Abstract
A method which allows us to find a polynomial model of an unknown function from a set of tuples with n independent variables and m tuples is presented. A plausible model of an explicit approximation is found. The number of monomials of the model is determined by a previously trained neural network (NNt). The degrees of the monomials are bounded from the Universal Approximation Theorem yielding a reduced polynomial form. The coefficients of the model are found by exploring the space of possible monomials with a genetic algorithm (GA). The polynomial defined by every training pair is approximated by the Ascent Algorithm which yields the coefficients of best fit with the L∞ norm. The L2 error corresponding to these coefficients is calculated and becomes the fitness function of the GA. A detailed example for a well known experimental data set is presented. It is shown that the model derived from the method yields a classification error of <7% for the cross-validation data set. A detailed description of the NNt is included. The approximation to 46 datasets tackled with our method is discussed. In all these cases the approximation accuracy was better than 94%. The tool described herein yields general models with explanatory characteristics not found in other methods.
You have full access to this open access chapter, Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
In this paper we address the problem of obtaining a general algebraic model for a set of given data pairs \( (\vec{x},y) \). By “model” we mean a function \( F(\vec{x}) \) derived from a set of n independent variables and m given data pairs assumed to stem from an unknown function \( y = f(\vec{x}) \), such that the error \( \varepsilon = \left\| {f(\vec{x}) - F(\vec{x})} \right\|_{2} \) is minimized.
In a previous work [1] a methodology is proposed to find a multivariate polynomial model to fit a dataset in which, instead of generating the full range of terms in (1), generates the combination of a preselected number of them. This is accomplished by the ensemble of a genetic algorithm and a regression algorithm, where the individuals of the genetic algorithm represent a set of monomials with different combination of variables and their exponents. The regression algorithm computes the coefficients of the terms in such way that the minimax error norm (or L∞) is minimized. The current work is based on this methodology. We complement it by finding a neural network (NNt) which yields the minimum needed number of terms.
The rest of the paper is organized as follows. In Sect. 2 we describe
-
1.
How to select the model’s form
-
2.
How to approximate, from an arbitrary training set of pairs \( (\vec{x},y) \), a polynomial model.
-
3.
How to avoid the exponential number of terms as the number of variables and the maximum degrees considered increases
-
4.
How to determine the degrees d1, …, dn of (1) needed for \( F(\vec{x}) \) to be a universal approximator.
In Sect. 3 we focus on the problem of how to determine the minimum number of terms in \( F(\vec{x}) \). In Sect. 4 we present the application of the methodology to a case of study. In Sect. 5 we present our conclusions.
2 Previous Results
In order to obtain a comprehensive methodology leading to the goal of finding closed mathematical (polynomial) models from experimental data we rely, at the offset, on previous works. The reader interested in the details is encouraged to dwell on the references.
2.1 Selecting the Model’s Form
A basic issue to be considered is how to select the form of \( F(\vec{x}) \). One possible approach to determine such form is suggested by the Weierstrass approximation theorem (WAT), which states that every continuous function defined on a closed interval [a, b] can be uniformly approximated as closely as desired by a polynomial function [2].
From the WAT we assume that there exists a full polynomial approximation
We consider all possible linear combinations of the products of the independent variables x1, …, xn raised to a given degree d1, …, dn. The set of allowed degrees d1, …, dn will be denoted with D, the set of coefficients \( c_{{i_{1} i_{2} \ldots i_{n} }} \) will be denoted with C, the number of terms of the polynomial will be denoted by T. The purpose of the algorithm we implemented is to express \( F(\vec{x}) \) as a more compact polynomial. By “algebraic model”, therefore, we mean a function of the form
2.2 Approximation of a Polynomial Model
The number of monomials in (1) is M = (d1 + 1)…(dn + 1). Consider, for example, the well known (number of web hits = 1,020,829) classification problem in [3]. The data pairs in it (n = 13 and m = 178) may be accommodated in a \( 178 \times 14 \) data base. This is a classification problem where the values of the tuples determine one of three classes. For convenience we shall assume that the number of the class (the dependent variable y) is denoted with 0, 1 and 2 and, furthermore, that it is placed on the last column of the matrix. Even assuming the conservative d1 = … = d13 = 2 we find that M = 413 = 67,108,864. The exponential growth of M has been referred to as “the curse of dimensionality” [4]. With the development of NNs this problem has been successfully circumvented [5] albeit without yielding an explicit model, i.e. one in which the relation between the independent variables is exposed. NNs are frequently cited as suffering from the “black box” disadvantage. The main motivation in this work is to solve this kind of problems without losing the explanatory advantages of an algebraic model. To achieve this we apply a method where the model of (1) is replaced by the one in (3), where the monomials (or “terms”) in (1) are selectively retained/discarded.
where
The basic idea is to set T and D which, immediately, allows us to avoid the curse of dimensionality. The way in which T and D are determined is discussed in what follows.
