Abstract
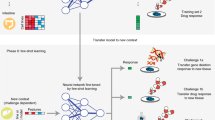
Data discrepancy between preclinical and clinical datasets poses a major challenge for accurate drug response prediction based on gene expression data. Different methods of transfer learning have been proposed to address such data discrepancy in drug response prediction for different cancers. These methods generally use cell lines as source domains, and patients, patient-derived xenografts or other cell lines as target domains; however, it is assumed that the methods have access to the target domain during training or fine-tuning, and they can only take labelled source domains as input. The former is a strong assumption that is not satisfied during deployment of these models in the clinic, whereas the latter means these methods rely on labelled source domains that are of limited size. To avoid these assumptions, we formulate drug response prediction in cancer as an out-of-distribution generalization problem, which does not assume that the target domain is accessible during training. Moreover, to exploit unlabelled source domain data—which tends to be much more plentiful than labelled data—we adopt a semi-supervised approach. We propose Velodrome, a semi-supervised method of out-of-distribution generalization that takes labelled and unlabelled data from different resources as input and makes generalizable predictions. Velodrome achieves this goal by introducing an objective function that combines a supervised loss for accurate prediction, an alignment loss for generalization and a consistency loss to incorporate unlabelled samples. Our experimental results demonstrate that Velodrome outperforms state-of-the-art pharmacogenomics and transfer learning baselines on cell lines, patient-derived xenografts and patients. Finally, we showed that Velodrome models generalize to different tissue types that were well-represented, under-represented or completely absent in the training data. Overall, our results suggest that Velodrome may guide precision oncology more accurately.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All the final preprocessed data employed in this paper are publicly available here: https://zenodo.org/record/4793442#.YK1HVqhKiUk (ref. 76). All the raw data before preprocessing are also publicly available as follows: (1) cell-line datasets with gene expression and drug response data, including CTRPv2, GDSCv2 and gCSI, were downloaded from ORCESTRA69; (2) TCGA cohorts with gene expression data were downloaded from Firehose (http://gdac.broadinstitute.org/) on 28 January 2016. Drug response data for TCGA cohorts was obtained from ref. 39; (3) PDX datasets (gene expression with drug response data) were obtained from the Supplementary Information of ref. 3; (4) Patient dataset (gene expression with drug response data) were obtained from the accession codes GSE25065 (Docetaxel and Paclitaxel) and GSE33072 (Erlotinib). Source data are provided with this paper.
Code availability
All the codes, model objects and supplementary material used to run and reproduce our experimental results are publicly available at https://github.com/hosseinshn/Velodrome (ref. 77). We also provided a conda environment to ensure version compatibility for future users.
References
Marquart, J., Chen, E. Y. & Prasad, V. Estimation of the percentage of US patients with cancer who benefit from genome-driven oncology. JAMA Oncol. 4, 1093–1098 (2018).
Pal, S. K. et al. Clinical cancer advances 2019: annual report on progress against cancer from the American society of clinical oncology. J. Clin. Oncol. 37, 834–849 (2019).
Gao, H. et al. High-throughput screening using patient-derived tumor xenografts to predict clinical trial drug response. Nat. Med. 21, 1318–1325 (2015).
Garnett, M. J. et al. Systematic identification of genomic markers of drug sensitivity in cancer cells. Nature 483, 570–575 (2012).
Barretina, J. et al. The cancer cell line encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–607 (2012).
Basu, A. et al. An interactive resource to identify cancer genetic and lineage dependencies targeted by small molecules. Cell 154, 1151–1161 (2013).
Seashore-Ludlow, B. et al. Harnessing connectivity in a large-scale small-molecule sensitivity dataset. Cancer Discov. 5, 1210–1223 (2015).
Klijn, C. et al. A comprehensive transcriptional portrait of human cancer cell lines. Nat. Biotechnol. 33, 306–312 (2015).
Iorio, F. et al. A landscape of pharmacogenomic interactions in cancer. Cell 166, 740–754 (2016).
Haverty, P. M. et al. Reproducible pharmacogenomic profiling of cancer cell line panels. Nature 533, 333–337 (2016).
Mourragui, S., Loog, M., van de Wiel, M. A., Reinders, M. J. T. & Wessels, L. F. A. PRECISE: a domain adaptation approach to transfer predictors of drug response from pre-clinical models to tumors. Bioinformatics 35, i510–i519 (2019).
Sharifi-Noghabi, H., Peng, S., Zolotareva, O., Collins, C. C. & Ester, M. AITL: Adversarial Inductive Transfer Learning with input and output space adaptation for pharmacogenomics. Bioinformatics 36, i380–i388 (2020).
