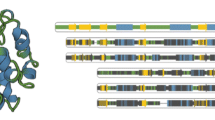
Abstract
The analysis of molecular cavities, which are transport pathways in protein structures, is critical to the understanding of molecular phenomena. However, this work is challenging due to the high complexity and diversity of the macromolecular shapes in dynamic processes. In this paper, we propose a novel multiscale visualization method for visualizing the interaction of protein cavities. We design a series of scales and visualizations of cavities based on both temporal and spatial perspectives to allow domain experts to process their work at any scale of semantic abstraction. These scales demonstrate the chemical and structural properties of cavities and span from a complete protein to a cavity at a specific moment in temporal and spatial dimensions. We also create a continuous interaction space for multiscale applications. Finally, the applicability of our approach is proven through experimental use cases, with cavities in proteins being visualized and analyzed in a focus-and-context manner. Our collaborating domain experts confirmed that our approach is an efficient and reliable method of analyzing cavities with great potential for large dynamic cavity data analysis.
Graphic abstract

















Similar content being viewed by others
References
Asbury TM, Mitman M, Tang J, Zheng WJ (2010) Genome3d: a viewer-model framework for integrating and visualizing multi-scale epigenomic information within a three-dimensional genome. BMC Bioinform 11(1):1–7
Bostock M, Ogievetsky V, Heer J (2011) \({\text{ D }}^{3}\) data-driven documents. IEEE Trans Vis Comput Graph 17(12):2301–2309
Brady GP Jr, Stouten PFW (2000) Fast prediction and visualization of protein binding pockets with pass. J Comput Aided Mol Des 14(4):383–401
Byska J, Jurcik A, Groller ME, Viola I, Kozlikova B (2015a) Molecollar and tunnel heat map visualizations for conveying spatio-temporo-chemical properties across and along protein voids. In: Eurographics conference on visualization
Byska J, Muzic ML, Groller ME, Viola I, Kozlikova B (2015b) Animoaminominer: exploration of protein tunnels and their properties in molecular dynamics. IEEE Trans Vis Comput Graph 22(1):747–756
Chovancova E, Pavelka A, Benes P, Strnad O, Brezovsky J, Kozlikova B, Gora A, Sustr V, Klvana M, Medek P (2012) Caver 3.0: a tool for the analysis of transport pathways in dynamic protein structures. PLoS Comput Biol 8(10):e1002708
Hensen U, Meyer T, Haas J, Rex R, Vriend G, Grubmller H (2012) Exploring protein dynamics space: the dynasome as the missing link between protein structure and function. PLoS ONE 7(5):e33931
Jurcik A, Bednar D, Byska J, Marques SM, Furmanova K, Daniel L, Kokkonen P, Brezovsky J, Strnad O, Stourac J (2018) Caver analyst 2.0: analysis and visualization of channels and tunnels in protein structures and molecular dynamics trajectories. Bioinformatics 34(20):3586–3588
Klvana M, Pavlova MT (2009) Pathways and mechanisms for product release in the engineered haloalkane dehalogenases explored using classical and random acceleration molecular dynamics simulations. J Mol Biol 392(5):1339–1356
Kouřil D, Čmolík L, Kozlíková B, Wu HY, Johnson G, Goodsell DS, Olson A, Gröller ME, Viola I (2019) Labels on levels: labeling of multi-scale multi-instance and crowded 3D biological environments. IEEE Trans Visual Comput Graphics 25(1):977–986
Krone M, Kauker D, Reina G, Ertl T (2014) Visual analysis of dynamic protein cavities and binding sites. In: IEEE pacific visualization symposium
Krone M, Kozlikova B, Lindow N, Baaden M, Baum D, Parulek J, Hege HC, Viola I (2016) Visual analysis of biomolecular cavities: state of the art. Comput Graph Forum 35(3):527–551
Li D, Mei H, Yi S, Shuang S, Zhang W, Wang J, Ming Z, Wei C (2018) Echarts: a declarative framework for rapid construction of web-based visualization. Vis Inform 2:136–146
