Abstract
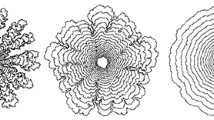
We develop two models for Myxobacteria swarming, a modified Lattice Gas Cellular Automata (LGCA) model and an off-lattice CA model. In the LGCA model each cell is represented by one node for the center of mass and an extended rod-shaped cell profile. Cells check the surrounding area and choose in which direction to move based on the local interactions. Using this model, we obtained a density vs. expansion rate curve with the shape similar to the experimental curve for the wild type Myxobacteria. In the off-lattice model, each cell is represented by a string of nodes. Cells can bend and move freely in the two-dimensional space. We use a phenomenological algorithm to determine the moving direction of cells guided by slime trail; the model allows for cell bending and alignment during collisions. In the swarming simulations for A+S- Myxobacteria, we demonstrate the formation of peninsula structures, in agreement with experiments.
Preview
Unable to display preview. Download preview PDF.
Similar content being viewed by others
References
Kaiser, D., Crosby, C.: Cell movement and its coordination in swarms of Myxococcus xanthus. Cell Motility 3, 227–245 (1983)
Kaiser, D.: Coupling Cell Movement to Multicellular Development in Myxobacteria. Nature Reviews Microbiology 1, 45–54 (2003)
Sozinova, O., Jiang, Y., Kaiser, D., Alber, M.: A three-dimensional model of myxobacterial aggregation by contact-mediated interactions. Proc. Natl. Acad. Sci. U.S.A. 102, 11308–11312 (2005)
Hodgkin, J., Kaiser, D.: Genetics of gliding motility in M. xanthus (Myxobacterales): two gene systems control movement. Mol. Gen. Genet. 171, 177–191 (1979)
Wolgemuth, C., Hoiczyk, E., Kaiser, D., Oster, G.: How Myxobacteria Glide. Curr. Biol. 12, 369–377 (2002)
Kaiser, D., Yu, R.: Reversing cell polarity: evidence and hypothesis. Cur. Opin. Microbiol. 8, 216–221 (2005)
Kaiser, D.: Signaling in Myxobacteria. Annu. Rev. Microbiol. 58, 75–98 (2004)
Burchard, R.P.: Trail following by gliding bacteria. J. Bacteriol. 152, 495–501 (1982)
Gallegos, A., Mazzag, B., Mogilner, A.: Mathematical analysis of the swarming behavior of myxobacteria. Bulletin of Mathematical Biology (in print)
Hardy, J., Pazzis, O., Pomeau, Y.: Molecular dynamics of a classical lattice gas: Transport properties and time correlation functions. Phys. Rev. A 13, 1949–1961 (1976)
Palsson, E., Othmer, H.G.: A model for individual and collective cell movement in Dictyostelium discoideum. Proc. Natl. Acad. Sci. USA. 97, 10448–10453 (2000)
Spormann, A.M., Kaiser, D.: Gliding movements in Myxococcus xanthus. J. Bacteriol. 177, 5846–5852 (1995)
Newman, T.J.: Modeling Multicellular systems using subcellular elements. Mathematical Biosciences and Engineering 2, 611–622 (2005)
Newman, M.E.J., Barkema, G.T.: Monte Carlo Methods in Statistical Physics. Clarendon Press, Oxford (1999)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2006 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Wu, Y., Chen, N., Rissler, M., Jiang, Y., Kaiser, D., Alber, M. (2006). CA Models of Myxobacteria Swarming. In: El Yacoubi, S., Chopard, B., Bandini, S. (eds) Cellular Automata. ACRI 2006. Lecture Notes in Computer Science, vol 4173. Springer, Berlin, Heidelberg. https://doi.org/10.1007/11861201_24
Download citation
DOI: https://doi.org/10.1007/11861201_24
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-540-40929-8
Online ISBN: 978-3-540-40932-8
eBook Packages: Computer ScienceComputer Science (R0)