Abstract
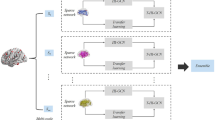
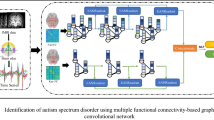
In recent studies, deep learning has shown great potential to explore topological properties of functional connectivity (FC), e.g., graph neural networks (GNNs), for brain disease diagnosis, e.g., Autism spectrum disorder (ASD). However, many of the existing methods integrate the information locally, e.g., among neighboring nodes in a graph, which hinders from learning complex patterns of FC globally. In addition, their analysis for discovering imaging biomarkers is confined to providing the most discriminating regions without considering individual variations over the average FC patterns of groups, i.e., patients and normal controls. To address these issues, we propose a unified framework that globally captures properties of inter-network connectivity for classification and provides individual-specific group characteristics for interpretation via prototype learning. In our experiments using the ABIDE dataset, we validated the effectiveness of the proposed framework by comparing with competing topological deep learning methods in the literature. Furthermore, we individually analyzed functional mechanisms of ASD for neurological interpretation.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
Notes
- 1.
The code of our proposed model is available at https://github.com/ku-milab/PL-FC.
- 2.
Scan procedure and protocols can be found at http://fcon_1000.projects.nitrc.org/indi/abide/.
- 3.
With a total loss of \(\mathcal {L}_{\text {all}} = \lambda _1\mathcal {L}_{\text {recon}} + \lambda _2\mathcal {L}_{\text {class}} + \lambda _3\mathcal {L}_{\text {proto}}\), where \(\lambda _{1/2/3}\) denote weight parameters.
References
American Psychiatric Association, D., Association, A.P., et al.: Diagnostic and statistical manual of mental disorders: DSM-5, vol. 5. American Psychiatric Association Washington, DC (2013)
Chen, C., Li, O., Tao, D., Barnett, A., Rudin, C., Su, J.K.: This looks like that: deep learning for interpretable image recognition. In: Advances in Neural Information Processing Systems, vol. 32 (2019)
Craddock, R.C., James, G.A., Holtzheimer, P.E., III., Hu, X.P., Mayberg, H.S.: A whole brain fMRI atlas generated via spatially constrained spectral clustering. Hum. Brain Mapp. 33(8), 1914–1928 (2012)
Dosovitskiy, A., et al.: An image is worth 16x16 words: transformers for image recognition at scale. In: International Conference on Learning Representations (2021)
Hamilton, W., Ying, Z., Leskovec, J.: Inductive representation learning on large graphs. In: Advances in Neural Information Processing Systems, vol. 30 (2017)
Jun, E., Kang, E., Choi, J., Suk, H.I.: Modeling regional dynamics in low-frequency fluctuation and its application to autism spectrum disorder diagnosis. Neuroimage 184, 669–686 (2019)
Kam, T.E., Suk, H.I., Lee, S.W.: Multiple functional networks modeling for autism spectrum disorder diagnosis. Hum. Brain Mapp. 38(11), 5804–5821 (2017)
Kawahara, J., et al.: BrainNetCNN: convolutional neural networks for brain networks; towards predicting neurodevelopment. Neuroimage 146, 1038–1049 (2017)
Kazeminejad, A., Sotero, R.C.: Topological properties of resting-state fMRI functional networks improve machine learning-based autism classification. Front. Neurosci. 12, 1018 (2019)
Kim, B.H., Ye, J.C.: Understanding graph isomorphism network for rs-fMRI functional connectivity analysis. Front. Neurosci. 14 (2020)
Li, X., et al.: BrainGNN: interpretable brain graph neural network for fMRI analysis. Med. Image Anal. 74, 102233 (2021)
Suk, H.I., Wee, C.Y., Lee, S.W., Shen, D.: Supervised discriminative group sparse representation for mild cognitive impairment diagnosis. Neuroinformatics 13(3), 277–295 (2015)
Vaswani, A., et al.: Attention is all you need. In: Advances in Neural Information Processing Systems, vol. 30 (2017)
Veličkovič, P., Cucurull, G., Casanova, A., Romero, A., Liò, P., Bengio, Y.: Graph attention networks. In: International Conference on Learning Representations (2018)
Yang, H.M., Zhang, X.Y., Yin, F., Liu, C.L.: Robust classification with convolutional prototype learning. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp. 3474–3482 (2018)
Ying, C., et al.: Do transformers really perform badly for graph representation? In: Advances in Neural Information Processing Systems, vol. 34 (2021)
Zhao, K., Duka, B., Xie, H., Oathes, D.J., Calhoun, V., Zhang, Y.: A dynamic graph convolutional neural network framework reveals new insights into connectome dysfunctions in ADHD. Neuroimage 246, 118774 (2022)
Zhao, Q., Liu, Z., Adeli, E., Pohl, K.M.: Longitudinal self-supervised learning. Med. Image Anal. 71, 102051 (2021)
Zhi, D., et al.: BNCPL: Brain-network-based convolutional prototype learning for discriminating depressive disorders. In: 2021 43rd Annual International Conference of the IEEE Engineering in Medicine & Biology Society, pp. 1622–1626 (2021)
Acknowledgements
This work was supported by Institute of Information & communications Technology Planning & Evaluation (IITP) grant funded by the Korea government (MSIT) No. 2022-0-00959 ((Part 2) Few-Shot Learning of Causal Inference in Vision and Language for Decision Making) and Institute of Information & communications Technology Planning & Evaluation (IITP) grant funded by the Korea government (MSIT) (No. 2019-0-00079, Artificial Intelligence Graduate School Program (Korea University))
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
1 Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Kang, E., Heo, DW., Suk, HI. (2022). Prototype Learning of Inter-network Connectivity for ASD Diagnosis and Personalized Analysis. In: Wang, L., Dou, Q., Fletcher, P.T., Speidel, S., Li, S. (eds) Medical Image Computing and Computer Assisted Intervention – MICCAI 2022. MICCAI 2022. Lecture Notes in Computer Science, vol 13433. Springer, Cham. https://doi.org/10.1007/978-3-031-16437-8_32
Download citation
DOI: https://doi.org/10.1007/978-3-031-16437-8_32
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-16436-1
Online ISBN: 978-3-031-16437-8
eBook Packages: Computer ScienceComputer Science (R0)