Abstract
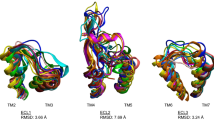
Some key concerns raised by molecular modeling and computational simulation of functional mechanisms for membrane proteins are discussed and illustrated for members of the family of G protein coupled receptors (GPCRs). Of particular importance are issues related to the modeling and computational treatment of loop regions. These are demonstrated here with results from different levels of computational simulations applied to the structures of rhodopsin and a model of the 5-HT2A serotonin receptor, 5-HT2AR. First, comparative Molecular Dynamics (MD) simulations are reported for rhodopsin in vacuum and embedded in an explicit representation of the membrane and water environment. It is shown that in spite of a partial accounting of solvent screening effects by neutralization of charged side chains, vacuum MD simulations can lead to severe distortions of the loop structures. The primary source of the distortion appears to be formation of artifactual H-bonds, as has been repeatedly observed in vacuum simulations. To address such shortcomings, a recently proposed approach that has been developed for calculating the structure of segments that connect elements of secondary structure with known coordinates, is applied to 5-HT2AR to obtain an initial representation of the loops connecting the transmembrane (TM) helices. The approach consists of a simulated annealing combined with biased scaled collective variables Monte Carlo technique, and is applied to loops connecting the TM segments on both the extra-cellular and the cytoplasmic sides of the receptor. Although this initial calculation treats the loops as independent structural entities, the final structure exhibits a number of interloop interactions that may have functional significance. Finally, it is shown here that in the case where a given loop from two different GPCRs (here rhodopsin and 5-HT2AR) has approximately the same length and some degree of sequence identity, the fold adopted by the loops can be similar. Thus, in such special cases homology modeling might be used to obtain initial structures of these loops. Notably, however, all other loops in these two receptors appear to be very different in sequence and structure, so that their conformations can be found reliably only by ab initio, energy based methods and not by homology modeling.
Similar content being viewed by others
References
Visiers, I., Ballesteros, J.A. and Weinstein, H., Meth. Enzymol., 343 (2002) 329.
Palczewski, K., Kumasaka, T., Hori, T., Behnke, C.A., Motoshima, A., Fox, B.A., LeTrong, I., Teller, D.C., Okada, T., Stenkamp, R.E., Yamamoto, M. and Miyano, M., Science, 289 (2000) 739.
Ballesteros, J.A., Shi, L. and Javitch, J.A., Mol. Pharmacol., 60(1) (2001) 1.
Shi, L. and Javitch, J.A., Annu. Rev. Pharmacol. Toxicol. 42 (2002) 437.
Almaula, N., Ebersole, B.J., Zhang, D., Weinstein, H. and Sealfon, S., J. Biol. Chem., 271 (1996) 14672.
Ebersole, B.J., Visiers, I., Weinstein, H. and Sealfon, S., Mol. Pharmacol., (2002), In press.
Chalmers, D.T. and Behan, D.P., Nature reviews-Drug Discovery, 1 (2002) 599.
George, S.R., O'Dowd, B.F. and Lee, S.P., Nature reviews-Drug Discovery, 1 (2002) 808.
Visiers, I., Ebersole, B.J., Dracheva, S., Ballesteros, J., Sealfon, S.C. and Weinstein, H., Intl. J. Quantum. Chem., 88 (2002) 65.
Prioleau, C., Visiers, I., Ebersole, B.J., Weinstein, H. and Sealfon, S., J. Biol. Chem., 277 (2002) 36577.
Visiers, I., Hassan, S.A. and Weinstein, H., Prot. Eng., 14 (2001) 409.
Huang, P., Visiers, I., Weinstein, H. and Liu-Chen, L.-Y., Biochemistry, 40 (2001) 13501.
Hassan, S.A. and Mehler, E.L., Int. J. Quant. Chem., 83 (2001) 193.
Schaefer, M., Bartels, C. and Karplus, M., J. Mol. Biol., 284 (1998) 835.
Mohanty, D., Elber, R., Thirumalai, D., Beglov, D. and Roux, B., J. Mol. Biol., 272 (1997) 423.
Ma, B. and Nussinov, R., PROTEINS: Structure, Function, and Genetics, 37 (1999) 73.
Fanelli, F., J. Mol. Biol., 296 (2000) 1333.
Chilmonczyk, Z., Cybulski, M., Iskra-Jopa, J., Chojnacka-Wójcik, E., Tatarczyska, E., Kodziska, A., Les, A., Bronowska, A. and Sylte, I. Il. Farmaco., 57 (2002) 285.
Bronowska, A., Chilmonczyk, Z., Les, A., Edvardsen, O., Østensen, R. and Sylte, I., J. Comp. Aid. Mol. Des., 15 (2001) 1005.
Hovelmann, S., Hoffmann, S.H., Kuhne, R., ter Laak, T., Reilaender, H. and Beckers, T., Biochemistry, 41 (2002) 1129.
Pierce, K.L., Premont, R.T. and Lefkowitz, R.J., Nature reviews-Mol. Cell Biol., 3 (2002) 639.
Hassan, S.A., Mehler, E.L. and Weinstein, H., In: Schlick, T. and Gan, H.H. (Eds.), Lecture Notes in Computational Science and Engineering, Vol. 24., Springer Verlag, New York, pp. 197–231 (2002).
Higo, J., Collura, V. and Garnier, J., Biopolymers, 32 (1992) 33.
Carlacci, L. and Englander, S.W., J. Comp. Chem., 17(8) (1996) 1002–1012.
Baysal, C. and Meirovitch, H., J. Comp. Chem., 20(15) (1999) 1659.
