Abstract
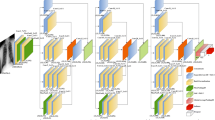
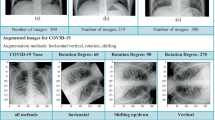
The coronavirus disease 2019 (COVID-19) caused by a novel coronavirus, turned into a pandemic and raised a serious concern to the global healthcare system. The reverse transcription polymerase chain reaction (RT-PCR) is the most widely used diagnostic tool to detect COVID-19. However, this test is time consuming and subject to availability of the test kits during a crisis. An automated method of screening chest x-ray images using convolutional neural network (CNN) Transfer Learning approach has been proposed as a relatively fast and cost-effective, decision support tool to detect pulmonary pathology due to COVID-19. In this study we have used Kaggle dataset with chest x-ray images of normal and pneumonia cases. We have added COVID-19 x-ray images from 5 different open-source datasets. The images were pre-processed based on the position of radiography images and greyscale was applied and subsequently the images were used for training. After consolidation, COVID-19 images comprised only 5% of the dataset. To address the class imbalance, we have used dynamic image augmentation technique to reduce the bias. We have then explored custom CNN and VGG-16, InceptionNet-V3, MobileNet-V2, ResNet-50, and DarkNet-53 transfer learning approaches to classify COVID-19, other pneumonia and normal x-ray images and compared their performances. So far, we have achieved the best score of F1 score 0.95, sensitivity 95% and specificity 95% for COVID-19 class with Darknet-53 feature extractor. Darknet-53 classifier is part of the state-of-the-art object detection algorithm named Yolo-v3. We have also done a McNemar-Bowker post-hoc test to compare Darknet-53 performance with the next best ResNet-50. This test suggests that Darknet-53 is significantly better skilled than ResNet-50 in differentiating COVID-19 from other pneumonia in chest x-ray images.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Corman, V.M., et al.: others, Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Eurosurveillance 25, 2000045 (2020)
Maguolo, G., Nanni, L.: A critic evaluation of methods for COVID-19 automatic detection from X-Ray images (2020). arXiv [eess.IV]
Narin, A., Kaya, C., Pamuk, Z.: Automatic detection of Coronavirus disease (COVID-19) using X-ray images and deep convolutional neural networks (2020). arXiv [eess.IV]
Ozturk, T., Talo, M., Yildirim, E.A., Baloglu, U.B., Yildirim, O., Rajendra Acharya, U.: Automated detection of COVID-19 cases using deep neural networks with X-ray images. Comput. Biol. Med. 121, 103792 (2020)
Cohen, J.P., Morrison, P., Dao, L., Roth, K., Duong, T.Q., Ghassemi, M.: COVID-19 Image Data Collection: Prospective Predictions Are the Future (2020). arXiv [q-bio.QM]
Wang, L., Lin, Z.Q., Wong, A.: COVID-Net: a tailored deep convolutional neural network design for detection of COVID-19 cases from chest X-ray images. Sci. Rep. 10(1), 19549 (2020)
Ng, M.-Y., et al.: Imaging profile of the COVID-19 infection: Radiologic findings and literature review. Radiol. Cardiothorac. Imaging. 2(1), 200034 (2020)
Abiyev, R.H., Maaitah, M.K.S.: Deep convolutional neural networks for chest diseases detection. J. Healthc. Eng. 2018, 1–11 (2018)
Shin, H.-C., et al.: Deep convolutional neural networks for computer-aided detection: CNN architectures, dataset characteristics and transfer learning. IEEE Trans. Med. Imaging 35(5), 1285–1298 (2016)
Redmon, J., Farhadi, A.: YOLOv3: An Incremental Improvement (2018). arXiv [cs.CV]
Convert Darknet-53 ImageNet weights and architecture from NVIDIA CUDA framework to Tensor Flow 2.0: Github repository - https://github.com/qqwweee/keras-yolo3
