Abstract
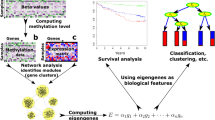
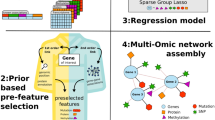
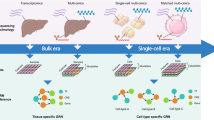
Multi-layer Complex networks are commonly used for modeling and analysing biological entities. This paper presents a new computational framework called COMBO (Combining Multi Bio Omics) for generating and analyzing heterogeneous multi-layer networks. Our model uses gene expression and DNA-methylation data. The power of COMBO relies on its ability to join different omics to study the complex interplay between various components in the disease. We tested the reliability and versatility of COMBO on colon and lung adenocarcinoma cancer data obtained from the TCGA database.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Boccaletti, S., Latora, V., Moreno, Y., Chavez, M., Hwang, D.: Complex networks: structure and dynamics. Phys. Rep., 175–308 (2006)
De Domenico, M.: Multilayer network modeling of integrated biological systems: comment on “network science of biological systems at different scales: a review” by Gosak et al. Phys. Life Rev., 149–52 (2018)
Rai, A., Pradhan, P., Nagraj, J., Lohitesh, K., Chowdhury, R., Jalan, S.: Understanding cancer complexome using networks, spectral graph theory and multilayer framework. Sci. Rep. 7, 41676 (2017)
Barabási, A.-L., Gulbahce, N., Loscalzo, J.: Network medicine: a network-based approach to human disease. Nature Rev. Genet., 56–68 (2011)
Goh, K.-I., Cusick, M.E., Valle, D., Childs, B., Vidal, M., Barabási, A.-L.: The human disease network [Internet]. In: Proceedings of the National Academy of Sciences, pp. 8685–90 (2007)
Lv, Y., Huang, S., Zhang, T., Gao, B.: Application of multilayer network models in bioinformatics. Front. Genet. 12, 664860 (2021)
Zheng, W., Wang, D., Zou, X.: Control of multilayer biological networks and applied to target identification of complex diseases. BMC Bioinform. 20, 271 (2019)
Boccaletti, S., Bianconi, G., Criado, R., del Genio, C.I., Gómez-Gardeñes, J., Romance, M., et al.: The structure and dynamics of multilayer networks [Internet]. Phys. Rep., 1–122 (2014)
Mangioni, G., Jurman, G., De Domenico, M.: Multilayer flows in molecular networks identify biological modules in the human proteome [Internet]. IEEE Trans. Netw. Sci. Eng., 411–20 (2020)
Kivela, M., Arenas, A., Barthelemy, M., Gleeson, J.P., Moreno, Y., Porter, M.A.: Multilayer networks [Internet]. J. Compl. Netw., 203–71 (2014)
Kenett, D.Y., Perc, M., Boccaletti, S.: Networks of Networks—An Introduction [Internet]. Chaos, Solitons and Fractals, pp. 1–6 (2015)
Hammoud, Z., Kramer, F.: Multilayer networks: aspects, implementations, and application in biomedicine. Big Data Anal. (2020)
McGee, F., Ghoniem, M., Melançon, G., Otjacques, B., Pinaud, B.: The state of the art in multilayer network visualization. Comp. Graph. Forum. 125–49 (2019)
Bersanelli, M., Mosca, E., Remondini, D., Giampieri, E., Sala, C., Castellani, G., et al.: Methods for the integration of multi-omics data: mathematical aspects. BMC Bioinform. (2016)
Tordini, F., Aldinucci, M., Milanesi, L., Liò, P., Merelli, I.: The Genome Conformation As an Integrator of Multi-Omic Data: The Example of Damage Spreading in Cancer. Frontiers in Genetics (2016)
Zitnik, M., Leskovec, J.: Predicting multicellular function through multi-layer tissue networks. Bioinformatics., i190–8 (2017)
Gligorijević, V., Pržulj, N.: Methods for biological data integration: perspectives and challenges. J. Roy. Soc. Interface., 20150571 (2015)
