Abstract
Cellular signaling (or signal transduction) is the process by which extracellular signals are converted into cellular responses. Mathematical models of signaling pathways may help understanding their general regulatory principles, as well as the roles of the different components often involved in more than one pathway.
The aim of the present work is to describe a discrete model in time and space to study the dynamics of the population of molecules involved in a specific pathway. To this purpose, we use a multi-compartment stochastic cellular automaton to simulate one of the best understood cell signaling pathways: the GPCR-pathway.
We then show the effect on the signal-carrying efficiency of condensing a molecular species, which is pivotal to the reaction pathway, in a closed region of the cytosol.
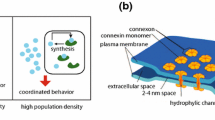
We find that localization increases the robustness of the translocation mechanism only above a density threshold. For homogeneous settings, by increasing the concentration above a critical value, the translocation is obstructed by mere physical occupation constraints of the signal-carrying molecule. This conclusion is in line with previous findings, according to which in highly dense regions of the cytosol, convection by autocatalytic activation of adjacent molecules might be more efficient than diffusion as a main signal propagation mechanism.
Similar content being viewed by others
References
Boon, J., Dab, D., Kapral, R., Lawniczak, A., 1996. Lattice gas automata for reactive systems. Phys. Rep. 273, 55–148.
Castiglione, F., Succi, S., Bernaschi, M., Heinrich, R., Kirschner, M., 2002. Intracellular signal propagation in a two-dimensional autocatalytic reaction model. Phys. Rev. E 66, 031905.
Chopard, B., Droz, M., 1998. Cellular Automata Modeling of Physical Systems. Cambridge University Press, Cambridge.
Endy, D., Brent, R., 2001. Modelling cellular behaviour. Nature 409, 391–395.
Gibbs, W., 2001. Cybernetic cells. Sci. Am. August, 53–57.
Hartwell, L., Hopfield, J., Leibler, S., Murray, A., 1999. From molecular to modular cell biology. Nature 402, C47-C52.
Lee, E., Salic, A., Krger, R., Heinrich, R., Kirschner, M., 2003. The roles of ape and axin derived from experimental and theoretical analysis of the wnt pathway. PLOS Biol. 1, 116–132.
Lodish, J., Berk, A., Zipurky, S., Matsudaira, P., Baltimore, D., Darnell, J., 2000. Molecular Cell Biology, fourth ed., W.H. Freeman and Company, New York, NY.
Murray, J., 1989. Mathematical Biology, Springer, Berlin.
Polakis, P., 2000. Wnt signalling and cancer. Genes Dev. 14, 1837–1851.
Robinson, M., Cobb, M., 1997. Mitogen-activated protein kinase pathway. Curr. Opin. Cell Biol. 9, 180–186.
Salic, A., Lee, E., Mayer, L., Kirschner, M., 2000. Control of β-catenin stability: reconstitution of the cytoplasmic steps of the wnt pathway in xenopus egg extract. Mol. Cell 5, 523–532.
Schaff, J., Loew, L., 1999. E-cell: software environment for whole cell simulation. In: Proceedings of the Pacific Symposium in Biocomputing, vol. 15, pp. 228–239.
Tomita, K., Hashimoto, K., Takahashi, K., Simizu, T., Matsuzaki, Y., Miyoshi, F., Saito, K., Tanida, S., Yugi, K., Venter, J., Hutchison, C.I., 1999. E-cell: software environment for whole cell simulation. Bioinformatics 15, 72–84.
Weimar, J., Boon, J., Dab, D., Succi, S., 1992. Fluctuation correlations in reaction-diffusion systems: reactive lattice gas automata approach. Europhys. Lett. 20, 627–632.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Castiglione, F., Succi, S. Simulating the G-protein cAMP pathway with a two-compartment reactive lattice gas. Theory Biosci. 123, 413–429 (2005). https://doi.org/10.1016/j.thbio.2004.10.003
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1016/j.thbio.2004.10.003