COVID-19 Image Data collection from Cohen et al.: [10] - https://github.com/ieee8023/covid-chestxray-dataset
COVID-19 datasets with Bounding box and Segmentation information: https://github.com/GeneralBlockchain
Kaggle RSNA Pneumonia detection challenge dataset: https://www.kaggle.com/c/rsna-pneumonia-detection-challenge
Data Source reference for Kaggle RSNA Pneumonia detection: https://nihcc.app.box.com/v/ChestXray-NIHCC
Figure 1 COVID-19 Chest X-ray Dataset Initiative - https://github.com/agchung/Figure1-COVID-chestxray-dataset
Actualmed COVID-19 Chest X-ray Dataset Initiative - https://github.com/agchung/Actualmed-COVID-chestxray-dataset
Italian Society of Medical and Interventional Radiology (SIRM). https://www.kaggle.com/tawsifurrahman/covid19-radiography-database
DarkNet-53 ImageNet weights in CUDA format. https://pjreddie.com/darknet/imagenet/#darknet53
Simonyan, K., Zisserman, A.: Very deep convolutional networks for large-scale image recognition (2014). arXiv [cs.CV]
Darapaneni, N., et al.: COVID 19 severity of pneumonia analysis using chest X rays. In: 2020 IEEE 15th International Conference on Industrial and Information Systems (ICIIS), pp. 381–386 (2020)
Szegedy, C., Vanhoucke, V., Ioffe, S., Shlens, J., Wojna, Z.: Rethinking the inception architecture for computer vision. In: 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (2016)
He, K., Zhang, X., Ren, S., Sun, J.: Deep residual learning for image recognition (2015). arXiv [cs.CV]
Dubey, R., Sen, K.K., Panda, S., Goyal, M., Menon, S.M., Arora, R.: Potential of Conventional Chest Radiography as a diagnostic and follow up imaging tool in COVID pandemic era Ejmcm.com. https://www.ejmcm.com/article_7273_4828eb9ff564ab3bc54bcfee5973b8b6.pdf. Accessed 10 Apr 2021
Ronneberger, O., Fischer, P., Brox, T.: U-Net: Convolutional Networks for Biomedical Image Segmentation (2015). arXiv [cs.CV]
Darapaneni, N., et al.: Inception C-net(IC-net): altered inception module for detection of covid-19 and pneumonia using chest X-rays. In: 2020 IEEE 15th International Conference on Industrial and Information Systems (ICIIS), pp. 393–398 (2020)
Albawi, S., Mohammed, T.A., Al-Zawi, S.: Understanding of a convolutional neural network. Int. Conf. Eng. Technol. (ICET) 2017, 1–6 (2017). https://doi.org/10.1109/ICEngTechnol.2017.8308186
Neuman, M.I., et al.: Variability in the interpretation of chest radiographs for the diagnosis of pneumonia in children. J. Hosp. Med. 7(4), 294–298 (2012)
Wikipedia reference for Evaluating McNemar test statistic. https://en.wikipedia.org/wiki/McNemar%27s_test
Reference book for Bowkers test – Basic Biostatistics for medical and Bio-Medical practioners (second edition) (2019). https://www.sciencedirect.com/book/9780128170847/basic-biostatistics-for-medical-and-biomedical-practitioners
Sandler, M., Howard, A., Zhu, M., Zhmoginov, A., Chen, L.-C.: MobileNetV2: Inverted residuals and linear bottlenecks (2018). arXiv [cs.CV]
Yolo-v3 author PJ Reddie’s website used for downloading DarkNet-53 weights. https://pjreddie.com/darknet/imagenet/#darknet53
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 IFIP International Federation for Information Processing
About this paper
Cite this paper
Darapaneni, N. et al. (2021). Deep Convolutional Neural Network (CNN) Design for Pathology Detection of COVID-19 in Chest X-Ray Images. In: Holzinger, A., Kieseberg, P., Tjoa, A.M., Weippl, E. (eds) Machine Learning and Knowledge Extraction. CD-MAKE 2021. Lecture Notes in Computer Science(), vol 12844. Springer, Cham. https://doi.org/10.1007/978-3-030-84060-0_14
Download citation
DOI: https://doi.org/10.1007/978-3-030-84060-0_14
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-84059-4
Online ISBN: 978-3-030-84060-0
eBook Packages: Computer ScienceComputer Science (R0)