Abstract
Early identification of the type of skin lesion, some of them carcinogenic, is of paramount importance. Currently, the use of Convolutional Neural Networks (CNNs) is the mainline of investigation for the automated analysis of such lesions. Most of the existing works, however, were designed by transfer learning general-purpose CNN architectures, adapting existing methods to the domain of dermatology. Despite effective, this approach poses inflexibility and high processing costs. In this work, we introduce a novel architecture that benefits from cutting-edge CNN techniques Aggregated Transformations combined to the mechanism of Squeeze-and-Excite organized in a residual block; our architecture is designed and trained from scratch to solve both the binary melanoma detection problem, as well as the multi-class skin-lesion classification problem. Our results demonstrate that such an architecture is competitive to major state-of-the-art architectures adapted to the domain of skin-lesion diagnosis. Our architecture is prone to evolve and to provide low processing cost for real-world in situ applications using a much smaller number of weights if compared to previous works.




Similar content being viewed by others
References
Cheng YI, Swamisai R, Umbaugh SE, Moss RH, Stoecker WV, Teegala S, Srinivasan SK. Skin lesion classification using relative color features. Skin Res Technol. 2008;14(1):53–64.
Menzies SW, Bischof L, Talbot H, Gutenev A, Avramidis M, Wong L, Lo SK, Mackellar G, Skladnev V, McCarthy W, et al. The performance of solarscan: an automated dermoscopy image analysis instrument for the diagnosis of primary melanoma. Arch Dermatol. 2005;141(11):1388–96.
T. A. C. Society, Key statistics for melanoma skin cancer. https://www.cancer.org/cancer/melanoma-skin-cancer/about/key-statistics.html. Accessed Dec 2020.
Schadendorf D, van Akkooi ACJ, Berking C, Griewank KG, Gutzmer R, Hauschild A, Stang A, Roesch A, Ugurel S. Melanoma. The Lancet. 2018;392(10151):971–84.
Jerant AF, Johnson JT, Sheridan CD, Caffrey TJ. Early detection and treatment of skin cancer. Am Fam Phys. 2000;62(2):357–68.
Esteva A, Kuprel B, Novoa RA, Ko J, Swetter SM, Blau HM, Thrun S. Dermatologist-level classification of skin cancer with deep neural networks. Nature. 2017;542:115–8.
Codella NC, Nguyen Q-B, Pankanti S, Gutman DA, Helba B, Halpern AC, Smith JR. Deep learning ensembles for melanoma recognition in dermoscopy images. IBM J Res Dev. 2017;61(4/5):5–1.
Fujisawa Y, Otomo Y, Ogata Y, Nakamura Y, Fujita R, Ishitsuka Y, Watanabe R, Okiyama N, Ohara K, Fujimoto M. Deep-learning-based, computer-aided classifier developed with a small dataset of clinical images surpasses board-certified dermatologists in skin tumour diagnosis. Br J Dermatol. 2018;180(2):373–81.
Menegola A, Tavares J, Fornaciali M, Li LT, de Avila SEF, Valle E. RECOD titans at ISIC challenge 2017. arXiv:1703.04819
Han SS, Kim MS, Lim W, Park GH, Park I, Chang SE. Classification of the clinical images for benign and malignant cutaneous tumors using a deep learning algorithm. J Invest Dermatol. 2018;138(7):1529–38.
Hameed N, Shabut AM, Hossain MA. Multi-class skin diseases classification using deep convolutional neural network and support vector machine. In 2018 12th International Conference on Software, Knowledge, Information Management Applications (SKIMA), 2018, pp. 1–7.
Kawahara J, Daneshvar S, Argenziano G, Hamarneh G. Seven-point checklist and skin lesion classification using multitask multimodal neural nets. IEEE J Biomed Health Inf. 2018; 23(2):538–546.
Harangi B, Baran A, Hajdu A. Assisted deep learning framework for multi-class skin lesion classification considering a binary classification support. Biomed Signal Process Control, 2020; v. 62, p. 102041.
Gessert N, Sentker T, Madesta F, Schmitz R, Kniep H, Baltruschat I, Werner R, Schlaefer A. Skin lesion classification using cnns with patch-based attention and diagnosis-guided loss weighting. IEEE Trans Biomed Eng., 2019; v. 67, n. 2, p. 495–503.
Torrey L, Shavlik J. Transfer learning. In Handbook of research on machine learning applications and trends: algorithms, methods, and techniques. IGI Global. 2010;242–264.
Russakovsky O, Deng J, Su H, Krause J, Satheesh S, Ma S, Huang Z, Karpathy A, Khosla A, Bernstein M, Berg A, Fei-Fei L. ImageNet large scale visual recognition challenge. Int J Comput Vis (IJCV). 2015;115(3):211–52.
Xie S, Girshick R, Dollar P, Tu Z, He K. Aggregated residual transformations for deep neural networks. In: IEEE conference on computer vision and pattern recognition, 2017;pp. 1492–1500.