Nakajima, N., Higo, J., Kidera, A. et al., J. Mol. Biol., 296 (2000) 197. 853
Kidera, A., Proc. Natl. Acad. Sci. U.S.A., 92 (1995) 9886.
Still, W.C., Tempczyk, A., Hawley, R.C. and Hendrickson, T., J. Am. Chem. Soc., 112 (1990) 6127.
Hassan, S.A., Guarnieri, F. and Mehler, E.L., J. Phys. Chem., B104 (2000) 6478.
Vasmatzis, L., Brower, R. and Delisi, C., Biopolymers, 34 (1994) 1669.
Zheng, Q., Rosenfeld, R., Delisi, C. and Kyle, D.J.,Protein Sci., 9 (1994) 493–506.
Fiser, A., Kinh Gian Do, R. and Sali, A., Protein Science, 9 (2000) 1753.
Berendsen, H.J.C., van der Spoel, D. and van Drunen, R., Comp. Phys. Comm., 91 (1995) 43–56.
van Gunsteren, W.F., Billeter, S.R., Eising, A.A., Huenenberger, P.H., Krueger, P., Mark, A.E., Scott, W.R.P. and Tironi, I.G., HochschulverlagAG an der ETH Zuerich.: (1996) Zuerich.
Darden, T., York, D. and Pederson, L., J. Chem. Phys., 98 (1993) 10089.
Okada, T., Fujiyoshi, Y., Silow, M., Navarro, J., Landau, E.M. and Shichida, Y., PNAS, 99 (2002) 5982.
Teller, D.C., Okada, T., Behnke, C.A., Palczewski, K. and Stenkamp, R.E., Biochemistry, (2001) 7761.
Brooks, B.R., Bruccoleri, R.E., Olafson, B.D., States, D.J., Swaminathan, S. and Karplus, M., J. Comput. Chem., 4 (1983) 187.
MacKerell, J., A. D., Bashford, D., Bellott, M., Dunbrack Jr., R.L., Evanseck, J.D., Field, M.J., Fischer, S., Gao, J., Guo, H., Ha, S., Joseph-McCarthy, D., Kuchnir, L., Kuczera, K., Lau, F.T.K., Mattos, C., Michnick, S., Ngo, T., Nguyen, D.T., Prodhom, B., Reiher, I., W.E., Roux, B., Schlenkrich, M., Smith, J.C., Stote, R., Straub, J., Watanabe, M., Wiorkiewicz-Kuczera, J., Yin, D. and Karplus, M., J. Phys. Chem. B, 102 (1998) 3586.
Hassan, S.A., Guarnieri, F. and Mehler, E.L., J. Phys. Chem., B104 (2000) 6490.
Ehrenson, S., J. Comp. Chem., 10 (1989) 77.
Bucher, M. and Porter, T.L., J. Phys. Chem., 90 (1986) 3406.
Webb, T.J., J. Am. Chem. Soc., 48 (1926) 2589.
Debye, P. and Pauling, L., J. Am. Chem. Soc., 47 (1925) 2129.
Schwarzenbach, G., Z. physik. Chem., A176 (1936) 133.
Harvey, S.C. and Hoekstra, P., J. Phys. Chem., 76 (1972) 2987.
Pennock, B.D. and Schwan, H.P., J. Phys. Chem., 73 (1969) 2600.
Takashima, S. and Schwan, H.P., J. Phys. Chem., 69 (1965) 4176.
Hasted, J.B., Ritson, D.M. and Collie, C.H., J. Chem. Phys., 16 (1948) 1.
Mehler, E.L., In: Murray, J.S. and Sen, K. (Eds), Molecular Electrostatic Potential: Concepts and Applications, Elsevier Science, Amsterdam, p. 371–405 (1996).
Hassan, S.A. and Mehler, E.L., PROTEINS: Stru. Func. Genet., 47(2002) 45.
Noguti, T. and Go, N., Biopolymers, 24 (1985) 527.
Lazaridis, T. and Karplus, M., PROTEINS: Struc. Func. Gen., 35 (1999) 133.
Shenkin,6P.S. and McDonald, D.Q., J. Comput. Chem., 15 (1994) 899.
Ballesteros, J.A. and Weinstein, H., Meth. Neurosci., 25 (1995) 366.
Visiers, I., Ballesteros, J.A. and Weinstein, H., In: Iyengar, I. and edJ. Hildebrandt, (Eds), Methods Enzymol, Academic Press: New York (2001).
Wong, S.K., Slaughter, C., Ruoho, A.E. and Ross, E.M., J. Biol. Chem., 263 (1988) 7925.
Konig, B., Arendt, A., McDowell, J.H., Kahlert, M., Hargrave, P.A. and Hoffman, K.P., Proc. Natl. Acad Sci., 86 (1989) 6878.
Cypess, A.M., Unson, C.G., Wu, C.R. and Sakmar, T.P., J. Biol. Chem., 274 (1999) 19455.
Niswender, C.M., Copeland, S.C., Herrick-Davis, K, Emeson, R.B. and Sanders-Bush, E., J. Biol. Chem., 274 (1999) 9472.
Tartaglia, M., Mehler, E.L., Goldberg, R., Zampino, G., Brunner, H.G., Kremer, H., vav der Burgt, I., Crosby, A.H., Ion, A., Jeffery, S., Kalidas, K., Patton, M.A., Kucherlapati, R.S. and Gelb, B.D., Nature Genet., 29 (2001) 465.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Mehler, E., Periole, X., Hassan, S. et al. Key issues in the computational simulation of GPCR function: representation of loop domains. J Comput Aided Mol Des 16, 841–853 (2002). https://doi.org/10.1023/A:1023845015343
Issue Date:
DOI: https://doi.org/10.1023/A:1023845015343