Abstract
One of the most prevailing types of lung cancer is non-small cell lung cancer (NSCLC). Differential diagnosis of NSCLC into adenocarcinoma (ADC) and squamous cell carcinoma (SCC) is important because of prognosis. Histological images are taken from a database consisting of 72 lung tissue samples collected indigenously with a core needle biopsy. In this work, a novel method has been developed where the features of ADC and SCC for a histological image are taken from various statistical and mathematical models implemented on the coefficients of the wavelet transform of an image. The method provides a precision of 95.1% and 96.2% in classifying malignant and non-malignant tissue type respectively. This methodology of classifying ADC and SCC without coding clinical diagnostic features into the system is a necessary step forward towards an autonomous decision system.
You have full access to this open access chapter, Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
The prevalence of Lung cancer in India can be observed by the fact that for the year 2018 a total of 67,795 new cases were registered for both the sexes and 63,475 deaths related to lung cancer were reported [1]. Non-small cell lung cancer (NSCLC) accounts for more than 80% of lung cancer spectrum. NSCLC can be further subtyped into two major types, viz. adenocarcinoma (ADC) and Squamous Cell Carcinoma (SCC).
It is imperative for differentiating the NSCLC into subtypes as the prognosis of adenocarcinoma (ADC) is found out to be low [2] as compared to the other subtype. Also, for NSCLC, disease management of different subtype is different. SCC outcome is worst [3] among the two NSCLC subtypes. A tissue biopsy can reveal whether the nodule is malignant or benign as well as the subtype if it is malignant. Tissue samples can be extracted by an invasive surgical procedure, core needle biopsy or fine needle aspiration biopsy. Needle biopsy is often favored over surgical biopsy [4]. The classification of lung tissue into ADC or SCC is not straightforward since the architecture that distinguishes them are complex and depends on the grade of the tumour [5].
The histopathological images are captured in RGB colour space. It is found in the literature that converting it into other colour space can improve the result. RGB to L*a*b* colour space is done for breast [6], lung [7] and prostate [8]. HSV (Hue, Saturation and Value), for head and neck cancer detection [9] and lung tissue type classification [7]. However, there are other colour spaces like YCbCr, which is used for prostate [10] and CMY for breast cancer [11].
The method described in this work is based on texture characterization of the lung tissue. Texture characterization of tissue can confirm the malignancy of a tumour [12] as filter banks are used for detecting breast cancer [11] and grading of prostate cancer [13]. The fractal dimension, along with the variance of the wavelet coefficient is implemented for prostate cancer grading [10]. Further, the wavelet coefficient as one of the feature to a support vector machine (SVM) for classification of prostate lesions [14] is implemented. Wavelets have been used to analyse texture effectively as they provide multiple scale partition of the image spectrum [15]. We have selected Continuous Wavelet Transform (CWT) as opposed to discrete wavelets as CWT gives a high degree of frequency selectivity [16] of a texture.
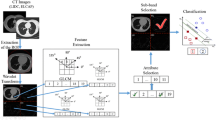
To the best of our knowledge, classification of ADC and SCC is done only using Raman scattering microscopy where they use domain specific clinical diagnosis knowledge as feature sets [17]. As the classification of these two subtypes is very complex, coding the domain specific rules into an algorithm like morphology feature may not always produce higher classification rate [17]. This paper is focused on classifying the two subtypes of NSCLC by quantitatively extracting the feature from each subtype automatically. Two dimensional Marr and isotropic Morlet wavelet are used to transform the image into wavelet domain so as to characterize the texture of the image. The wavelet coefficients are modelled with Generalised Gaussian Distribution (GGD). SVM as a classifier, is used, the features for SVM are selected through Recursive Feature Elimination (RFE) [18] method.
2 Methods and Materials
2.1 Data Set
The slides containing the lung tissue are collected from an NABL (National Accreditation Board for Testing and Calibration, India) laboratory. The slide containing lung tissue were prepared from core needle biopsy samples, these samples are stained with Hematoxylin and Eosin (H&E) and are sectioned 5 μm thickness. The images are captured using Leica ICC50 HD digital microscope, digitisation is done at magnification of 20x which forms image pixel resolution of 0.32 μm, the images are of 2048 × 1536 resolution and each pixel is of 24 bit to incorporate 3 channels i.e. Red (R), Green (G) and Blue (B) of 8 bit each. The histopathological images of the lung contain various cellular-based tissue types, like but not limited to: the tumour, red blood cells, fibrosis, necrosis, carbon particles, normal cells.