Haibe-Kains, B. et al. Inconsistency in large pharmacogenomic studies. Nature 504, 389–393 (2013).
Mpindi, J. P. et al. Consistency in drug response profiling. Nature 540, E5–E6 (2016).
Geeleher, P., Gamazon, E. R., Seoighe, C., Cox, N. J. & Huang, R. S. Consistency in large pharmacogenomic studies. Nature 540, E1–E2 (2016).
Pan, S. J. & Yang, Q. A survey on transfer learning. IEEE Trans. Knowl. Data Eng. 22, 1345–1359 (2010).
Neyshabur, B., Sedghi, H. & Zhang, C. What is being transferred in transfer learning? In 34th Conference on Neural Information Processing Systems (NeurIPS, 2020).
Raghu, M. et al. Transfusion: understanding transfer learning for medical imaging. In 33rd Conference on Neural Information Processing System (eds, Wallach, H. et al.) 3347–3357 (Curran Associates, 2019).
Hu, J. et al. Iterative transfer learning with neural network for clustering and cell type classification in single-cell RNA-seq analysis. Nat. Mach. Intell. 2, 607–618 (2020).
Sharifi-Noghabi, H., Zolotareva, O., Collins, C. C. & Ester, M. MOLI: multi-omics late integration with deep neural networks for drug response prediction. Bioinformatics 35, i501–i509 (2019).
Snow, O. et al. Interpretable Drug Response Prediction using a Knowledge-based Neural Network. In Proc. 27th ACM SIGKDD Conference on Knowledge Discovery & Data Mining (2021).
Kuenzi, B. M. et al. Predicting drug response and synergy using a deep learning model of human cancer cells. Cancer Cell 38, 672–684.e6 (2020).
Mourragui, S. et al. Predicting clinical drug response from model systems by non-linear subspace-based transfer learning. Preprint at https://www.biorxiv.org/content/10.1101/2020.06.29.177139v3 (2020).
Ma, J. et al. Few-shot learning creates predictive models of drug response that translate from high-throughput screens to individual patients. Nat. Cancer 2, 233–244 (2021).
Zhu, Y. et al. Ensemble transfer learning for the prediction of anti-cancer drug response. Sci. Rep. 10, 18040 (2020).
Salvadores, M., Fuster-Tormo, F. & Supek, F. Matching cell lines with cancer type and subtype of origin via mutational, epigenomic, and transcriptomic patterns. Sci. Adv. 6, aba1862 (2020).
Najgebauer, H. et al. CELLector: genomics-guided selection of cancer in vitro models. Cell Syst. 10, 424–432.e6 (2020).
Peres da Silva, R., Suphavilai, C. & Nagarajan, N. TUGDA: task uncertainty guided domain adaptation for robust generalization of cancer drug response prediction from in vitro to in vivo settings. Bioinformatics 37, i76–i83 (2021).
Warren, A. et al. Global computational alignment of tumor and cell line transcriptional profiles. Nat. Commun. 12, 22 (2021).
Gulrajani, I. & Lopez-Paz, D. In search of lost domain generalization. In International Conference on Learning Representations (2021).
Wang, J. et al. Generalizing to unseen domains: a survey on domain generalization. In Proc. Thirtieth International Joint Conference on Artificial Intelligence (2021).
Zhou, K., Liu, Z., Qiao, Y., Xiang, T. & Loy, C. C. Domain generalization: a survey. Preprint at https://arxiv.org/abs/2103.02503 (2021).
Zhang, H. et al. An empirical framework for domain generalization in clinical settings. In Proc. Conference on Health, Inference, and Learning (ACM, 2021); https://doi.org/10.1145/3450439.3451878
Zhao, S., Gong, M., Liu, T., Fu, H. & Tao, D. Domain generalization via entropy regularization. In 33rd Conference on Neural Information Processing Systems (NeurIPS, 2020).
Wang, Z., Loog, M. & van Gemert, J. Respecting domain relations: hypothesis invariance for domain generalization. In 2020 25th International Conference on Pattern Recognition 9756–9763 (ICPR, 2021).
Cancer Genome Atlas Research Network et al. The cancer genome atlas pan-cancer analysis project. Nat. Genet. 45, 1113–1120 (2013).
Schwartz, L. H. et al. RECIST 1.1—update and clarification: from the RECIST committee. Eur. J. Cancer 62, 132–137 (2016).
Hatzis, C. et al. A genomic predictor of response and survival following taxane-anthracycline chemotherapy for invasive breast cancer. JAMA 305, 1873–1881 (2011).
Ding, Z., Zu, S. & Gu, J. Evaluating the molecule-based prediction of clinical drug responses in cancer. Bioinformatics 32, 2891–2895 (2016).