2.3 Determining the Coefficients of the Monomials
Once D and T are determined we may determine the \( \mu_{{i_{1} i_{2} \ldots i_{n} }} \) indirectly by applying the genetic algorithm (EGA) described in [6], as follows. The individuals of the population of the EGA represent the combination of powers \( i_{1} , \ldots ,i_{n} \) corresponding to each of the T monomials in (2) for which the μi1i2…in are retained. This is exemplified in Fig. 1 where n = 3, 0 ≤ di ≤ 9 and T = 5. For simplicity, the coefficients \( c_{{i_{1} i_{2} \ldots i_{n} }} \) in (2) have been replaced with C1, …, C5.
Every individual of the EGA represents a polynomial consisting of T monomials. From them we find a data base of size \( T \times m \) where every one of its tuples is a map from the original \( \vec{x} \) into the monomials proposed by the EGA plus the value of the dependent variable, as illustrated in Fig. 2. The data base from which this figure is originated consists of 50 \( (\vec{x},y) \) pairs, in this case of the form \( (x_{1} ,x_{2} ,x_{3} ,F) \).
From the original data a mapping analogous to the one illustrated in Fig. 2 is obtained.
To find C we implemented an approximation algorithm known as the Ascent Algorithm (AA) [7]. Cij is the coefficient induced by the i-th individual during the j-th generation. Using the AA, we find Cij and its associated fit error εij. After the evolutionary process the coefficients of the best individual (corresponding to εij) will yield \( F(\vec{x}) \). The \( \mu_{{i_{1} i_{2} \ldots i_{n} }} \) are never explicitly obtained. The C resulting from the EGA tacitly determines the T retained monomials whose coefficients minimize \( \varepsilon \).
2.4 Degree of the Terms in the Polynomial
In [1] it was shown that, from the Universal Approximation Theorem [8], any unknown function \( f(\vec{x}) \) may be approximated with a polynomial of the form of (5)
if
where \( \sum {dj} \in L \) means that the maximum degree (maxdeg) of any term (the summation of the degrees associated to each of the independent variables) must belong to set L = {1, 3, 5, 7, 9, 11, 15, 21, 25, 33, 35, 37, 45, 49, 55, 63, 77, 81, 99, 121}.
The way in which L was arrived at may be found in [1]. Notice, however, that even if maxdeg of the i-th monomial is odd, the powers of the variables in it may take any value leading to one of those in L.
The algebraic model of (1) is thusly simplified and the EGA can be made to include in its population only individuals whose coefficients comply with (6). To illustrate this fact, the data of Fig. 1 [consisting of 50 pairs \( (\vec{x},y) \)] was approximated with the EGA now considering Eq. (5). The resulting coefficients are as shown in Fig. 3. In an “indexed coefficient” like Ci1, …, in, the indices denote the degrees associated to the i-th variable. The degree of the term is shown on the first column.
Approximation coefficients for the data in Fig. 2
3 Minimum Number of Terms
Once we have a method to determine, via EGA, the most adequate coefficient for every term in (5) it only remains to specify the most adequate value for T. The solution to this problem completes the specification of the method. An attempt to solve this problem is to appeal to statistical criteria.
The first step was to set lower and upper practical limits on the number of terms. We reasoned that we were not interested in very small values for T. Very low values are hardly prone to yield good models. Therefore, we set a lower value of T = 3. On the other side of the spectrum we decided to focus on T ≤ 13. Higher values are seldom of practical interest since large sets of coefficients are cumbersome and difficult to analyze. At the end of the day algebraic models allow us to search for patterns in the relations which are exposed by the monomials and the possible relations embedded. Very large Ts will make this search too complex in general.
Therefore, we collected 46 datasets from the University of California Machine Learning dataset repository [9] and the Knowledge Extraction Evolutionary Learning dataset repository [10]. To begin with, 32 of these datasets were chosen and were subjected to the procedure outlined above. For every \( {\mathbf{T}} \in [3,13] \) a polynomial was found and the number of terms corresponding to the best fit was recorded. A total of \( 352\;(11 \times 32) \) polynomials, therefore, were calculated. Table 1 shows the values of best T from the 32 selected data sets. For every one of the 32 datasets 11 polynomials (T = 3, …, 13) were found. From these the lower bound of the number of terms was determined. The best values of T were then used to train NNt. The polynomials were calculated as a function of the following variables: (a) Number of attributes, (b) Number tuples, (c) Optimally encoded size, (d) Compression ratio [the quotient of the (dataset’s original size)/(optimally compressed size)]. These attributes were selected from a much larger original subset of such attributes. The ones we selected were shown to be the more relevant from a Principal Components Analysis.