A surgical pathologist delineates the malignant tumour portion on the digital image, this Region of Interest (ROI) is stored as a binary mask. The subtype of the NSCLC is also mentioned and stored as a label for the binary mask, the rest of the portion of the image containing tissue structures are labelled as Unspecified Lung Region (ULR). As the aim of the study is to classify subclasses of NSCLC from the background, i.e. the portion of the image not containing any lung tissue structure is also labelled as ULR. The ROI and the ULR are segmented into overlapping blocks of size 256 × 256, it is empirically found out that this size gives the highest level of classification for both the classifiers. The number of blocks for ROI and ULR is almost equal to avoid biasness in classification [19]. A visual inspection is done on all the images and only focused images are taken (due to human error unfocused images are sometimes produced). In total slides from 72 core needle biopsy are used of which 34 are of SCC and 38 are of ADC type. Table 1 illustrate the data set.
2.2 Colour Space Transform and Normalization
In H&E stain the nucleus of the cell is stained blue whereas cytoplasm and extracellular materials are stained with varying degree of the colour pink. The colour appearance of the tissue in light microscopy varies due to a wide range of factors, the sensor type of the digital camera, H&E reagents from different manufacturers or from different batches, the concentration of the stain, time for which stains are applied, and many more factors are there. If most of the procedure of staining is standardised still the colour fades with time. Normalization of colour is a necessary pre-processing for histopathological images, in this work we have selected a method that nonlinearly maps a source image (an image that needs to be normalized) to a target image (an image that is used to train the system) [20]. This method shows stable representations as it is not sensitive to the imaging condition and digital sensor used.
The captured digital images of lung histology are in RGB colour space. The 3 colour channels of the RGB colour space are not independent and change in one channel changes other channels. To avoid such pitfall of RGB colour space and to closely resemble the colour perception of human L*a*b* colour space is chosen. The L* channel corresponds to luminance, a* and b* channel represent the variance of red to green and yellow to blue respectively, a* and b* together define the chrominance. Figure 1, shows the representation of different channels. Only two levels of headings should be numbered. Lower level headings remain unnumbered; they are formatted as run-in headings.
2.3 Wavelet Coefficients Modelling and Similarity Measurement
Marr wavelet is a real, rotation invariant wavelet. In some literature, it is also known as Mexican hat wavelet. A 2D Marr wavelet is defined as (see Fig. 2(a) for representation):
Marr wavelet was selected for its good localization feature and also its affinity towards representing the nucleus in an effective way in the transform domain as evident from Fig. 2(c) and (d). As nucleus features are essential and they are available only in the L* and b* colour channels since L* is for luminance and nucleus are coloured blue/purple with H&E stain which is darker than the pale pink/red the colour of cytoplasm.
The b* channel record changes from bluish to yellowish colour component thus nucleus are available in this channel (as evident from Fig. 1). Marr wavelet is used for these two colour channels.
Isotropic Morlet wavelet is a complex-valued wavelet. A simple Morlet wavelet can be a plain wave modulated by a Gaussian envelope with a well-localized frequency domain with power only near its fundamental frequency. An isotropic wavelet is given by [21]:
where \( \omega_{0} = \left( {0,\omega_{0} } \right) \) is a wave vector with \( \omega_{0} > 5.5 \). Phase information is important for texture characterisation since Morlet wavelet are complex valued we can compute the phase as well as magnitude information.
This wavelet is used on the a* channel, which codes non nucleus information. Since a* is for change of colour information from reddish to greenish and most non nucleus material are pale pink and pale red in appearance (see Fig. 1).
