Abstract
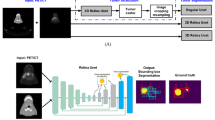
Several recent PET/CT radiomics studies have shown promising results for the prediction of patient outcomes in Head and Neck (H&N) cancer. These studies, however, are most often conducted on relatively small cohorts (up to 300 patients) and using manually delineated tumors. Recently, deep learning reached high performance in the automatic segmentation of H&N primary tumors in PET/CT. The automatic segmentation could be used to validate these studies on larger-scale cohorts while obviating the burden of manual delineation. We propose a complete PET/CT processing pipeline gathering the automatic segmentation of primary tumors and prognosis prediction of patients with H&N cancer treated with radiotherapy and chemotherapy. Automatic contours of the primary Gross Tumor Volume (GTVt) are obtained from a 3D UNet. A radiomics pipeline that automatically predicts the patient outcome (Disease Free Survival, DFS) is compared when using either the automatically or the manually annotated contours. In addition, we extract deep features from the bottleneck layers of the 3D UNet to compare them with standard radiomics features (first- and second-order as well as shape features) and to test the performance gain when added to them. The models are evaluated on the HECKTOR 2020 dataset consisting of 239 H&N patients with PET, CT, GTVt contours and DFS data available (five centers). Regarding the results, using Hand-Crafted (HC) radiomics features extracted from manual GTVt achieved the best performance and is associated with an average Concordance (C) index of 0.672. The fully automatic pipeline (including deep and HC features from automatic GTVt) achieved an average C index of 0.626, which is lower but relatively close to using manual GTVt (p-value = 0.20). This suggests that large-scale studies could be conducted using a fully automatic pipeline to further validate the current state of the art H&N radiomics. The code will be shared publicly for reproducibility.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
Notes
- 1.
We can unconventionally detail the number of HC features as follows: \(274 = 2 \text { modalities}\times (2\text { bin}\times 56 \text { 2}^{nd}\text {order} + 18 \text { 1}^{st}\text {order}) + 14 \text { shape}\) (see Table 2).
References
Gillies, R.J., Kinahan, P.E., Hricak, H.: Radiomics: images are more than pictures, they are data. Radiology 278(2), 563–577 (2016)
Zwanenburg, A., et al.: The image biomarker standardization initiative: standardized quantitative radiomics for high-throughput image-based phenotyping. Radiology 295(2), 328–338 (2020)
Vallieres, M., et al.: Radiomics strategies for risk assessment of tumour failure in head-and-neck cancer. Sci. Rep. 7(1), 1–14 (2017)
Andrearczyk, V., et al.: Automatic segmentation of head and neck tumors and nodal metastases in PET-CT scans. In International Conference on Medical Imaging with Deep Learning (MIDL) (2020)
Apostolova, I., et al.: Asphericity of pretherapeutic tumour FDG uptake provides independent prognostic value in head-and-neck cancer. Eur. Radiol. 24(9), 2077–2087 (2014)
Andrearczyk, V., et al.: Overview of the HECKTOR challenge at MICCAI 2020: automatic head and neck tumor segmentation in PET/CT. In: Lecture Notes in Computer Science (LNCS) Challenges (2021)
Ronneberger, O., Fischer, P., Brox, T.: U-Net: Convolutional Networks for Biomedical Image Segmentation. In: Navab, N., Hornegger, J., Wells, W.M., Frangi, A.F. (eds.) MICCAI 2015. LNCS, vol. 9351, pp. 234–241. Springer, Cham (2015). https://doi.org/10.1007/978-3-319-24574-4_28
Menze, B.H., et al.: The multimodal brain tumor image segmentation benchmark (BRATS). IEEE Trans. Med. Imaging 34(10), 1993–2024 (2014)
Havaei, M.: Brain tumor segmentation with deep neural networks. Med. Image Anal. 35, 18–31 (2017)
