Abstract
Stress is a body reaction that is one of the principal causes of many physical and mental disorders, including cardiovascular disease and depression. Developing robust methods for rapid and accurate stress detection plays an important role in improving people’s life quality and wellness. Prior research shows that analyzing physiological signals collected from wearable sensors is a reliable predictor of stress. For stress detection, methods based on machine learning techniques have been defined in the literature. However, they require hand-crafted features to be effective. Deep learning-based approaches overcome these limitations.
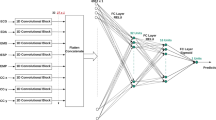
In this work, we introduce STREDWES, a method for stress detection that analyzes biosignals obtained from wearable sensor data. STREDWES extracts signal fragments using a sliding windows approach and converts them into Gramian Angular Fields images. These images are then classified using a Convolutional Neural Network, a deep learning algorithm. We apply our method to a publicly available dataset. The analysis of the performance values shows that our method outperforms other state-of-the-art competitors.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
Notes
- 1.
the tuning algorithm chooses the best scale for the hyperparameter exploration among linear, logarithmic, and reverse logarithmic.
References
Bara, C.P., Papakostas, M., Mihalcea, R.: A deep learning approach towards multimodal stress detection. In: AffCon@ AAAI, pp. 67–81 (2020)
Chakraborty, S., Aich, S., Joo, M.I., Sain, M., Kim, H.C.: A multichannel convolutional neural network architecture for the detection of the state of mind using physiological signals from wearable devices. J. Healthc. Eng. 2019, 5397814 (2019)
Chollet, F., et al.: Keras (2015). https://keras.io
Girija, S.S.: Tensorflow: large-scale machine learning on heterogeneous distributed systems. Software available from tensorflow. org 39(9), (2016)
Health and Safety Executive: HSE on work-related stress (2021). http://www.hse.gov.uk/statistics/causdis/-ffstress/index.htm. Accessed 7 Mar 2022
Hovsepian, K., Al’Absi, M., Ertin, E., Kamarck, T., Nakajima, M., Kumar, S.: cStress: towards a gold standard for continuous stress assessment in the mobile environment. In: Proceedings of the 2015 ACM International Joint Conference on Pervasive and Ubiquitous Computing, pp. 493–504 (2015)
Koldijk, S., Sappelli, M., Verberne, S., Neerincx, M.A., Kraaij, W.: The swell knowledge work dataset for stress and user modeling research. In: Proceedings of the 16th International Conference on Multimodal Interaction, pp. 291–298 (2014)
Lee, E.H.: Review of the psychometric evidence of the perceived stress scale. Asian Nurs. Res. 6(4), 121–127 (2012)
Lee, H., Yang, K., Kim, N., Ahn, C.R.: Detecting excessive load-carrying tasks using a deep learning network with a Gramian angular field. Autom. Constr. 120, 103390 (2020)
Lin, J., Pan, S., Lee, C.S., Oviatt, S.: An explainable deep fusion network for affect recognition using physiological signals. In: Proceedings of the 28th ACM International Conference on Information and Knowledge Management, pp. 2069–2072 (2019)
McEwen, B.S.: Protective and damaging effects of stress mediators. N. Engl. J. Med. 338(3), 171–179 (1998)
Piangerelli, M., Maestri, S., Merelli, E.: Visualising 2-simplex formation in metabolic reactions. J. Mol. Graph. Model. 97, 107576 (2020)
Plarre, K., et al.: Continuous inference of psychological stress from sensory measurements collected in the natural environment. In: Proceedings of the 10th ACM/IEEE International Conference on Information Processing in Sensor Networks, pp. 97–108. IEEE (2011)
Quadrini, M., Cavallin, M., Daberdaku, S., Ferrari, C.: ProSPs: protein sites prediction based on sequence fragments. In: International Conference on Machine Learning, Optimization, and Data Science, pp. 568–580. Springer, Cham (2021). https://doi.org/10.1007/978-3-030-95467-3_41
Quadrini, M., Daberdaku, S., Ferrari, C.: Hierarchical representation and graph convolutional networks for the prediction of protein–protein interaction sites. In: Nicosia, G., et al. (eds.) LOD 2020. LNCS, vol. 12566, pp. 409–420. Springer, Cham (2020). https://doi.org/10.1007/978-3-030-64580-9_34
Quadrini, M., Daberdaku, S., Ferrari, C.: Hierarchical representation for PPI sites prediction. BMC Bioinf. 23(1), 1–34 (2022)
Quadrini, M., Merelli, E., Piergallini, R.: Loop grammars to identify RNA structural patterns. In: BIOINFORMATICS, pp. 302–309 (2019)
Quadrini, M., Tesei, L., Merelli, E.: ASPRAlign: a tool for the alignment of RNA secondary structures with arbitrary pseudoknots. Bioinformatics 36(11), 3578–3579 (2020)
Schmidt, P., Reiss, A., Duerichen, R., Marberger, C., Van Laerhoven, K.: Introducing WESAD, a multimodal dataset for wearable stress and affect detection. In: Proceedings of the 20th ACM International Conference on Multimodal Interaction, pp. 400–408 (2018)
Uddin, M.T., Canavan, S.: Synthesizing physiological and motion data for stress and meditation detection. In: 2019 8th International Conference on Affective Computing and Intelligent Interaction Workshops and Demos (ACIIW), pp. 244–247. IEEE (2019)
Wang, Z., Oates, T.: Encoding time series as images for visual inspection and classification using tiled convolutional neural networks. In: Workshops at the Twenty-Ninth AAAI Conference on Artificial Intelligence (2015)
Wang, Z., Oates, T.: Imaging time-series to improve classification and imputation. In: Twenty-Fourth International Joint Conference on Artificial Intelligence (2015)
Acknowledgements
MQ is supported by the “GNCS - INdAM”. The authors are grateful to Simone Cardis @ Sorint.Tek for his helpful insights and to Zerina Koplikaj @ Sorint.Tek for proofreading the manuscript.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Quadrini, M., Daberdaku, S., Blanda, A., Capuccio, A., Bellanova, L., Gerard, G. (2022). Stress Detection from Wearable Sensor Data Using Gramian Angular Fields and CNN. In: Pascal, P., Ienco, D. (eds) Discovery Science. DS 2022. Lecture Notes in Computer Science(), vol 13601. Springer, Cham. https://doi.org/10.1007/978-3-031-18840-4_13
Download citation
DOI: https://doi.org/10.1007/978-3-031-18840-4_13
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-18839-8
Online ISBN: 978-3-031-18840-4
eBook Packages: Computer ScienceComputer Science (R0)