Let \( {\text{I}}_{h} \) be an image, the 2D CWT of the image is given by:
where \( {\mathcal{C}}_{\psi } \left( {a,x^{\prime},y^{\prime}} \right) \) is the wavelet coefficient at location \( x^{\prime},y^{\prime} \) and having scale \( \varvec{a} \) (\( a > 0 \)) (for a M × N image MN number of coefficients are extracted). \( \psi \) is the complex conjugate of those defined in Eqs. (1) and (2). The scale parameter range plays and important part in our analysis. Since it is computationally intensive to use many scale parameters and also not all scale parameter represents the details of the lung histopathology, small scale and large scale parameters will have over or low detailed information about the image. We empirically found \( a = 3 \) to 6 suited for our work.
2.4 Wavelet Coefficients Modelling and Similarity Measurement
The marginal distribution of the Marr and isotropic Morlet wavelet coefficients are long tailed, bell-shaped and centered around zero (see Fig. 3). To model such a distribution GGD [22] is used. Two varying parameters can be used to approximate the coefficient of the wavelet transform as shown below:
(a) Marr wavelet coefficient for a particular subband is represented as a histogram for a random 256 × 265 image in L* colour space and selected from ADC and SCC data set. (b) Histogram representing the magnitude of isotropic Morlet wavelet coefficient for a subband, random images of size 256 × 256 each in a* colour space for ADC, SCC and ULR is used.
Where \( \Gamma \) is a gamma function and \( \alpha \) is a scale parameter which model the width of the Probability density function(PDF) peak, while \( \beta \) is the shape parameter which is inversely proportional to the decreasing rate of the peak. In Fig. 3(b) the distributions are created from the images using isotropic Morlet wavelet, to model shapes described in the figure Laplace distribution can be use. Since Laplace distribution is a special case of GGD (\( \beta = 0 \)), so only the latter distribution is considered. The parameters \( \alpha \) and \( \beta \) are estimated by maximum likelihood estimator (MLE) [22]. The scale and shape parameters are used as features for the classifier. Various statistical measures [23] of the GGD such as variance, kurtosis and entropy are used also used as an input feature vector to the classifier.
To quantify the difference between two empirical distributions a distance measure was used. The distance between two distributions was calculated using Kullback-Leibler divergence (KL-D), KL-D cannot be used as a metric since it is not symmetric and a symmetric version [24] of KL-D was implemented for this work. Jensen-Shannon divergence (J-divergence) with multiple probability distributions [25] is used to calculate the similarity of more than two distributions, J-divergence is symmetric. To quantify the goodness of fit of the GGD model to the observed distribution, symmetric KL-D and the \( \chi^{2} \) test is used.
3 Results and Discussion
This section provides evidence that the method that is proposed in this work is implemented correctly and the features that are used for classification are indeed can classify different lung tissue textures. The results reveal the accuracy of our method in classifying ADC, SCC and ULR from a data set. The specifics of the data set are tabled in Table 1, the SVM classifier is used to classify the subtype of the NSCLC with input feature vectors obtained from various methods used in this study.
3.1 Goodness of Fit
The goodness of fit of the model into the empirical distribution of the coefficients is calculated with symmetric KL-divergence and \( \chi^{2} \) test at 5% confidence level. The model which we were trying to fit to the observed distribution was assumed as the null hypothesis. Considering all the data 96.75% have accepted the null hypothesis, i.e. the chi square values in these cases are found to be lower than the upper limit of \( \chi^{2} \) distribution i.e. \( \chi_{{\left( {0.05} \right)}}^{2} = 3.841 \) with degree of freedom equals 1. Table 2 represent the goodness of fit of the data represented by symmetric KL-divergence and Pearson’s \( \chi^{2} \) values of the distribution. The GGD model fits the isotropic Morlet distribution more accurately since the distributions produced by isotropic Morlet for the given data set have near symmetric values on the both sides around zero (see Fig. 3(b)).
3.2 Similarity Measurement of Empirical Distributions of Various Classes
The similarity of distributions within a class (ADC, SCC or ULR) is high since the J-Divergence (J-D) of all the distributions of a class using a particular wavelet function and scale parameter is low as shown in Table 3.
As intra class similarity is high to calculate inter class similarity a distribution need not be compared with the all the distribution of the comparing class, a distribution from one class is taken and KL-D was applied with \( n^{\prime} \) (we take the value of \( n^{\prime} \) such that \( n^{\prime}\,{<<}\,N \), where N is the total number of distribution for the comparing class) number of distribution from the other class and the average of these KL-D values is taken as the KL-D value of the distribution with the other class. Table 4 shows the inter class KL-D variations, as variations exists these values were used as feature vector for the classifier.