Lindow N, Baum D, Bondar AN, Hege HC (2013) Exploring cavity dynamics in biomolecular systems. BMC Bioinform 14(S19):S5
Liskova V, Bednar D, Prudnikova T, Rezacova P, Damborsky J (2015) Balancing the stability-activity trade-off by fine-tuning dehalogenase access tunnels. Chemcatchem 7(4):648–659
Martin K, Andrew MJ (2002) Molecular dynamics simulations of biomolecules. Nat Struct Biol 9(9):646–52
Masood TB, Sandhya S, Chandra N, Natarajan V (2015) Chexvis: a tool for molecular channel extraction and visualization. BMC Bioinform 16(1):1–19
Medek P, Benes P, Sochor J (2007) Computation of tunnels in protein molecules using delaunay triangulation. J WSCG 15:107–114
Miao H, De Llano E, Sorger J, Ahmadi Y, Kekic T, Isenberg T, Gröller ME, Barišić I, Viola I (2017) Multiscale visualization and scale-adaptive modification of DNA nanostructures. IEEE Trans Vis Comput Graph 24(1):1014–1024
Miao H, Klein T, Kouril D, Mindek P, Schatz K, Groller ME, Kozlikova B, Isenberg T, Viola I (2018) Multiscale molecular visualization. J Mol Biol 431(6):1049–1070
Parulek J, Turkay C, Reuter N, Viola I (2012) Implicit surfaces for interactive graph based cavity analysis of molecular simulations. In: IEEE symposium on biological data visualization
Parulek J, Turkay C, Reuter N, Viola I (2013) Visual cavity analysis in molecular simulations. BMC Bioinform 19(14 Suppl 19):S4
Petrek M, Otyepka M, Banas P, Kosinova P, Koca J, Damborsky J (2006) Caver: a new tool to explore routes from protein clefts, pockets and cavities. BMC Bioinform 7(1):316
Petrek M, Kosinova P, Koca J, Otyepka M (2007) Mole: a voronoi diagram-based explorer of molecular channels, pores, and tunnels. Structure 15(11):1357–1363
Sehnal D, Varekova RS, Berka K, Pravda L, Navratilova V, Banas P, Ionescu CM, Otyepka M, Koca J (2013) Mole 2.0: advanced approach for analysis of biomacromolecular channels. J Cheminform 5(1):39–39
Sehnal D, Deshpande M, Varekova RS, Mir S, Berka K, Midlik A, Pravda L, Velankar S, Koca J (2017) Litemol suite: interactive web-based visualization of large-scale macromolecular structure data. Nat Methods 14(12):1121. https://doi.org/10.1038/nmeth.4499
Smart OS, Neduvelil JG, Wang X, Wallace BA, Sansom MS (1996) Hole: a program for the analysis of the pore dimensions of ion channel structural models. J Mol Graph 14(6):354–360
Sykora J, Brezovsky J, Koudelakova T, Lahoda M, Fortova A, Chernovets T, Chaloupkova R, Stepankova V, Prokop Z, Smatanova IK (2014) Dynamics and hydration explain failed functional transformation in dehalogenase design. Nat Chem Biol 10(6):S90
Waldin N, Muzic ML, Waldner M, Groller E, Goodsell D, Ludovic A, Viola I (2016) Chameleon—dynamic color mapping for multi-scale structural biology models. In: Bruckner S, Preim B, Vilanova A, Hauser H, Hennemuth A, Lundervold A (eds) Eurographics workshop on visual computing for biology and medicine. The Eurographics Association. https://doi.org/10.2312/vcbm.20161266
Zhou HX, Mccammon JA (2010) The gates of ion channels and enzymes. Trends Biochem Sci 35(3):179–185
Acknowledgements
Our deepest gratitude goes to the developers of Caver Analyst 2.0 (Jurcik et al. 2018) for the experimental data provided. This work was supported in part by grants from the National Key R&D Program of China and the National Science Foundation of China (Grant No. 61463042).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary material 1 (mp4 21687 KB)
Rights and permissions
About this article
Cite this article
Guo, D., Han, D., Xu, X. et al. Spatiotemporal multiscale molecular cavity visualization and visual analysis. J Vis 23, 661–676 (2020). https://doi.org/10.1007/s12650-020-00646-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12650-020-00646-x