Abstract
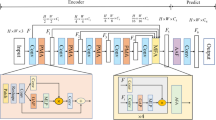
We present a new encoder-decoder Vision Transformer architecture, Patcher, for medical image segmentation. Unlike standard Vision Transformers, it employs Patcher blocks that segment an image into large patches, each of which is further divided into small patches. Transformers are applied to the small patches within a large patch, which constrains the receptive field of each pixel. We intentionally make the large patches overlap to enhance intra-patch communication. The encoder employs a cascade of Patcher blocks with increasing receptive fields to extract features from local to global levels. This design allows Patcher to benefit from both the coarse-to-fine feature extraction common in CNNs and the superior spatial relationship modeling of Transformers. We also propose a new mixture-of-experts (MoE) based decoder, which treats the feature maps from the encoder as experts and selects a suitable set of expert features to predict the label for each pixel. The use of MoE enables better specializations of the expert features and reduces interference between them during inference. Extensive experiments demonstrate that Patcher outperforms state-of-the-art Transformer- and CNN-based approaches significantly on stroke lesion segmentation and polyp segmentation. Code for Patcher is released to facilitate related research. (Code: https://github.com/YanglanOu/patcher.git.).
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Cao, H., et al.: Swin-Unet: Unet-like pure transformer for medical image segmentation. arXiv preprint arXiv:2105.05537 (2021)
Chen, J., et al.: TransUNet: Transformers make strong encoders for medical image segmentation. arXiv preprint arXiv:2102.04306 (2021)
Çiçek, Ö., Abdulkadir, A., Lienkamp, S.S., Brox, T., Ronneberger, O.: 3D U-Net: learning dense volumetric segmentation from sparse annotation. In: Ourselin, S., Joskowicz, L., Sabuncu, M.R., Unal, G., Wells, W. (eds.) MICCAI 2016. LNCS, vol. 9901, pp. 424–432. Springer, Cham (2016). https://doi.org/10.1007/978-3-319-46723-8_49
Dosovitskiy, A., et al.: An image is worth \(16\times 16\) words: Transformers for image recognition at scale. arXiv preprint arXiv:2010.11929 (2020)
Hatamizadeh, A., et al.: UNETR: Transformers for 3D medical image segmentation. In: Proceedings of the IEEE/CVF Winter Conference on Applications of Computer Vision, pp. 574–584 (2022)
Huang, C.H., Wu, H.Y., Lin, Y.L.: HarDNet-MSEG: a simple encoder-decoder polyp segmentation neural network that achieves over 0.9 mean dice and 86 FPS. arXiv preprint arXiv:2101.07172 (2021)
Jha, D., et al.: Kvasir-SEG: a segmented polyp dataset. In: Ro, Y.M., et al. (eds.) MMM 2020. LNCS, vol. 11962, pp. 451–462. Springer, Cham (2020). https://doi.org/10.1007/978-3-030-37734-2_37
Jha, D., et al.: ResUNet++: An advanced architecture for medical image segmentation. In: Proceedings of the IEEE International Symposium on Multimedia (ISM), pp. 225–2255. IEEE (2019)
Kingma, D.P., Ba, J.: Adam: A method for stochastic optimization. arXiv preprint arXiv:1412.6980 (2014)
Liu, Z., et al.: Swin Transformer: Hierarchical vision transformer using shifted windows. Proceedings of the International Conference on Computer Vision (ICCV) (2021)
Loshchilov, I., Hutter, F.: Decoupled weight decay regularization. arXiv preprint arXiv:1711.05101 (2017)
Oktay, O., et al.: Attention U-Net: Learning where to look for the pancreas. arXiv preprint arXiv:1804.03999 (2018)
Ou, Y., et al.: LambdaUNet: 2.5D Stroke lesion segmentation of diffusion-weighted MR images. In: de Bruijne, M., et al. (eds.) MICCAI 2021. LNCS, vol. 12901, pp. 731–741. Springer, Cham (2021). https://doi.org/10.1007/978-3-030-87193-2_69
Petit, O., Thome, N., Rambour, C., Themyr, L., Collins, T., Soler, L.: U-Net transformer: self and cross attention for medical image segmentation. In: Lian, C., Cao, X., Rekik, I., Xu, X., Yan, P. (eds.) MLMI 2021. LNCS, vol. 12966, pp. 267–276. Springer, Cham (2021). https://doi.org/10.1007/978-3-030-87589-3_28
Ronneberger, O., Fischer, P., Brox, T.: U-Net: Convolutional networks for biomedical image segmentation. In: Navab, N., Hornegger, J., Wells, W.M., Frangi, A.F. (eds.) MICCAI 2015. LNCS, vol. 9351, pp. 234–241. Springer, Cham (2015). https://doi.org/10.1007/978-3-319-24574-4_28
Srivastava, R.K., Greff, K., Schmidhuber, J.: Highway networks. arXiv preprint arXiv:1505.00387 (2015)
Vaswani, A., et al.: Attention is all you need. In: Advances in Neural Information Processing Systems, vol. 30 (2017)
Xie, E., Wang, W., Yu, Z., Anandkumar, A., Alvarez, J.M., Luo, P.: SegFormer: Simple and efficient design for semantic segmentation with transformers. arXiv preprint arXiv:2105.15203 (2021)
Zhang, Y., Liu, H., Hu, Q.: TransFuse: Fusing transformers and cnns for medical image segmentation. In: de Bruijne, M., et al. (eds.) MICCAI 2021. LNCS, vol. 12901, pp. 14–24. Springer, Cham (2021). https://doi.org/10.1007/978-3-030-87193-2_2
Zheng, S., et al.: Rethinking semantic segmentation from a sequence-to-sequence perspective with transformers. In: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, pp. 6881–6890 (2021)
Zhou, Z., Rahman Siddiquee, M.M., Tajbakhsh, N., Liang, J.: UNet++: A nested U-Net architecture for medical image segmentation. In: Stoyanov, D., et al. (eds.) DLMIA/ML-CDS -2018. LNCS, vol. 11045, pp. 3–11. Springer, Cham (2018). https://doi.org/10.1007/978-3-030-00889-5_1
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
1 Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Ou, Y. et al. (2022). Patcher: Patch Transformers with Mixture of Experts for Precise Medical Image Segmentation. In: Wang, L., Dou, Q., Fletcher, P.T., Speidel, S., Li, S. (eds) Medical Image Computing and Computer Assisted Intervention – MICCAI 2022. MICCAI 2022. Lecture Notes in Computer Science, vol 13435. Springer, Cham. https://doi.org/10.1007/978-3-031-16443-9_46
Download citation
DOI: https://doi.org/10.1007/978-3-031-16443-9_46
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-16442-2
Online ISBN: 978-3-031-16443-9
eBook Packages: Computer ScienceComputer Science (R0)