Abstract
Objective
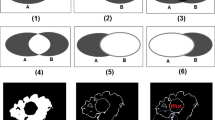
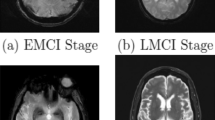
Statistical models for medical images have been developed to increase robustness in the segmentation process. In this project, a fully automatic approach to build a statistical shape-intensity model and combine this model with level set segmentation was designed, implemented and tested by applying the algorithm to clinical image data.
Methods
By using a hierarchical registration approach based on mutual information and demons registration, 3D statistical shape-intensity models were created by applying Principal Component Analysis. Using these models in combination with level set segmentation results in a fully automatic modeling and segmentation pipeline.
Results
Examples for shape-intensity models were synthesized and these models were used to automatically segment 3D MRI and CT images. Quantitative evaluation of the framework was performed by comparing automatic segmentation results to segmentation results of medical experts.
Conclusion
Evaluation tests in which this method was used for the automatic segmentation of femora and cardiac MRI endocardial surfaces are very promising. The implementation of an additional cost function term and the addition of information about the surroundings of an organ in the model are currently under development.
Similar content being viewed by others
References
Kass M, Witkin A, Terzopoulos D (1987) Snakes: active contour models. Int J Comput Vis 1:321–331
Cootes TF, Hill A, Taylor CJ, Haslam J (1994) The use of active shape models for locating structures in medical images. Image Vis Comput 12:355–366
Caselles V, Kimmel R, Sapiro G (1995) Geodesic active contours. International journal of Computer Vision 22:61–79
Zeng X, Staib LH, Schultz RT, Duncan JS (1999) Segmentation and measurement of the cortex from 3-DMRimages using coupled surfaces propagation. IEEE Trans Med Imaging 18:927–937
Leventon M, Grimson E, Faugeras O (2000) Statistical shape influence in geodesic active contours. Comput Vis Patt Recog 1:316–323
Fritscher KD, Schubert R (2006) 3D image segmentation by using statistical deformation models and level sets. Int J Comput Assist Radiol Surg 1:123–135
Chen T, Metaxas D (2000) Image segmentation based on the integration of markov random fields and deformable models. Lecture Notes in Computer Science 1935:256–265
Cremers D, Schnorr C, Weickert J (2001) Diffusion in snakes: combining statistical shape knowledge and image information in a variational framework. IEEE Workshop on Variational Level Set Methods, pp 137–144
Rousson M, Derriche R (2002) A variational framework for active and adaptive segmentation of vector valued images. Presented at IEEE Workshop on Motion and Video Computing, Orlando, Florida
Cootes TF, Edwards GJ, Taylor CJ (2001) Active appearance models. IEEE Trans Patt Anal Mach Intell 6:681–685
Yang J, Duncan JS (2003) 3D image segmentation of deformable objects with shape-appearance joint prior models. Lecture Notes in Comput Science 2878:573–580
Paragios N, Deriche R (2002) Geodesic active regions and level set methods for supervised texture segmentation. Int J Comput Vis 46:223–247
Zhu SC, Yuille A (1996) Region competition: unifying snakes, region growing, and Bayes/MDL for multiband image segmentation. IEEE Trans Patt Anal Mach Intell 18:884–900
Huang X, Li Z, Metaxas D (2004) Learning coupled prior shape and appearance models for segmentation. Lecture Noets in Comput Science 3216:60–69
Huang X, Metaxas D, Chen T (2004) Metamorphs: deformable shape and texture models. In: Presented at IEEE Computer Vision and Pattern Recognition
Tsai A, Yezzi A, Wells W, Tempany C, Tucker D, Fan A, Grimson WE, Willsky A (2003) A shape-based approach to the segmentation of medical imagery using level sets. IEEE Trans Med Imaging 22:137–154
Tsai A, Wells W, Tempany C, Grimson E, Willsky A (2004) Mutual information in coupled multi-shape model for medical image segmentation. Med Image Anal 8:429–445
Wells W, Viola P, Atsumi H, Nakajima S, Kikinis R (1996) Multi-modal volume registration by maximization of mutual information. Med Image Abal 1:35–51
Parzen E (1962) On estimation of probability density function and mode. Ann Math Stat 33:1065–1076
Thirion JP (1998) Image matching as a diffusion process: an analogy with Maxwell’s demons. Med Image Anal 2:243–260
Rueckert D, Frangi AF, Schnabel JA (2001) Automatic construction of 3-D statistical deformation models of the brain using nonrigid registration. IEEE Trans Med Imaging 22:1014–1025
Mattes D, Haynor Dr, Veselle H, Lewellen TK, Eubank W (2003) PET-CT image registration in the chest using free-form deformations. IEEE Trans Med Imaging 22:120–128
Styner M, Gerig G, Brechbuehler Ch, Szekely G (2000) Parametric estimate of intensity inhomogeneities applied to MRI. IEEE Trans Med Imaging 19:153–165
Zijdenbos AP, Dawant BM, Margolin RA, Palmer AC (1994) Morphometric analysis of white matter lesions in MR images: method and validation. IEEE Trans Med Imaging 13:716–724
Cattell RB (1966) The scree test for the number of factors. Multivar Behav Res 1:245–276
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Fritscher, K.D., Grünerbl, A. & Schubert, R. 3D image segmentation using combined shape-intensity prior models. Int J CARS 1, 341–350 (2007). https://doi.org/10.1007/s11548-007-0070-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11548-007-0070-z