Hu J, Shen L, Sun G. Squeeze-and-excitation networks. In: Proceedings of the IEEE conference on computer vision and pattern recognition, 2018;pp. 7132–7141.
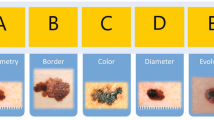
Abbasi, Naheed R., et al. Early diagnosis of cutaneous melanoma: Revisiting the ABCD criteria. JAMA 2004; 292(22), 2771–2776.
Krizhevsky A, Sutskever I, Hinton G. “Imagenet classification with deep convolutional neural networks,” in Advances in Neural Information Processing Systems. Curran Associates. 2012;1097–105.
Szegedy C, Vanhoucke V, Ioffe S, Shlens J, Wojna Z. “Rethinking the inception architecture for computer vision,” in The IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2016; pp. 2818–2826.
Codella NCF, Gutman D, Celebi ME, Helba B, Marchetti MA, Dusza SW, Kalloo A, Liopyris K, Mishra N, Kittler H, Halpern A. Skin lesion analysis toward melanoma detection: A challenge at the 2017 international symposium on biomedical imaging. In IEEE Biomedical Imaging, 2018; pp. 168–172.
Simonyan K, Zisserman A. Very deep convolutional networks for large-scale image recognition. 2014. arXiv:1409.1556
He K, Zhang X, Ren S, Sun J. Deep residual learning for image recognition. In Proceedings of the IEEE conference on computer vision and pattern recognition, 2016;pp. 770–778
Zakhem GA, Motosko CC, Ho RS. How should artificial intelligence screen for skin cancer and deliver diagnostic predictions to patients? JAMA Dermatol. 2018;154(12):1383–4.
Ruiz D, Berenguer V, Soriano A, Sanchez B. A decision support system for the diagnosis of melanoma: a comparative approach. Expert Syst Appl. 2011;38(12), 15217–15223
Yu L, Chen H, Dou Q, Qin J, Heng PA. Automated melanoma recognition in dermoscopy images via very deep residual networks. IEEE Trans Med Imaging. 2017;36(4):994–1004.
Ganzeli H, Bottesini J, Paz L, Ribeiro M. Skan: Skin scanner - system for skin cancer detection using adaptive techniques. IEEE Lat Am Trans. 2011;9(2):206–12.
Aswin RB, Jaleel JA, Salim S. Hybrid genetic algorithm - artificial neural network classifier for skin cancer detection. In Control instrumentation: communication and computational technologies, 2014; 1304–1309.
Nasr-Esfahani E, Samavi S, Karimi N, Soroushmehr SMR, Jafari MH, Ward K, Najarian K. Melanoma detection by analysis of clinical images using convolutional neural network. In IEEE Engineering in Medicine and Biology Society, 2016;pp. 1373–1376.
Majtner T, Yildirim-Yayilgan S, Hardeberg JY. Combining deep learning and hand-crafted features for skin lesion classification. In: International Conference on Image Processing Theory, Tools and Applications, 2016; pp. 1–6.
Gonzalez R, Woods R, Eddins S. Digital image processing using MATLAB. Pearson, 2004.
Barata C, Ruela M, Francisco M, Mendonca T, Marques J. Two systems for the detection of melanomas in dermoscopy images using texture and color features. IEEE systems Journal, 2013; v. 8, n. 3, p. 965–979.
Li Y, Shen L. Skin lesion analysis towards melanoma detection using deep learning network. Sensors. 2018;18(2):556.
Tan M, Le QV. Efficientnet: Rethinking model scaling for convolutional neural networks. In: Proceedings of the 36th International Conference on Machine Learning (ICML), 2019; p. 6105–6114.
Acknowledgements
This research was financed by French agency Multidisciplinary Institute in Artificial Intelligence (Grenoble Alpes, ANR-19-P3IA-0003); and by Brazilian agencies Fundacao de Amparo a Pesquisa do Estado de Sao Paulo (2018/17620-5, and 2016/17078-0); Conselho Nacional de Desenvolvimento Cientifico e Tecnologico (406550/2018-2, and 305580/2017-5); and Coordenacao de Aperfeicoamento de Pessoal de Nivel Superior (CAPES, Finance Code 001). We thank NVidia for donating the GPUs that supported this work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest statement
On behalf of all the authors, the corresponding author states that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the topical collection “AI and Deep Learning Trends in Healthcare” guest edited by KC Santosh, Paolo Soda and Zalelam Temesgen.
Rights and permissions
About this article
Cite this article
Lima, D.M., Rodrigues-Jr, J.F., Brandoli, B. et al. DermaDL: Advanced Convolutional Neural Networks for Computer-Aided Skin-Lesion Classification. SN COMPUT. SCI. 2, 253 (2021). https://doi.org/10.1007/s42979-021-00641-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42979-021-00641-5