Abstract
Purpose
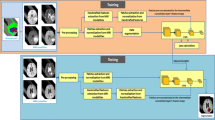
Fully convolutional neural networks (FCNNs) have achieved good performance in the field of medical image segmentation. FCNNs that use multimodal images and multi-scale feature extraction have higher accuracy for brain tumor segmentation. Therefore, we have made some improvements to U-Net for fully automated segmentation of gliomas using multimodal images. And we named it multi-scale dilate network with deep supervision (MSD-Net).
Methods
MSD-Net is a symmetrical structure composed of a down-sampling process and an up-sampling process. In the down-sampling process, we use the multi-scale feature extraction block (ME) to extract multi-scale features and focus on primary features. Unlike other methods, ME consists of dilate convolution and standard convolution. Dilate convolution extracts multi-scale informations and standard convolution merges features of different scales. Hence, the output of the ME contains local information and global information. During the up-sampling process, we add a deep supervision block (DSB), which can shorten the length of back-propagation. In this paper, we pay more attention to the importance of shallow features for feature restoration.
Results
Our network validated in the BraTS17’s validation dataset. The DSC scores of MSD-Net for complete tumor, tumor core and enhancing tumor were 0.88, 0.81 and 0.78, respectively, which outperforms most networks.
Conclusion
This study shows that ME enhances the feature extraction ability of the network and improves the accuracy of segmentation results. DSB speeds up the convergence of the network. In addition, we should also pay attention to the contribution of shallow features to feature restoration.






Similar content being viewed by others
References
Ahir BK, Engelhard HH, Lakka SS (2020) Tumor development and angiogenesis in adult brain tumor: Glioblastoma. Mol Neurobiol 57:2461–2478. https://doi.org/10.1007/s12035-020-01892-8
Goetz M, Weber C, Binczyk F, Polanska J, Tarnawski R, Bobek-Billewicz B, Koethe U, Kleesiek J, Stieltjes B, Maier-Hein KH (2015) DALSA: domain adaptation for supervised learning from sparsely annotated MR images. IEEE Trans Med Imaging 35:184–196
Gupta M , Gayatri KS , Harika K , Rao PBVVSN, Rajagopalan V, Das A, Kesavadas C (2015) Brain tumor segmentation by integrating symmetric property with region growing approach. 2015 Annual IEEE India Conference (INDICON) 2015:1-5
Khorram B, Yazdi M (2019) A new optimized thresholding method using ant colony algorithm for MR brain image segmentation. J Digit Imaging 32:162–174
Popuri K, Cobzas D, Murtha A, Jägersand M (2012) 3D variational brain tumor segmentation using Dirichlet priors on a clustered feature set. Int J Comput Assist Radiol Surg 7:493–506. https://doi.org/10.1007/s11548-011-0649-2
Soltaninejad M, Yang G, Lambrou T, Allinson N, Jones TL, Barrick TR, Howe FA, Ye X (2017) Automated brain tumour detection and segmentation using superpixel-based extremely randomized trees in FLAIR MRI. Int J Comput Assist Radiol Surg 12(2):183–203. https://doi.org/10.1007/s11548-016-1483-3
Zeineldin RA, Karar ME, Coburger J, Wirtz CR, Burgert O (2020) DeepSeg: deep neural network framework for automatic brain tumor segmentation using magnetic resonance FLAIR images. Int J Comput Assist Radiol Surg 15:909–920. https://doi.org/10.1007/s11548-020-02186-z
Pereira S, Pinto A, Alves V, Silva CA (2016) Brain tumor segmentation using convolutional neural networks in MRI images. IEEE Trans Med Imaging 35:1240–1251
Hussain S, Anwar SM, Majid M (2018) Segmentation of glioma tumors in brain using deep convolutional neural network. Neurocomputing 282:248–261
Long J, Shelhamer E, Darrell T (2017) Fully convolutional networks for semantic segmentation. IEEE Trans Pattern Anal Mach Intell 39(4):640–651
Ronneberger O (2017) U-net: convolutional networks for biomedical image segmentation. Springer, Berlin
Kong XM, Sun GX, Wu Q, Liu J, Lin FM (2018) Hybrid pyramid U-net model for brain tumor segmentation. In: IFIP Advances in information and communication technology, vol 538
