Abstract
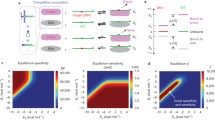
Since they minimize errors from cross-hybridizations, DNA oligonucleotides that annealas designed are beneficial to DNA computing. By in vitro selection, huge libraries of non-crosshybridizing oligonucleotides might be evolved in the test tube. As a first step, a fitness function corresponding to non-crosshybridization was based upon the duplex stability of randomly matched oligonucleotides. By melting pairs that have a low thermal stability, a protocol based on DNA polymerization selectively amplifies maximally mismatched oligonucleotides over those that were more closely matched. Experiments confirmed this property of the protocol, and in addition, a reaction temperature window was identified in which discrimination between matched and mismatched might be obtained. The protocol was iterated on a set of random starting material, and there was evidence that non-crosshybridizing libraries were in fact being created. These results are a step toward practical manufacture of very large libraries of non-crosshybridizing oligonucleotides in the test tube.
Similar content being viewed by others
References
Adleman LM (1994) Molecular computation of solutions to combinatorial problems. Science 266: 1021–1024
Baum EB (1998) DNA sequences useful for computation. In: Landweber LF and Baum EB (eds) Proceedings of the Second Annual Meeting on DNA Based Computers, Volume 44, Providence, RI, DIMACS, American Mathematical Society, pp. 122–127. DIMACS Workshop, Princeton, NJ, June 10-12, 1996
Deaton R, Murphy RC, Rose JA, Garzon M, Franceschetti DR, Stevens SE Jr (1997) A DNA based implementation of an evolutionary search for good encodings for DNA computation. In: Proceedings of the 1997 IEEE International Conference on Evolutionary Computation, IEEE, pp. 267–272. Indianapolis, IN, April 13-16
Deaton R, Chen J, Bi H and Rose J (2002) A software tool for generating non-crosshybridizing libraries of DNA oligonucleotides. In: Preliminary Proceedings of the Eighth International Meeting on DNA Based Computers, Sapporo, Japan, June 2002, pp. 211–220
Lipton RJ (1995) DNA solution of hard computational problems. Science 268: 542–545
Lipton R (2001) DNA computing: does it compute? Plenary address at the Seventh International Meeting on DNA Based Computers
Marathe A, Condon AE and Corn RM (1999) On combinatorial DNA word design. In: Winfree E and Gifford DK (eds) DNA Based Computers V, Providence, RI, DIMACS, American Mathematical Society, pp. 75–90. DIMACS Workshop, Massachusetts Institute of Technology, Cambridge, MA, June 14-16, 1999
Osborne SE and Ellington AD (1997) Nucleic acid selection and the challenge of combinatorial chemistry. Chemical Review 97: 349–370
SantaLucia J Jr. (1998) A unified view of polymer, dumbbell, and oligonucleotide DNA nearest-neighbor thermodynamics. Proc. Natl. Acad. Sci. 95: 1460–1465
Smith TF and Waterman MS (1981) The identification of common molecular subsequences. J. Mol. Biol. 147: 195–197
Winfree E, Liu F, Wenzler LA and Seeman NC (1998) Design and self-assembly of twodimensional DNA crystals. Nature 394: 539–544
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Bi, H., Chen, J., Deaton, R. et al. In vitro selection of non-crosshybridizing oligonucleotides for computation. Natural Computing 2, 417–426 (2003). https://doi.org/10.1023/B:NACO.0000006772.32105.46
Issue Date:
DOI: https://doi.org/10.1023/B:NACO.0000006772.32105.46