The remaining 14 datasets were used as cross validation data for the trained NN model. The corresponding results are shown in Table 2. From the results a statistically highly significant lower bound on T may be inferred. NNt may be fully specified from: (a) Its architecture, (b) The associated weights, (c) The definition of the activation function.
We can calculate the average RMS error and the standard deviation. From \( P(\mu T - k\sigma T \le T \le \mu T + k\sigma T) > \mathit{1} - \mathit{1}/k^{\mathit{2}} \) (Chebyshev’s theorem [13]). Making k = 3, the probability of finding the smallest expected value of T above the average value at k = −3 standard deviations is ≈88%. Thereafter the mathematical model for the minimum number of terms as a function of the RMS error within the indicated limits may be calculated. We trained a NN from these data yielding NNt. The architecture of NNt is shown in Fig. 4.
NNt was trained using the backpropagation algorithm. To improve convergence the learning rate is set in the hidden and output layers. The learning rate was tested in the hidden layer with the values .1, .2, .3 and .25; the learning rate in the output layer was tested for the values .1, .01 and .001.
Activation Function.
Every neuron is a perceptron whose activation function is \( 1/(1 + e^{ - x} ) \).
Momentum.
Amomentum term was added, which allows the adjustments on the weights to depend also on the previous changes. If the momentum is equal to 0, the change depends only on the error gradient, while a value of 1 means that the change will only depend on the last change. The value of the momentum in the hidden and output layers was tested for values 0.3, 0.5 and .8. The weights were initialized with uniform random numbers in the range [−0.1, 0.1] ∈ R. The learning strategy used was stochastic in which each input creates a weight adjustment. The NN was trained for 10,000 epochs with a test every 100 epochs. NNt was validated vs. the remaining 14 data sets.
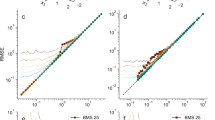
The above information was fed to NNt to get T. In Table 3 we see the estimated T which are expected to be near the optimum fit. RMS(T-i) denotes the RMS error when NNt yields i for the value of T. The actual best value of T was always within i ± 2 of the T suggested by NNt.
The weights of NNt are shown in Fig. 5. Every neuron is, as stated, a perceptron whose activation function is \( 1/(1 + e^{ - x} ) \). Because of this the input values were scaled into [0, 1). Given the values of the 4 input attributes a simple spreadsheet may easily calculate the suggested number of terms. An example is shown in Fig. 6.
-
Since the values must be scaled the upper and lower values of the variables are needed. These are shown in the lines titled “Max” and “Min”. The corresponding scaled values are shown in the line “Scaled_01”.
-
“Compressed size” refers to the size (bytes) of the original dataset after it is compressed using the PPMZ Algorithm [11]. This yields a practical approximation to the information contained in the datasetFootnote 1.
4 Case of Study
To illustrate the process we take the data base from the U.S. Census [14]. From the web page we get the data illustrated in Fig. 7.
A partial view of the data set is shown in Fig. 8. This data is both numerical and categorical (i.e. non-numerical). The numerical attributes are scaled into [0, 1) and, subsequently, the non-numerical attributes were encoded into [0, 1) using the CESAMO algorithm [15]. The original DB is illustrated in Fig. 8. The corresponding encoded DB is illustrated in Fig. 9. This DB was further divided into a training data (TRN) set and a test data set (TST).
TRN was used to train (find) a polynomial and cross validated with TST. The number of terms T of the polynomial, as illustrated in Fig. 6, was originally determined to be 9. The actual resulting polynomial is illustrated in Fig. 10.
EpsiTh denotes the worst case approximation error in TST. In every column starting with the second row we find (a) The degree of the monomial, (b) The powers of the independent variables in every monomial, (c) The coefficient associated to the monomial. The value of T is 7 because the EGA may yield monomials of identical monomials are odd (1, 3, 5, 7). The genetic algorithm was allowed to choose from any combination of the degrees in set L. It may be seen that it picked terms whose highest power for any one variable is 3 and the corresponding highest degree is 7.
degrees and these are merged in the end. As specified, the degrees of every one of the are odd (1, 3 ,5, 7). The genetic algorithm was allowed to choose from any combination of the degrees in set L. It may be seen that it picked terms whose highest power for any one variable is 3 and the corresponding highest degree is 7.
When the polynomial was evaluated on the test data we determined that, of the 235 tuples of TST, 231 were correctly classified for an effectiveness ratio of 93.83%.
