Abstract
Purpose
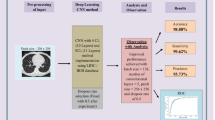
Lung cancer is the most frequent cancer worldwide and is the leading cause of cancer-related deaths. Its early detection and treatment at the stage of a lung nodule improve the prognosis. In this study was proposed a new classification approach named bilinear convolutional neural network (BCNN) for the classification of lung nodules on CT images.
Methods
Convolutional neural network (CNN) is considered as the leading model in deep learning and is highly recommended for the design of computer-aided diagnosis systems thanks to its promising results on medical image analysis. The proposed BCNN scheme consists of two-stream CNNs (VGG16 and VGG19) as feature extractors followed by a support vector machine (SVM) classifier for false positive reduction. Series of experiments are performed by introducing the bilinear vector features extracted from three BCNN combinations into various types of SVMs that we adopted instead of the original softmax to determine the most suitable classifier for our study.
Results
The method performance was evaluated on 3186 images from the public LUNA16 database. We found that the BCNN [VGG16, VGG19] combination with and without SVM surpassed the [VGG16]2 and [VGG19]2 architectures, achieved an accuracy rate of 91.99% against 91.84% and 90.58%, respectively, and an area under the curve (AUC) rate of 95.9% against 94.8% and 94%, respectively.
Conclusion
The proposed method improved the outcomes of conventional CNN-based architectures and showed promising and satisfying results, compared to other works, with an affordable complexity. We believe that the proposed BCNN can be used as an assessment tool for radiologists to make a precise analysis of lung nodules and an early diagnosis of lung cancers.






Similar content being viewed by others
References
Cancer Facts and Figures (2020) Atlanta: American Cancer Society. https://www.cancer.org/research/cancer-facts-statistics/all-cancer-facts-figures/cancer-facts-figures-2020.html. Accessed 06 June 2020
Lung Cancer Fact Sheet (2020). American Lung Association. https://www.lung.org/lung-health-and-diseases/lung-disease-lookup/lung-cancer/learn-about-lung-cancer/lung-cancer-factsheet.html. Accessed 06 June 2020
Mastouri R, Khlifa N, Neji H, Hantous-Zannad S (2020) Deep learning-based CAD schemes for the detection and classification of lung nodules from CT images: a survey. J X Ray Sci Technol 28(4):591–617. https://doi.org/10.3233/XST-200660
Ben Jabra M, Guetari R, Chetouani A, Tabia H, Khlifa N (2020) Facial expression recognition using the bilinear pooling. In: 15th international joint conference on computer vision, imaging and computer graphics theory and applications (VISIGRAPP 2020), pp 294–301. https://doi.org/10.5220/0008928002940301
Monkam P, Qi S, Xu M, Han F, Zhao X, Qian W (2018) CNN models discriminating between pulmonary micro-nodules and non-nodules from CT images. BioMed Eng OnLine 17(1):96. https://doi.org/10.1186/s12938-018-0529-x
Monkam P, Qi S, Xu M, Li H, Han F, Teng Y, Qian W (2019) Ensemble learning of multiple-view 3D-CNNs model for micro-nodules identification in CT images. IEEE Access 7:5564–5576
Shen W, Zhou M, Yang F, Yu D, Dong D, Yang C, Zang Y, Tian J (2017) Multi-crop convolutional neural networks for lung nodule malignancy suspiciousness classification. Pattern Recogn 61:663–673
Onishi Y, Teramoto A, Tsujimoto M, Tsukamoto T, Saito K, Toyama H, Imaizumi K, Fujita H (2019) Automated pulmonary nodule classification in computed tomography images using a deep convolutional neural network trained by generative adversarial networks. Biomed Res Int 6:1–9. https://doi.org/10.1155/2019/6051939
Wu P, Sun X, Zhao Z, Wang H, Pan S, Schuller B (2020) Classification of lung nodules based on deep residual networks and migration learning. Comput Intell Neurosci. https://doi.org/10.1155/2020/8975078
