Abstract
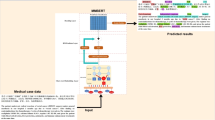
Identifying named entities from unstructured biomedical text is an important part of information extraction. The irrelevant words in long biomedical sentences and the complex composition of the entity make LSTM used in the general domain less effective. We find that emphasizing the local connection between words in a biomedical entity can improve performance. Based on the above observation, this paper proposes two novel neural network architectures combining bidirectional LSTM and CNN. In the first architecture S-CLSTM, a CNN structure is built on the top of bidirectional LSTM to keep both long dependencies in a sentence and local connection between words. The second architecture P-CLSTM combines bidirectional LSTM and CNN in parallel with the weighted loss to take advantage of the complementary features of two networks. Experimental results indicate that our architectures achieve significant improvements compared with baselines and other state-of-the-art approaches.
This paper is supported by the National Key Research and Development Program of China (Grant No. 2016YFB1001102), the National Natural Science Foundation of China (Grant Nos. 61876080, 61502227), the Fundamental Research Funds for the Central Universities No.020214380040, the Collaborative Innovation Center of Novel Software Technology and Industrialization at Nanjing University.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
Notes
- 1.
The character representation is generated by BiLSTM, with 100 hidden states.
- 2.
References
Zhu, Q., Li, X., Conesa, A., Pereira, C.: GRAM-CNN: a deep learning approach with local context for named entity recognition in biomedical text. Bioinformatics 34(9), 1547–1554 (2017)
Wu, Z., Dai, X.-Y., Yin, C., Huang, S., Chen, J.: Improving review representations with user attention and product attention for sentiment classification. In: Proceedings of the Thirty-Second AAAI Conference on Artificial Intelligence, New Orleans, Louisiana, USA, 2–7 February 2018
Crichton, G., Pyysalo, S., Chiu, B., Korhonen, A.: A neural network multi-task learning approach to biomedical named entity recognition. BMC Bioinform. 18(1), 368 (2017)
Ma, X., Hovy, E.: End-to-end sequence labeling via bi-directional LSTM-CNNS-CRF. In: Proceedings of the 54th Annual Meeting of the Association for Computational Linguistics. ACL, Berlin, Germany, Volume 1: Long Papers, 7–12 August 2016
Habibi, M., Weber, L., Neves, M., Wiegandt, D.L., Leser, U.: Deep learning with word embeddings improves biomedical named entity recognition. Bioinformatics 33(14), i37–i48 (2017)
Rei, M., Crichton, G.K.O., Pyysalo, S.: Attending to characters in neural sequence labeling models. In: COLING 2016, 26th International Conference on Computational Linguistics, Proceedings of the Conference: Technical Papers, Osaka, Japan, pp. 309–318, 11–16 December 2016
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this paper
Cite this paper
Lu, Q., Xu, Y., Yang, R., Li, N., Wang, C. (2019). Serial and Parallel Recurrent Convolutional Neural Networks for Biomedical Named Entity Recognition. In: Li, G., Yang, J., Gama, J., Natwichai, J., Tong, Y. (eds) Database Systems for Advanced Applications. DASFAA 2019. Lecture Notes in Computer Science(), vol 11448. Springer, Cham. https://doi.org/10.1007/978-3-030-18590-9_62
Download citation
DOI: https://doi.org/10.1007/978-3-030-18590-9_62
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-18589-3
Online ISBN: 978-3-030-18590-9
eBook Packages: Computer ScienceComputer Science (R0)