Abstract
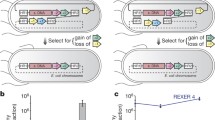
The emerging field of synthetic biology moves beyond conventional genetic manipulation to construct novel life forms which do not originate in nature. We explore the problem of designing the provably shortest genomic sequence to encode a given set of genes by exploiting alternate reading frames. We present an algorithm for designing the shortest DNA sequence simultaneously encoding two given amino acid sequences. We show that the coding sequence of naturally occurring pairs of overlapping genes approach maximum compression. We also investigate the impact of alternate coding matrices on overlapping sequence design. Finally, we discuss an interesting application for overlapping gene design, namely the interleaving of an antibiotic resistance gene into a target gene inserted into a virus or plasmid for amplification.



Similar content being viewed by others
References
Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2002) Molecular biology of the cell. Garland Science, New York
Ball P (2004) Starting from scratch. Nature 431:624–626
Cann AJ (1993) Principles of molecular virology. Academic Press, London
Cello J, Paul AV, Wimmer E (2002) Chemical synthesis of poliovirus cDNA: generation of infectious virus in the absence of natural template. Science 297:1016–1018
Cohen B, Skiena S (2003) Natural selection and algorithmic design of mrna. J Comput Biol 10:419–432
Daley M, McQuillan I (2005a) Formal modelling of viral gene compression. Int J Found Comput Sci 16(3):453–469
Daley M, McQuillan I (2005b) Viral gene compression: complexity and verification. Lect Notes Comput Sci 3317:102–112
Elber R, Karplus M (1990) Enhanced sampling in molecular dynamics: use of the time-dependent hartree approximation for a simulation of carbon monoxide diffusion through myoglobin. J Am Chem Soc 112:9161–9175
Freeland S, Hurst L (2004) Evolution encoded. Sci Am 290(4):84–91
Fukuda Y, Washio T, Tomita M (1998) Evolution of overlapping genes: Comparative genomics of mycoplasma genitalium and mycoplasma pneumoniae. Genome Inform 9:254–255
Fukuda Y, Nakayama Y, Tomita M (2003) On dynamics of overlapping genes in bacterial genomes. Gene 323:181–187
Gilis D, Massar D, Cerf NJ, Rooman M (2001) Optimality of the genetic code with respect to protein stability and amino-acid frequencies. Genome Biol 2(11):1–12
Hornak V, Simmerling C (2003) Generation of accurate protein loop conformations through low-barrier molecular dynamics. Proteins 51:577–590
Karlin S, Chen C, Gentles A, Cleary M (2002) Associations between human disease genes and overlapping gene groups and multiple amino acid runs. Proc Natl Acad Sci 99(26):17008–17013
Keese P, Gibbs A (1992) Origins of genes: “big bang” or continuous creation? Proc Natl Acad Sci 89:9489–9493
Kodumal S, Pael K, Reid R, Menzella H, Welch M, Santi D (2004) Total synthesis of long DNA sequences: synthesis of a contiguous 32-kb polyketide synthase gene cluster. Proc Natl Acad Sci 44:15573–15578
Krakauer DC (2000) Stability and evolution of overlapping genes. Evolution 54(3):731–739
Krakauer D (2002) Evolutionary principles of genomic compression. Comments Theor Biol 7:215–236
Levitt M (1976) A simplified representation of protein conformations for rapid simulation of protein folding. J Mol Biol 104:59–107
Marti-Renom MA, Stuart AC, Fiser A, Sanchez R, Melo F, Sali A (2000) Comparative protein structure modeling of genes and genomes. Annu Rev Biophys Biomol Struct 29:291–325
Miyata T, Yasunaga T (1978) Evolution of overlapping genes. Nature 272:532–535
Oppenheim D, Yahofsky C (1980) Translational coupling during expression of the tryptophan operon of E. coli. Genetics 95:785–795
Rogozin I, Spiridonov A, Sorokin A, Wolf Y, King J, Tatusov R, Koonin E (2002) Purifying and directional selection in overlapping prokaryotic genes. Trends Genet 18(5):228–232
Skiena S (2001) Designing better phages. Bioinformatics 17:253–261
Skiena S, Wimmer E (2003) Gene design for vaccines and theraputic phages. NSF ITR Award 0325123
Smith H, Hutchison C, Pfannkoch C, Venter JC (2003) Generating a synthetic genome by whole genome assembly: phix174 bacteriophage from synthetic oligonucleotides. Proc Natl Acad Sci 100:15440–15445
Tian J, Gong H, Sheng N, Zhou Z, Gulari E, Gao X, Church G (2004) Accurate multiplex gene synthesis from programmable DNA microchips. Nature 432:1050–1054
Veeramachaneni V, Makalowski W, Galdzicki M, Sood R, Makalowska I (2004) Mammalian overlapping genes: the comparative method. Genome Res 14:280–286
Acknowledgments
This research was partially supported by NSF grants EIA-0325123 and DBI-0444815. We thank Eckard Wimmer for his interest and support. We also thank Chen Zhao, Huei-Chi Chen, and Rahul Sinha for discussions and contributions to this research.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, B., Papamichail, D., Mueller, S. et al. Two proteins for the price of one: the design of maximally compressed coding sequences. Nat Comput 6, 359–370 (2007). https://doi.org/10.1007/s11047-006-9031-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11047-006-9031-7