Abstract
Purpose
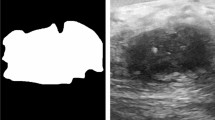
Ultrasound-based navigation is a promising method in breast-conserving surgery, but tumor contouring often requires a radiologist at the time of surgery. Our goal is to develop a real-time automatic neural network-based tumor contouring process for intraoperative guidance. Segmentation accuracy is evaluated by both pixel-based metrics and expert visual rating.
Methods
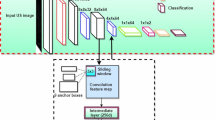
This retrospective study includes 7318 intraoperative ultrasound images acquired from 33 breast cancer patients, randomly split between 80:20 for training and testing. We implement a u-net architecture to label each pixel on ultrasound images as either tumor or healthy breast tissue. Quantitative metrics are calculated to evaluate the model’s accuracy. Contour quality and usability are also assessed by fellowship-trained breast radiologists and surgical oncologists. Additionally, the viability of using our u-net model in an existing surgical navigation system is evaluated by measuring the segmentation frame rate.
Results
The mean dice similarity coefficient of our u-net model is 0.78, with an area under the receiver-operating characteristics curve of 0.94, sensitivity of 0.95, and specificity of 0.67. Expert visual ratings are positive, with 93% of responses rating tumor contour quality at or above 7/10, and 75% of responses rating contour quality at or above 8/10. Real-time tumor segmentation achieved a frame rate of 16 frames-per-second, sufficient for clinical use.
Conclusion
Neural networks trained with intraoperative ultrasound images provide consistent tumor segmentations that are well received by clinicians. These findings suggest that neural networks are a promising adjunct to alleviate radiologist workload as well as improving efficiency in breast-conserving surgery navigation systems.




Similar content being viewed by others
References
Alkabban FM, Ferguson T. Breast Cancer. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2021 [cited 2021 Dec 24]. Available from: http://www.ncbi.nlm.nih.gov/books/NBK482286/
Walters S, Maringe C, Butler J, Rachet B, Barrett-Lee P, Bergh J, Boyages J, Christiansen P, Lee M, Warnberg F, Allemani C, Engholm G, Fornander T, Gjertstorff ML, Johannesen TB, Lawrence G, McGahan CE, Middleton R, Steward J, Tracey E, Turner D, Richards MA, Coleman MP (2013) Breast cancer survival and stage at diagnosis in Australia, Canada, Denmark, Norway, Sweden and the UK, 2000–2007: a population-based study. Br J Cancer 108(5):1195–1208
Chen K, Li S, Li Q, Zhu L, Liu Y, Song E, Fengxi Su (2016) Breast-conserving surgery rates in breast cancer patients with different molecular subtypes: an observational study based on surveillance, epidemiology, and end results (SEER) database. Medicine 95(8):e2593
Maloney BW, McClatchy DM, Pogue BW, Paulsen KD, Wells WA, Barth RJ (2018) Review of methods for intraoperative margin detection for breast conserving surgery. JBO 23(10):100901
Wu W, Su Z, Ma L, Chang J, Cui J (2020) Process analysis and application summary of surgical navigation system. J Complexity Health Sci 3(1):52–61
Janssen N, Eppenga R, VranckenPeeters M-J, Duijnhoven F, Oldenburg H, van der Hage J, Rutgers E, Sonke J, Kuhlmann K, Ruers T, Nijkamp J (2017) Real-time wireless tumor tracking during breast conserving surgery. Int J Comput Assist Radiol Surg 13:13
Gauvin G, Yeo CT, Ungi T, Merchant S, Lasso A, Jabs D, Vaughan T, Rudan JF, Walker R, Fichtinger G, Engel CJ (2020) Real-time electromagnetic navigation for breast-conserving surgery using NaviKnife technology: a matched case-control study. Breast J 26(3):399–405
Pan H, Wu N, Ding H, Ding Q, Dai J, Ling L, Chen L, Zha X, Liu X, Zhou W, Wang S (2013) Intraoperative ultrasound guidance is associated with clear lumpectomy margins for breast cancer: a systematic review and meta-analysis. PLoS ONE 8(9):e74028