Kaya A (2018) Cascaded classifiers and stacking methods for classification of pulmonary nodule characteristics. Comput Methods Programs Biomed 166:77–89
Kido S, Hirano Y, Hashimoto N (2018) Detection and classification of lung abnormalities by use of convolutional neural network (CNN) and regions with CNN features (R-CNN). Int Workshop Adv Image Technol. https://doi.org/10.1109/iwait.2018.8369798
Liu J, Yang Z, Zhang T, Xiong H (2017) Multi-part compact bilinear CNN for person re-identification. In: 2017 IEEE international conference on image processing (ICIP)
Ustinova E, Ganin Y, Lempitsky V (2015) Multiregion bilinear convolutional neural networks for person re-identification. arXiv preprint arXiv:1512.05300
Chen H, Wang J, Qi Q, Li Y, Sun H (2017) Bilinear CNN models for food recognition. In: 2017 International conference on digital image computing: techniques and applications (DICTA)
Wang C, Shi J, Zhang Q, Yin S (2017) Histopathological image classification with bilinear convolutional neural networks. In: 39th Annual international conference of the IEEE engineering in medicine and biology society (EMBC)
LUng Nodule Analysis (2016). Grand challenge. https://luna16.grand-challenge.org/Data/. Accessed 26 Feb 2019
Wang W, Luo J, Yang X, Lin H (2015) Data analysis of the lung imaging database consortium and image database resource initiative. Acad Radiol 22(4):488–495. https://doi.org/10.1016/j.acra.2014.12.004
Lin TY, RoyChowdhury A, Maji S (2015) Bilinear cnn models for fine-grained visual recognition. In: Proceedings of the IEEE international conference on computer vision, pp 1449–1457
Simonyan K; Zisserman A (2015) Very deep convolutional networks for large-scale image recognition. In: International conference on learning representations (ICLR). arXiv:1409.1556
Shi Z, Hao H, Zhao M, Feng Y, He L, Wang Y, Suzuki K (2018) A deep CNN based transfer learning method for false positive reduction. Multimed Tools Appl. https://doi.org/10.1007/s11042-018-6082-6
Zhao X, Liu L, Qi S, Teng Y, Li J, Qian W (2018) Agile convolutional neural network for pulmonary nodule classification using CT images. Int J Comput Assist Radiol Surg 13(4):585–595. https://doi.org/10.1007/s11548-017-1696-0
Zhao X, Qi S, Zhang B, Ma H, Qian H, Yao Y, Sun J (2019) Deep CNN models for pulmonary nodule classification: model modification, model integration, and transfer learning. J X-Ray Sci Technol. https://doi.org/10.3233/xst-180490
Shen S, Han SX, Aberle DR, Bui AA, Hsu W (2019) An interpretable deep hierarchical semantic convolutional neural network for lung nodule malignancy classification. Expert Syst Appl 128:84–95. https://doi.org/10.1016/j.eswa.2019.01.048
López-Sánchez D, Arrieta AG, Corchado JM (2020) Compact bilinear pooling via kernelized random projection for fine-grained image categorization on low computational power devices. Neurocomputing 398:411–421. https://doi.org/10.1016/j.neucom.2019.05.104
Gao Y, Beijbom O, Zhang N, Darrell T (2016) Compact bilinear pooling. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp 317–326
Sun Q, Wang Q, Zhang J, Li P (2018) Hyperlayer bilinear pooling with application to fine-grained categorization and image retrieval. Neurocomputing 282:174–183
Moussa O, Khachnaoui H, Guetari R, Khlifa N (2019) Thyroid nodules classification and diagnosis in ultrasound images using fine-tuning deep convolutional neural network. Int J Imaging Syst Technol 30(1):185–195. https://doi.org/10.1002/ima.22363
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants performed by any of the authors.
Informed consent
This article does not contain patient data.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Mastouri, R., Khlifa, N., Neji, H. et al. A bilinear convolutional neural network for lung nodules classification on CT images. Int J CARS 16, 91–101 (2021). https://doi.org/10.1007/s11548-020-02283-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11548-020-02283-z