3.3 Classification Results
The features for the SVM are selected through Recursive Feature Elimination (RFE), there are eight distinct features: shape and size parameter from the GGD modelling, three symmetric KL-D values for the three class and three statistical measures (variance, kurtosis and entropy). These eight values are extracted for four scale (a = 3, 4, 5, 6) and each colour channel. Data set as defined in Sect. 2.1 is used and to validate the system a ten-fold cross validation method is employed. The effect of different combinations of features for classifying ULR and malignant tissue (ADC and SCC) is shown in Fig. 4(a).
To classify the ULR, a subset of 41 features from 96 features is used. Using the 41 features an accuracy of 96.2% for ULR and 95.1% malignant tissue is achieved (Fig. 5(a)). Few texture structure of ULR might be similar to malignant texture representation since a ULR consists of many different tissue elements some texture represented by mild necrosis while other may be normal tissue but get damaged due to sample preparation or undergoing mitosis.
Classification accuracy of SVM is maximum with 62 features out of total 96 features used for classifying ADC and SCC, Fig. 4(b) shows the variation of classification accuracy between ADC and SCC for various feature sets. An accuracy of 77.2% is achieved in classifying ADC and the method gives an accuracy of 75.8% for classifying SCC, refer Fig. 5(b). These results are satisfactory since conclusive diagnosis of these two subtypes of NSCLC is even contradictory to different pathologists, since the complex organisation of the tissue structures can be seen for different stages of cancer. To have a concrete diagnostic answer often molecular analysis is carried out.
Proposed method is also compared with Gray Level Co-occurrence Matrix (GLCM), with angle \( \left( {0^{ \circ } ,45^{ \circ } ,90^{ \circ } \;and\; 135^{ \circ } } \right) \) and four properties viz. energy, contrast, correlation and homogeneity. The accuracy for ULR, Malignant tissue, ADC, SCC are 78.8%, 81.1%, 65.9%,67.8% respectively for GLCM.
The effect of shape and scale parameter of the GGD on the classification accuracy is very acute, features from L* and b* plays an important role in differentiating ULR from ADC and SCC in our proposed method.
4 Conclusion
In this work, we proposed a method to classify the two important subtypes of NSCLC i.e. Adenocarcinoma and Squamous cell carcinoma. The features for classifying ADC and SCC are not clinical diagnostic features, rather features extracted automatically by a wavelet function from an image. Since colour plays a role in understanding the histological slides by a pathologist, we used the colour information provided by H&E stain. The digitized colour images were transformed into a L*a*b* colour space, this colour space helped in segregating nucleus of a cell from its surrounding. The results we obtained are very promising as characterization of these subtypes of lung cancer are done without any prior knowledge about their morphology coded into the system.