5 Conclusions
A method allowing us to find a polynomial model of an unknown function from a set of experimental tuples was explored. The allowed powers of the monomials in such polynomial were derived from the Universal Approximation Theorem. The coefficients of the monomials were gotten by exploring the space of such monomials with the EGA. Every individual’s represented polynomial is found by the Ascent Algorithm. The AA yields the coefficients of best fit with the L∞ norm. From these coefficients the L2 error is calculated and becomes the fitness function of the GA. In it, every individual represents a plausible polynomial in which the number of terms (T) is determined by a trained neural network (NNt) with the best value of T (3 ≤ T ≤ 13) for 32 representative real world data sets, i.e. \( 352\;(11 \times 32) \) polynomials were calculated. NNt yields T when the number of tuples, the number of attributes, the size of the compressed data set and the compression ratio are input. The method was used to solve a case study where the evaluated polynomial over the test data set yielded an effectiveness ratio of 93.83%. This illustrates a method which yields the number of terms and the coefficients and degree of every one of the independent variables for each term purely by machine learning.
Notes
- 1.
The true amount of information in a data set is exactly expressible by Kolmogorov’s Complexity (KC) which corresponds to the most compact representation of the set under scrutiny. Since KC is known to be incomputable [12] we have chosen the PPM (Prediction by Partial Matching) algorithm as our compression standard; a good practical approximation to KC.
References
Kuri-Morales, A., Cartas-Ayala, A.: Polynomial multivariate approximation with genetic algorithms. In: Sokolova, M., van Beek, P. (eds.) AI 2014. LNCS (LNAI), vol. 8436, pp. 307–312. Springer, Cham (2014). https://doi.org/10.1007/978-3-319-06483-3_30
Cotter, N.E.: The Stone-Weierstrass theorem and its application to neural networks. IEEE Trans. Neural Netw. 1(4), 290–295 (1990)
https://archive.ics.uci.edu/ml/datasets/wine. Wine Data Set: Accessed 27 Nov 2018
Haykin, S.: Neural Networks: A Comprehensive Foundation. Prentice Hall PTR, Upper Saddle River (1994)
Bastian, M., Heymann, S., Jacomy, M.: Gephi: an open source software for exploring and manipulating networks. In: ICWSM, vol. 8, no. 2009, pp. 361–362 (2009)
Kuri-Morales, A.F., Aldana-Bobadilla, E., López-Peña, I.: The best genetic algorithm II. In: Castro, F., Gelbukh, A., González, M. (eds.) MICAI 2013. LNCS (LNAI), vol. 8266, pp. 16–29. Springer, Heidelberg (2013). https://doi.org/10.1007/978-3-642-45111-9_2
Cheney, W., Kincaid, D.: Linear Algebra: Theory and Applications. The Australian Mathematical Society, p. 110 (2009)
Cybenko, G.: Approximation by superpositions of a sigmoidal function. Math. Control Signals Syst. 2(4), 303–314 (1989)
Dheeru, D., Karra Taniskidou, E.: UCI machine learning repository (2017). http://archive.ics.uci.edu/ml
Alcalá-Fdez, J., et al.: Keel data-mining software tool: data set repository, integration of algorithms and experimental analysis framework. J. Multiple-Valued Logic Soft Comput. 17 (2011)
Moffat, A.: Implementing the PPM data compression scheme. IEEE Trans. Commun. 38(11), 1917–1921 (1990)
Vitányi, P.M.B., Li, M.: An Introduction to Kolmogorov Complexity and Its Applications. TCS, vol. 34. Springer, New York (2008). https://doi.org/10.1007/978-0-387-49820-1
Saw, J.G., Yang, M.C.K., Mo, T.C.: Chebyshev inequality with estimated mean and variance. Am. Stat. 38(2), 130–132 (1984)
Kohavi, R., Becker, B.: Data Mining and Visualization. Silicon Graphics. https://archive.ics.uci.edu/ml/datasets/Census+Income
Kuri-Morales, A.: Transforming mixed data bases for machine learning: a case study. In: Batyrshin, I., Martínez-Villaseñor, M., Ponce Espinosa, H. (eds.) MICAI 2018. LNCS (LNAI), vol. 11288, pp. 157–170. Springer, Cham (2018). https://doi.org/10.1007/978-3-030-04491-6_12
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this paper
Cite this paper
Kuri-Morales, A. (2019). A Comprehensive Methodology to Find Closed Mathematical Models from Experimental Data. In: Carrasco-Ochoa, J., Martínez-Trinidad, J., Olvera-López, J., Salas, J. (eds) Pattern Recognition. MCPR 2019. Lecture Notes in Computer Science(), vol 11524. Springer, Cham. https://doi.org/10.1007/978-3-030-21077-9_3
Download citation
DOI: https://doi.org/10.1007/978-3-030-21077-9_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-21076-2
Online ISBN: 978-3-030-21077-9
eBook Packages: Computer ScienceComputer Science (R0)