Ronneberger O, Fischer P, Brox T (2015) U-Net: Convolutional Networks for Biomedical Image Segmentation. In: Navab N, Hornegger J, Wells WM, Frangi AF (eds) Medical image computing and computer-assisted intervention – MICCAI 2015. Springer International Publishing, Cham, pp 234–241
Zhuang Z, Li N, Joseph Raj AN, Mahesh VGV, Qiu S (2019) An RDAU-NET model for lesion segmentation in breast ultrasound images. PLoS ONE 14(8):e0221535
Byra M, Jarosik P, Szubert A, Galperin M, Ojeda-Fournier H, Olson L, O’Boyle M, Comstock C, Andre M (2020) Breast mass segmentation in ultrasound with selective kernel U-Net convolutional neural network. Biomed Signal Process Control 61:102027
Negi A, Raj ANJ, Nersisson R, Zhuang Z, Murugappan M (2020) RDA-UNET-WGAN: an accurate breast ultrasound lesion segmentation using wasserstein generative adversarial networks. Arab J Sci Eng 45(8):6399–6410
Wang Y, Qin C, Lin C, Lin D, Xu M, Luo X, Wang T, Li A, Ni D (2020) 3D Inception U-net with asymmetric loss for cancer detection in automated breast ultrasound. Med Phys 47(11):5582–5591
Amiri M, Brooks R, Behboodi B, Rivaz H (2020) Two-stage ultrasound image segmentation using U-Net and test time augmentation. Int J CARS 15(6):981–988
Punn NS, Agarwal S (2021) RCA-IUnet: A residual cross-spatial attention guided inception U-Net model for tumor segmentation in breast ultrasound imaging. arXiv:210802508
Guo Y, Duan X, Wang C, Guo H (2021) Segmentation and recognition of breast ultrasound images based on an expanded U-Net. PLoS ONE 16(6):e0253202
Yap MH, Goyal M, Osman FM, Martí R, Denton E, Juette A, Zwiggelaar R (2018) Breast ultrasound lesions recognition: end-to-end deep learning approaches. JMI 6(1):011007
Liao WX, He P, Hao J, Wang XY, Yang RL, An D, Cui LG (2019) Automatic identification of breast ultrasound image based on supervised block-based region segmentation algorithm and features combination migration deep learning model. IEEE J Biomed Health Inform 24(4):984–993
Lazo JF, Moccia S, Frontoni E, De Momi E (2020) Comparison of different CNNs for breast tumor classification from ultrasound images. arXiv:201214517. p 1–6
Kim J, Kim HJ, Kim C, Lee JH, Kim KW, Park YM et al (2021) Weakly-supervised deep learning for ultrasound diagnosis of breast cancer. Sci Rep 11(1):24382
Lasso A, Heffter T, Rankin A, Pinter C, Ungi T, Fichtinger G (2014) PLUS: open-source toolkit for ultrasound-guided intervention systems. IEEE Trans Biomed Eng 61(10):2527–2537
Fedorov A, Beichel R, Kalpathy-Cramer J, Finet J, Fillion-Robin JC, Pujol S, Bauer C, Jennings D, Fennessy F, Sonka M, Buatti J, Aylward S, Miller JV, Pieper S, Kikinis R (2012) 3D Slicer as an image computing platform for the quantitative imaging network. Magn Reson Imaging 30(9):1323–1341
Ungi T, Lasso A, Fichtinger G (2016) Open-source platforms for navigated image-guided interventions. Med Image Anal 33:181–186
Trevethan R (2017) Sensitivity, specificity, and predictive values: foundations, pliabilities, and pitfalls in research and practice. Front Public Health 5:307
Qualtrics. Qualtrics, https://www.qualtrics.com/ (2021, accessed December 1 2020)
Funding
Queen’s University, Department of Pathology and Molecular Medicine, Dean’s Doctoral Award (PVNF). Gabor Fichtinger was supported as Canada Research Chair of the Natural Sciences and Engineering Research Council of Canada.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in this study involving human participants were in accordance with the ethical standards of the institutional research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in this study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Hu, Z., Nasute Fauerbach, P.V., Yeung, C. et al. Real-time automatic tumor segmentation for ultrasound-guided breast-conserving surgery navigation. Int J CARS 17, 1663–1672 (2022). https://doi.org/10.1007/s11548-022-02658-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11548-022-02658-4