References
Bray, F., Jacques, F., et al.: Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 68, 394–424 (2018)
Ma, L.H., Li, G., et al.: The effect of nonsmall cell lung cancer histology on survival as measured by the graded prognostic assessment in patients with brain metastases treated by hypofractionated stereotactic radiotherapy. Radiat. Oncol. 11, 92 (2016)
Yano, M., Yoshida, J., et al.: The outcomes of a limited resection for nonsmall cell lung cancer based on differences in pathology. World J. Surg. 40(11), 2688–2697 (2016)
Yao, X., Gomes, M.M., et al.: Fine-needle aspiration biopsy versus core-needle biopsy in diagnosing lung cancer: a systematic review. Curr. Oncol. 19(1), 16–27 (2012)
Webb, W.R., Muller, N.L., Naidich, D.P.: High–Resolution CT of the Lung. Lippincott Williams & Wilkins, Philadelphia (2001)
Dundar, M.M., Badve, S.: Computerized classification of intraductal breast lesions using histopathological images. IEEE Trans. Biomed. Eng. 58(7), 1977–1984 (2011)
Sieren, J.C., Weydert, J., et al.: An automated segmentation approach for highlighting the histological complexity of human lung cancer. Ann. Biomed. Eng. 38(12), 3581–3591 (2010)
Nguyen, K., Sabata, B., Jain, A.K.: Prostate cancer grading: gland segmentation and structural features. Pattern Recognit. Lett. 33(7), 951–961 (2012)
Mete, M., Xu, X., et al.: Head and neck cancer detection in histopathological slides. In: 6th IEEE International Conference on Data Mining—Workshops (2006)
Tabesh, A., Teverovskiy, M.: Multifeature prostate cancer diagnosis and Gleason grading of histological images. IEEE Trans. Med. Imaging 26(10), 1366–1378 (2007)
Chekkoury, A., Khurd, P., et al.: Automated malignancy detection in breast histopathological images. In: Medical Imaging 2012: Computer-Aided Diagnosis, San Diego, California, vol. 8315 (2012)
Jafari-Khouzani, K., Soltanian-Zadeh, H.: Multiwavelet grading of pathological images of prostate. IEEE Trans. Biomed. Eng. 50(6), 697–704 (2003)
Khurd, P., Bahlmann, C., Gibbs-Strauss, S.: Computer-aided Gleason grading of prostate cancer histopathological images using Texton forests. In: 2010 IEEE International Symposium on Biomedical Imaging: From Nano to Macro (2010)
Wang, W., John, A., et al.: Detection and classification of thyroid follicular lesions based on nuclear structure from histopathology images. Cytometry Part A 77(5), 485–494 (2010)
Smith, J.R., Chang, S.F.: Transform features for texture classification and discrimination in large image databases. In: Proceedings of the IEEE International Conference on Image Processing (1994)
Scheunders, P., Livens S., et al.: Wavelet-based texture analysis. Int. J. Comput. Sci. Inf. Manag. (1997)
Gao, L., Li, F., Thrall, M.J.: On-the-spot lung cancer differential diagnosis by label-free, molecular vibrational imaging and knowledge-based classification. J. Biomed. Opt. 16(9), 096004 (2011). https://doi.org/10.1117/1.3619294
Sambl, M.L., Camara1, F.: A novel RFE-SVM-based feature selection approach for classification. Int. J. Adv. Sci. Technol. 43, 27–36 (2012)
Batuwita, R., Palade, V.: Class imbalance learning methods for support vector machines. In: He, H., Ma, Y. (eds.) Imbalanced Learning: Foundations Algorithms and Applications. Wiley, New York (2013)
Khan, A.M., Rajpoot, N.: A nonlinear mapping approach to stain normalization in digital histopathology images using image-specific color deconvolution. IEEE Trans. Biomed. Imaging 61(6), 1729–1738 (2014)
Kumar, P.: A wavelet based methodology for scale-space anisotropic analysis. Geophys. Res. Lett. 22(20), 2777–2780 (1995)
Do, M.N., Vetterli M.: Wavelet-based texture retrieval using generalized Gaussian density and Kullback–Leibler distance. IEEE Trans. Image Process. 11(2), 146–158 (2002)
Nadarajah, S.: A generalized normal distribution. J. Appl. Stat. 32(7), 685–694 (2005)
Johnson, D., Sinanovic, S.: Symmetrizing the Kullback-Leibler distance. IEEE Trans. Inf. Theory (2000)
Lin, J.: Divergence measures based on the Shannon entropy. IEEE Trans. Inf. Theory 37(1), 145–151 (1991)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this paper
Cite this paper
Das, M.J., Mahanta, L.B. (2019). Design and Analysis of an Isotropic Wavelet Features-Based Classification Algorithm for Adenocarcinoma and Squamous Cell Carcinoma of Lung Histological Images. In: Deka, B., Maji, P., Mitra, S., Bhattacharyya, D., Bora, P., Pal, S. (eds) Pattern Recognition and Machine Intelligence. PReMI 2019. Lecture Notes in Computer Science(), vol 11942. Springer, Cham. https://doi.org/10.1007/978-3-030-34872-4_6
Download citation
DOI: https://doi.org/10.1007/978-3-030-34872-4_6
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-34871-7
Online ISBN: 978-3-030-34872-4
eBook Packages: Computer ScienceComputer Science (R0)