Tarvainen, A. & Valpola, H. Mean teachers are better role models: weight-averaged consistency targets improve semi-supervised deep learning results. In 31st Conference on Neural Information Processing Systems (2017).
Yang, Y. & Xu, Z. Rethinking the value of labels for improving class-imbalanced learning. In Conference on Neural Information Processing Systems (2020).
Geeleher, P. et al. Discovering novel pharmacogenomic biomarkers by imputing drug response in cancer patients from large genomics studies. Genome Res. 27, 1743–1751 (2017).
Noghabi, H. S. et al. Drug sensitivity prediction from cell line-based pharmacogenomics data: guidelines for developing machine learning models. Briefings Bioinformatics https://doi.org/10.1093/bib/bbab294 (2021).
Renner, W., Langsenlehner, U., Krenn-Pilko, S., Eder, P. & Langsenlehner, T. BCL2 genotypes and prostate cancer survival. Strahlenther. Onkol. 193, 466–471 (2017).
Chaudhary, K. S., Abel, P. D. & Lalani, E. N. Role of the Bcl-2 gene family in prostate cancer progression and its implications for therapeutic intervention. Environ. Health Perspect. 107, 49–57 (1999).
Paraf, F., Gogusev, J., Chrétien, Y. & Droz, D. Expression of Bcl-2 oncoprotein in renal cell tumours. J. Pathol. 177, 247–252 (1995).
Bhat, K. M. R. & Setaluri, V. Microtubule-associated proteins as targets in cancer chemotherapy. Clin. Cancer Res. 13, 2849–2854 (2007).
He, Z., Liu, H., Moch, H. & Simon, H.-U. Machine learning with autophagy-related proteins for discriminating renal cell carcinoma subtypes. Sci. Rep. 10, 720 (2020).
Martin, S. K., Kamelgarn, M. & Kyprianou, N. Cytoskeleton targeting value in prostate cancer treatment. Am. J. Clin. Exp. Urol. 2, 15–26 (2014).
Kelly, R. S. et al. The role of tumor metabolism as a driver of prostate cancer progression and lethal disease: results from a nested case-control study. Cancer Metab. 4, 22 (2016).
Numakura, K. et al. Successful mammalian target of rapamycin inhibitor maintenance therapy following induction chemotherapy with gemcitabine and doxorubicin for metastatic sarcomatoid renal cell carcinoma. Oncol. Lett. 8, 464–466 (2014).
Pignon, J.-C. et al. Androgen receptor controls EGFR and ERBB2 gene expression at different levels in prostate cancer cell lines. Cancer Res. 69, 2941–2949 (2009).
Reid, A., Vidal, L., Shaw, H. & de Bono, J. Dual inhibition of ErbB1 (EGFR/HER1) and ErbB2 (HER2/neu). Eur. J. Cancer 43, 481–489 (2007).
Gordon, M. S. et al. Phase II study of Erlotinib in patients with locally advanced or metastatic papillary histology renal cell cancer: SWOG S0317. J. Clin. Oncol. 27, 5788–5793 (2009).
Chen, Y.-H. et al. No more discrimination: cross city adaptation of road scene segmenters. In Proc. IEEE International Conference on Computer Vision 1992–2001 (IEEE, 2017).
Costello, J. C. et al. A community effort to assess and improve drug sensitivity prediction algorithms. Nat. Biotechnol. 32, 1202–1212 (2014).
Jiang, Y., Rensi, S., Wang, S. & Altman, R. B. DrugOrchestra: jointly predicting drug response, targets, and side effects via deep multi-task learning. Preprint at https://www.biorxiv.org/content/10.1101/2020.11.17.385757v1 (2020).
Pozdeyev, N. et al. Integrating heterogeneous drug sensitivity data from cancer pharmacogenomic studies. Oncotarget 7, 51619–51625 (2016).
Xia F, et al. A cross-study analysis of drug response prediction in cancer cell lines. Brief. Bioinform. (2021).
Sharifi-Noghabi, H., Liu, Y., Erho, N. & Shrestha, R. Deep genomic signature for early metastasis prediction in prostate cancer. Preprint at https://www.biorxiv.org/content/10.1101/276055v2 (2019).
Torrente, A. et al. Identification of cancer related genes using a comprehensive map of human gene expression. PLoS ONE 11, e0157484 (2016).
Villicaña, C., Cruz, G. & Zurita, M. The basal transcription machinery as a target for cancer therapy. Cancer Cell Int. 14, 18 (2014).
Bailey, M. H. et al. Comprehensive characterization of cancer driver genes and mutations. Cell 174, 1034–1035 (2018).
