Abstract
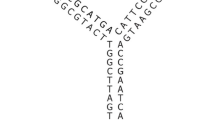
DNA tile nanostructures have lately attracted a lot of attention as a new calculation technique and material on the nanometer scale. In forming DNA tiles, sequences need to bond in tile conformation. Conventional work can design sequences using overlapping subsequence. In this paper, we design tile sequences based on free energy. As a result of optimization, we show that we can design tile sequences as stable as conventional tiles. Moreover, we illustrate that the tile designed by the proposed method is perhaps more stable than conventional one. This method will be useful to design many tiles when forming large scale and complex DNA nanostructures.
Preview
Unable to display preview. Download preview PDF.
Similar content being viewed by others
References
Winfree, E., Liu, F., Wenzler, L.A., Seeman, N.C.: Design and self-assembly of two-dimensional DNA crystals. Nature 394, 539–544 (1998)
Winfree, E.: DNA Computing by Self-assencly. NAE’s TheBredge 33(4), 31–38 (2003)
Yan, H., Feng, L., LaBean, T.H., Reif, J.H.: Parallel Molecular Computations of Pairwise Exclusive-Or (XOR) Using DNA ”String Tile” Self-Assembly. J. Am. Chem. Soc. 125(47), 14246–14247 (2003)
Yan, H., Park, S.H., Finkelstein, G., Reif, J.H., LaBean, T.H.: DNA-Templated Self-Assembly of Protein Arrays and Highly Conductive Nanowires. Science 301, 1882–1884 (2003)
Seeman, N.C.: De Nove Design of Sequence for Nucleic Acid Structual Engineering. Journal of Biomolecular Structure & Dynamics, ISSNO 8, 739–1102 (1990)
Tanaka, F., Kameda, A., Yamamoto, M., Ohuchi, A.: Design of nucleic acid sequences for DNA computing based on a thermodynamic approach. Nucleic Acids Research 33(3), 903–911 (2005)
Iimura, N., Yamamoto, M., Tanaka, F., Kameda, A., Ohuchi, A.: Stability evaluation method of DNA tile Structure. In: Proceedings of the 11th International Symposium on Artificial Life and Robotics (AROB 11th 2006), CD-ROM (2006)
Suppoting Online Material, www.sciencemag.org/cgi/content/full/301/5641/1882/DC1
Zuker, A.M., Mathews, B.D.H., Turner, C.D.H.: Algorithms And Thermodynamics For RNA Secondary Structure Prediction: A Practical Guide. In: Barciszewski, J., Clark, B.F.C. (eds.) RNA Biochemistryand Biotechnology. NATO ASI Series, Kluwer Academic Publisers, Dordrecht (1999)
Zuker, M.: Mfold web server for nulceic acid folding and hybridization prediction. Nucleic Acids Reserch 31(13), 3406–3415 (2003)
Andronescu, M., Aguirre-Hernandez, R., Condon, A., Hoos, H.H.: RNA soft: a suite of RNA secondary structure prediction and design software tools. Nucleic Acids Research 31(13), 3416–3422 (2003)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2006 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Iimura, N., Yamamoto, M., Tanaka, F., Kameda, A., Ohuchi, A. (2006). Sequence Design for Stable DNA Tiles. In: Mao, C., Yokomori, T. (eds) DNA Computing. DNA 2006. Lecture Notes in Computer Science, vol 4287. Springer, Berlin, Heidelberg. https://doi.org/10.1007/11925903_13
Download citation
DOI: https://doi.org/10.1007/11925903_13
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-540-49024-1
Online ISBN: 978-3-540-68423-7
eBook Packages: Computer ScienceComputer Science (R0)