Ding Y, Li C, Yang QQ, Qin Z, Qin ZG (2019) How to improve the deep residual network to segment multi-modal brain tumor images. IEEE Access 7:152821–152831. https://doi.org/10.1109/ACCESS.2019.2948120
Cheng GH, Ji HL, Ding ZX (2020) Spatial-channel relation learning for brain tumor segmentation. Med Phys 47:4885–4894
Wang LS, Xie C, Zeng NY (2019) RP-net: A 3D convolutional neural network for brain segmentation from magnetic resonance imaging. IEEE Access 7:39670–39679
Menze BH, Jakab A, Bauer S, Kalpathy CJ, Farahani K, Kirby J, Burren Y, Porz N, Slotboom J, Wiest R, Lanczi L, Gerstner E, Weber MA, Arbel T, Avants BB, Ayache N, Buendia P, Collins DL, Cordier N, Corso JJ, Criminisi A, Das T, Delingette H, Demiralp Ç, Durst CR, Dojat M, Doyle S, Festa J, Forbes F, Geremia E, Glocker B, Golland P, Guo X, Hamamci A, Iftekharuddin KM, Jena R, John NM, Konukoglu E, Lashkari D, Mariz JA, Meier R, Pereira S, Precup D, Price SJ, Raviv TR, Reza SMS, Ryan M, Sarikaya D, Schwartz L, Shin HC, Shotton J, Silva CA, Sousa N, Subbanna Nagesh K, Szekely G, Taylor TJ, Thomas OM, Tustison NJ, Unal G, Vasseur F, Wintermark M, Ye DH, Zhao L, Zhao B, Zikic D, Prastawa M, Reyes M, Van LK (2015) The Multimodal Brain Tumor Image Segmentation Benchmark(BRATS). IEEE Trans Med Imaging 34(10):1993–2024. https://doi.org/10.1109/TMI.2014.2377694
Wang P, Chen P, Yuan Y, Liu D, Huang Z, Hou X, Cottrell G (2018) Understanding convolution for semantic segmentation. In: 2018 IEEE winter conference on applications of computer vision 2018:1451–1460. doi: https://doi.org/10.1109/WACV.2018.00163
Szegedy C, Vanhoucke V, Ioffe S, Shlens J, Wojna Z (2016) Rethinking the inception architecture for computer vision. In: 2016 IEEE conference on computer vision and pattern recognition 2016:2818–2826. Doi: https://doi.org/10.1109/CVPR.2016.308
He KM, Zhang X, Ren S, Sun J (2016) Deep residual learning for image recognition. In: 2016 IEEE conference on computer vision and pattern recognition (CVPR) 2016:770–778. Doi: https://doi.org/10.1109/CVPR.2016.90
Huang G, Liu Z, Laurens VDM, Weinberger KQ (2017) Densely connected convolutional networks. In: 2017 IEEE conference on computer vision and pattern recognition (CVPR) 2017:2261–2269. Doi: https://doi.org/10.1109/CVPR.2017.243
Zhao LJ, Lu ZX, Jiang J, Zhou YJ, Wu Y, Feng QJ (2019) Automatic nasopharyngeal carcinoma segmentation using fully convolutional networks with auxiliary paths on dual-modality PET-CT images. J Digit Imaging 32(3):462–470. https://doi.org/10.1007/s10278-018-00173-0
Li ZJ, Wang YJ, Yu JH (2018) Brain tumor segmentation using an adversarial network. MICCAI Workshop Cham: Springer 2018:123–132
Kamnitsas K, Bai W, Ferrante E, McDonagh S, Matthew S, Pawlowski N, Rajchl M, Lee M, Kainz B, Rueckert D, Glocker B (2018) Ensembles of multiple models and architectures for robust brain tumour segmentation. In: Crimi A, Bakas S, Kuijf H, Menze B, Reyes M (eds) BrainLes 2017. Springer, Cham, pp 450–462
Ben M-N, Saouli R, Akil M, Kachouri R (2018) Fully automatic brain tumor segmentation using end-to-end incremental deep neural networks in MRI images. Comput Methods Programs Biomed 166:39–49
Author information
Authors and Affiliations
Contributions
The design and verification of MSD-Net are mainly done by BY. At the same time, the first draft is also completed by BY. Miao Cao participated in the polishing and revision of the paper. The preprocessing of the data is done by WG. BW participated in the result analysis.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declared that there is no conflict of interest.
Data availability
The data used in this paper are a public dataset.
Code availability
Python.
Ethical approval
The data used in this paper are a public dataset.
Informed consent
The data used in this paper are a public dataset.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Yan, B., Cao, M., Gong, W. et al. Multi-scale brain tumor segmentation combined with deep supervision. Int J CARS 17, 561–568 (2022). https://doi.org/10.1007/s11548-021-02515-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11548-021-02515-w