Baid, U., et al.: Deep Learning Radiomics Algorithm for Gliomas (DRAG) Model: A Novel Approach Using 3D UNET Based Deep Convolutional Neural Network for Predicting Survival in Gliomas. In: Crimi, A., Bakas, S., Kuijf, H., Keyvan, F., Reyes, M., van Walsum, T. (eds.) BrainLes 2018. LNCS, vol. 11384, pp. 369–379. Springer, Cham (2019). https://doi.org/10.1007/978-3-030-11726-9_33
Isensee, F., Kickingereder, P., Wick, W., Bendszus, M., Maier-Hein, K.H.: Brain tumor segmentation and radiomics survival prediction: contribution to the BRATS 2017 challenge. In: Crimi, A., Bakas, S., Kuijf, H., Menze, B., Reyes, M. (eds.) BrainLes 2017. LNCS, vol. 10670, pp. 287–297. Springer, Cham (2018). https://doi.org/10.1007/978-3-319-75238-9_25
Andrearczyk, V., Oreiller, V., Depeursinge, A.: Oropharynx detection in PET-CT for tumor segmentation. In: Irish Machine Vision and Image Processing (2020)
Harrell Jr, F.E., Lee, K.L., Mark, D.B.: Multivariable prognostic models: issues in developing models, evaluating assumptions and adequacy, and measuring and reducing errors. Stat. Med. 15(4), 361–387 (1996)
Lambin, P., et al.: Radiomics: the bridge between medical imaging and personalized medicine. Nat. Rev. Clin. Oncol. 14(12), 749–762 (2017)
Suter, Y., et al.: Radiomics for glioblastoma survival analysis in pre-operative MRI: exploring feature robustness, class boundaries, and machine learning techniques. Cancer Imaging 20(1), 1–13 (2020)
David, C.R., et al.: Regression models and life tables (with discussion). J. R. Stat. Soc. 34(2), 187–220 (1972)
Ishwaran, H., et al.: Random survival forests. Ann. Appl. Stat. 2(3), 841–860 (2008)
Lowekamp, B.C., Chen, D.T., Ibáñez, L., Blezek, D.: The design of simpleitk. Front. Neuroinf. 7, 45 (2013)
Van Griethuysen, J.J., et al.: Computational radiomics system to decode the radiographic phenotype. Cancer Res. 77(21), e104–e107 (2017)
Pedregosa, F., et al.: Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011)
Pölsterl, S.: scikit-survival: a library for time-to-event analysis built on top of scikit-learn. J. Mach. Learn. Res. 21(212), 1–6 (2020)
Andrearczyk, V., et al.: Multi-task deep segmentation and radiomics for automatic prognosis in head and neck cancer. In: Rekik, I., Adeli, E., Park, S.H., Schnabel, J. (eds.) Predictive Intelligence in Medicine. PRIME 2021. Lecture Notes in Computer Science, vol. 12928, Springer, Cham (2021). https://doi.org/10.1007/978-3-030-87602-9_14
Bouckaert, R.R., Frank, E.: Evaluating the replicability of significance tests for comparing learning algorithms. In: Dai, H., Srikant, R., Zhang, C. (eds.) PAKDD 2004. LNCS (LNAI), vol. 3056, pp. 3–12. Springer, Heidelberg (2004). https://doi.org/10.1007/978-3-540-24775-3_3
Vorwerk, H., et al.: The delineation of target volumes for radiotherapy of lung cancer patients. Radiother. Oncol. 91(3), 455–460 (2009)
Fontaine, P., Acosta, O., Castelli, J., De Crevoisier, R., Müller, H., Depeursinge, A.: The importance of feature aggregation in radiomics: a head and neck cancer study. Sci. Rep. 10(1), 1–11 (2020)
Zhai, T.T., et al.: Improving the prediction of overall survival for head and neck cancer patients using image biomarkers in combination with clinical parameters. Radiother. Oncol. 124(2), 256–262 (2017)
Acknowledgments
This work was partially supported by the Swiss National Science Foundation (SNSF, grant 205320_179069), the Swiss Personalized Health Network (SPHN via the IMAGINE and QA4IQI projects), and the Hasler Foundation (via the EPICS project, grant 20004).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Switzerland AG
About this paper
Cite this paper
Fontaine, P. et al. (2021). Fully Automatic Head and Neck Cancer Prognosis Prediction in PET/CT. In: Syeda-Mahmood, T., et al. Multimodal Learning for Clinical Decision Support. ML-CDS 2021. Lecture Notes in Computer Science(), vol 13050. Springer, Cham. https://doi.org/10.1007/978-3-030-89847-2_6
Download citation
DOI: https://doi.org/10.1007/978-3-030-89847-2_6
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-89846-5
Online ISBN: 978-3-030-89847-2
eBook Packages: Computer ScienceComputer Science (R0)