Joshi, S. K. et al. ERBB2/HER2 mutations are transforming and therapeutically targetable in leukemia. Leukemia 34, 2798–2804 (2020).
Thomas, R. & Weihua, Z. Rethink of EGFR in cancer with its kinase independent function on board. Front. Oncol. 9, 800 (2019).
Nath, S. et al. The prognostic impact of epidermal growth factor receptor (EGFR) in patients with acute myeloid leukaemia. Indian J. Hematol. Blood Transfus. 36, 749–753 (2020).
Iqbal, N. & Iqbal, N. Human epidermal growth factor receptor 2 (HER2) in cancers: overexpression and therapeutic implications. Molecular Biol. Int. 2014, 1–9 (2014).
Goss, G. D. et al. Association of ERBB mutations with clinical outcomes of Afatinib- or Erlotinib-treated patients with lung squamous cell carcinoma: Secondary analysis of the LUX-lung 8 randomized clinical trial. JAMA Oncol. 4, 1189–1197 (2018).
Mammoliti, A. et al. Orchestrating and sharing large multimodal data for transparent and reproducible research. Nature Communications volume 12, Article number: 5797 (2021).
Smirnov, P. et al. PharmacoGx: an R package for analysis of large pharmacogenomic datasets. Bioinformatics 32, 1244–1246 (2016).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Erratum: near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 888 (2016).
Manica, M. et al. Toward explainable anticancer compound sensitivity prediction via multimodal attention-based convolutional encoders. Mol. Pharm. 16, 4797–4806 (2019).
Sun, B. & Saenko, K. Deep CORAL: correlation alignment for deep domain adaptation. In Computer Vision—ECCV 2016 Workshops 443–450 (Springer, 2016).
Sakellaropoulos, T. et al. A deep learning framework for predicting response to therapy in cancer. Cell Rep. 29, 3367–3373.e4 (2019).
Smirnov, P. et al. PharmacoDB: an integrative database for mining in vitro anticancer drug screening studies. Nucl. Acids Res. 46, D994–D1002 (2018).
Sarifi-Noghabi, H,. Harjandi, P. A., Zolotareva, O., Collins, C. C. & Ester, M. Velodrome: Out-of-Distribution Generalization from Labeled and Unlabeled Gene Expression Data for Drug Response Prediction (Zenodo, 2021); https://doi.org/10.5281/zenodo.4793442
Sharifi-Noghabi, H. Code Repository hosseinshn/Velodrome: DOI (v1.0.0) (Zenodo, 2021); https://doi.org/10.5281/zenodo.5164625
Acknowledgements
We would like to thank H. Asghari (Ocean Genomics) and S. Peng (Simon Fraser University) for their support. We also would like to thank the Vancouver Prostate Centre and Compute Canada (West Grid) for providing the computational resources for this research. This work was supported by a Discovery Grant from the National Science and Engineering Research Council of Canada (to M.E.), Canada Foundation for Innovation (33440 to C.C.C.), The Canadian Institutes of Health Research (PJT-153073 to C.C.C.), Terry Fox Foundation (201012TFF to C.C.C.) and The Terry Fox New Frontiers Program Project Grants (1062 to C.C.C.).
Author information
Authors and Affiliations
Contributions
H.S-.N. and M.E. conceived the study concept and design. H.S-.N. was responsible for the deep learning design, implementations and analysis. H.S-.N. and O.Z. performed data preprocessing, analysis and interpretation. H.S-.N. and P.A.H. performed the experiments. H.S-N., P.A.H. and O.Z. analysed and interpreted the results. C.C.C. and M.E. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Machine Intelligence thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1
The percentage of tissue types in CTRPv2 and GDSCv2 cell line datasets combined.
Supplementary information
Supplementary Information
Supplementary Tables 1–3.
Source data
Source Data Fig. 2
Results of prediction performance for cell lines, PDXs and patients.
Source Data Fig. 3
Results of multiple runs for cell lines, PDXs and patients (Fig. 3A sheets) and the ablation study (Fig. 3B sheet).
Source Data Fig. 4
Results of prediction performance compared with baseline correlations.
Source Data Extended Data Fig. 1
The percentage of tissue types in CTRPv2 and GDSCv2 cell line datasets combined.
Rights and permissions
About this article
Cite this article
Sharifi-Noghabi, H., Harjandi, P.A., Zolotareva, O. et al. Out-of-distribution generalization from labelled and unlabelled gene expression data for drug response prediction. Nat Mach Intell 3, 962–972 (2021). https://doi.org/10.1038/s42256-021-00408-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42256-021-00408-w
This article is cited by
-
A context-aware deconfounding autoencoder for robust prediction of personalized clinical drug response from cell-line compound screening
Nature Machine Intelligence (2022)