely, a distributed memory (file) system, a kinetic engine, and input/output mechanisms. Finally, the ability of this environment in preserving major features of the wet counterpart in vitro is evaluated via an implementation on a cluster of PCs. The results of several simulations are summarized that establish the soundness, utility, applicability, and cost efficiency of the software to facilitate experimentation in vitro.
Similar content being viewed by others
References
LM Adleman (1994) ArticleTitleMolecular computation of solutions to combinatorial problems Science 266 1021–1024 Occurrence Handle1:CAS:528:DyaK2MXitFSjs7k%3D Occurrence Handle7973651
E Baum (1995) ArticleTitleBuilding an associative memory vastly larger than the brain Science 268 583–585
Blain D, Garzon M, Shin SY, Zhang BT, Kashiwamura S, Yamamoto M, Kameda K and Ohuchi A (2004) Development, evaluation and benchmarking of simulation software for biomolecular computing. This issue
Braich RS, Johnson C, Rothemund PWK, Hwang D, Chelyapov N and Adleman LA (2000) Solution to a satisfiability problem on a gel-based DNA computer. In: Lecture Notes in Computer Science LNCS, Vol. 2054, pp. 27–42, Springer-Verlag
C Cantor P Schimmel (1980) Biophysical Chemistry, Part III: The Behavior of Biological Macromolecules Freeman New York
Chen J and Reif J eds (2003) Proceedings of 9th international workshop on DNA-based computers DNA 2003 (Revised Papers). In: Lecture Notes in Computer Science LNCS, Vol. 2943, Springer-Verlag
Chen J, Deaton R, Garzon M, Kim JW, Wood D, Bi H, Carpenter D and Wang YZ (2004) Characterization of non-crosshybridizing DNA oligonucleotides manufactured in vitro. In: (Ferreti et al., 2003), 132–141
Deaton R, Chen J, Bi H, Garzon M, Rubin H and Wood D, (2002) A PCR-based protocol for in vitro selection of non-crosshybridizing oligonucleotides. In: (Chen and Reif, 2003), 105–114
L Deng (2004) Generalized mersenne prime numbers and its application to random number generation H Niederreiter (Eds) Monte Carlo and Quasi-Monte Carlo Methods. Springer-Verlag ␣ 167–180
A Einstein (1956) Investigations on the theory of the Brownian movement Reprint, Dover Publications Inc New York 49
Ferreti C, Mauri G and Zandron C eds (2004) Proceedings 10th international workshop on DNA-based computers DNA 2004. In: Lecture Notes in Computer Science LNCS, Springer-Verlag, in press
Fogel G, and Garry Greenwood et al. (eds) (2004) Proceedings of the IEEE Conference on Evolutionary Computation CEC04. Computer Society Press
Garzon M and Oehmen C (2001) Biomolecular computation in virtual test tubes. Proceedings of 7th International workshop on DNA-based computers DNA 2002 (Revised Papers). In: Lecture Notes in Computer Science LNCS, Vol. 2340, pp. 117–128, Springer-Verlag
M Garzon (1995) Models of Massive Parallelism (Analysis of Cellular Automata and Neural Networks) Springer-Verlag Berlin
Garzon M, Bobba K and Hyde B (2004) Digital information encoding on DNA. In: Lecture Notes in Computer Science, Vol. 2590, pp. 152–166, Springer-Verlag
Garzon M, Blain D, Bobba K, Neel A and West M (2003a) Self-Assembly of DNA-like structures In-Silico. In (Garzon, 2003), 185–200
Garzon M, Drumwright E, Deaton R and Renault D (2000) Virtual test tubes: a new methodology for computing. In Proceedings of 7th International Symposium on String Processing and Information Retrieval. pp. 116–121, A Coruña, Spain. IEEE Computer Society Pzzress
Garzon M, Deaton R, Rose J, and Franceschetti D (1999) Soft molecular computing. In: Proceedings of the 5th workshop, MIT, Vol. 54, pp. 89–98, DIMACS Series American Mathematical Society
M Garzon (Eds) (2003) Biomolecular Machines and Artificial Evolution Genetic Programming and Evolvable Machines Kluwer Academic Publishers
Garzon M, Neathery P, Deaton R, Murphy R, Franceschetti D and Stevens S Jr. (1997) A new metric for DNA computing. In: (Koza et al., 1997), 472–478
Garzon M, Neel A and Bobba K (2003b) Efficiency and reliability of semantic retrieval in DNA-based memories. In: (Chen and Reif, 2003), 157–169
E Gentle (2003) Random Number Generation and Monte Carlo Methods Springer-Verlag New York 51
Hagiya M and Ohuchi A eds (2002) Proceedings of 8th international workshop on DNA-based computers DNA 2002 (Revised Papers). In: Lecture Notes in Computer Science LNCS, Vol. 2568, Springer-Verlag.
S Ji (1998) ArticleTitleThe cell as the smallest DNA-based molecular computer Biosystems 52 123–133
Jonoska N, sa-Ardyen P and Seeman AN (1997) Computation by self-assembly of DNA graphs. In (Garzon, 2003), 123–137
Kari L, Winfree E and Gifford D eds (1999) Proceedings of 5th workshop on DNA Computers, MIT, Cambridge MA, 1999, Vol. 54, pp. 247–258. DIMACS series of the American Mathematical Society
Koza J, Deb K, Dorigo M, Fogel D, Garzon M, Iba H and Riolo R eds (1997). Proceedings of 2nd Annual Genetic Programming Conference. Morgan Kaufmann, San Mateo, California
SantaLucia J Jr (1998) A unified view of polymer, dumbbell, and oligonucleotideDNA nearest-neighbor thermodynamics. Proceedings of the National Academic Science 95(1998), 1460
T Schlick (2002) Molecular Modeling and Simulation Springer-Verlag New York 33
T Toffoli N Margolus (1987) Cellular Automata Machines. In: A New Environment for Modeling MIT Press (1987) Cambridge Massachusetts
West M and Garzon MH (2003) Self-Aseembly of DNA Structures In Silico for 3-Colorability. Poster at the 10th Int. Workshop on DNA-Based Computers. Milan, Italy, 2004
West M, Garzon M, and Blain D (2003) DNA-like Genomes for Evolution in silico. In: Proceedings of GECCO-2003, The Genetic and Evolutionary Programming Conference Springer-Verlag Lecture Notes in Computer Science Vol. 2723, pp. 413–424
Wetmur J (1997) Preliminary Proceedings of the Third Annual Meeting on DNA Based Computers, University of Pennsylvania, DIMACS Series American Mathematical Society 48, Providence, RI
W Winfree F Liu LA Wenzler NC Seeman (1998) ArticleTitleDesign and self-assembly of two-dimensional DNA crystals Nature 394 539–544 Occurrence Handle10.1038/28998 Occurrence Handle1:CAS:528:DyaK1cXltlyitrg%3D Occurrence Handle9707114
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Garzon, M.H., Blain, D.R. & Neel, A.J. Virtual test tubes. Nat Comput 3, 461–477 (2004). https://doi.org/10.1007/s11047-004-2642-y
Issue Date:
DOI: https://doi.org/10.1007/s11047-004-2642-y