Domenico, M.D., De Domenico, M., Porter, M.A., Arenas, A.: MuxViz: a tool for multilayer analysis and visualization of networks. J. Compl. Netw., 159–76 (2015)
De Bacco, C., Power, E.A., Larremore, D.B., Moore, C.: Community detection, link prediction, and layer interdependence in multilayer networks. Phys. Rev. E. 95, 042317 (2017)
Škrlj, B., Kralj, J., Lavrač, N.: Py3plex toolkit for visualization and analysis of multilayer networks. Appl. Netw. Sci. (2019)
Hammoud, Z., Kramer, F.M.: An R Package to Create, Modify and Visualize Multilayered Graph. Genes, p. 519 (2018)
Sahoo, D., Dill, D.L., Tibshirani, R., Plevritis, S.K.: Extracting binary signals from microarray time-course data. Nucl. Acids Res., 3705–12 (2007)
Sahoo, D., Dill, D.L., Gentles, A.J., Tibshirani, R., Plevritis, S.K.: Boolean implication networks derived from large scale, whole genome microarray datasets. Genome Biol. 9, R157 (2008)
Ritchie, M.E., Phipson, B., Wu, D., Hu, Y., Law, C.W., Shi, W., et al.: limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucl. Acids Res. 43, e47 (2015)
Du, P., Zhang, X., Huang, C.-C., Jafari, N., Kibbe, W.A., Hou, L., et al.: Comparison of Beta-value and M-value methods for quantifying methylation levels by microarray analysis. BMC Bioinform. BioMed. Central 11, 1–9 (2010)
Sgariglia, D., Conforte, A.J., Pedreira, C.E., de Carvalho, L.A.V., Carneiro, F.R.G., Carels, N., et al.: Data-Driven Modeling of Breast Cancer Tumors Using Boolean Networks. Frontiers in Big Data [Internet]. Frontiers Media SA (2021)
Xu, X., Zhu, L., Yang, Y., Pan, Y., Feng, Z., Li, Y., et al.: Low tumour PPM1H indicates poor prognosis in colorectal cancer via activation of cancer-associated fibroblasts. Br J Cancer. Nature Publ. Group 120, 987–995 (2019)
Dabydeen, S.A., Desai, A., Sahoo, D.: Unbiased Boolean Analysis of Public Gene Expression Data for Cell Cycle Gene Identification. The American Society for Cell Biology, Mol Biol Cell (2019)
Sahoo, D., Wei, W., Auman, H., Hurtado-Coll, A., Carroll, P.R., Fazli, L., et al.: Boolean analysis identifies CD38 as a biomarker of aggressive localized prostate cancer. Oncotarget. Impact J. 9, 6550–6561 (2018)
da Mata, A.S., da Mata, A.S.: Complex networks: a mini-review [Internet]. Brazilian J. Phys. 658–72 (2020)
Kinsley, A.C., Rossi, G., Silk, M.J., VanderWaal, K.: Multilayer and multiplex networks: an introduction to their use in veterinary epidemiology. Front. Vet. Sci. 7, 596 (2020)
Alaimo, S., Giugno, R., Acunzo, M., Veneziano, D., Ferro, A., Pulvirenti, A.: Post-transcriptional knowledge in pathway analysis increases the accuracy of phenotypes classification. Oncotarget 7, 54572–54582 (2016)
Alaimo, S., Marceca, G.P., Ferro, A., Pulvirenti, A.: Detecting disease specific pathway substructures through an integrated systems biology approach. Noncoding RNA. 3 (2017)
Alaimo, S., Rapicavoli, R.V., Marceca, G.P., La Ferlita, A., Serebrennikova, O.B., Tsichlis, P.N., et al.: PHENSIM: phenotype simulator. PLoS Comput. Biol. 17, e1009069 (2021)
Silva, T.C., Colaprico, A., Olsen, C., D’Angelo, F., Bontempi, G., Ceccarelli, M., et al.: TCGA workflow: analyze cancer genomics and epigenomics data using bioconductor packages. F1000 Res., 1542 (2016)
Lambert, S.A., Jolma, A., Campitelli, L.F., Das, P.K., Yin, Y., Albu, M., et al.: The human transcription factors. Cell 175, 598–599 (2018)
Yu, G., He, Q.-Y.: ReactomePA: an R/Bioconductor package for reactome pathway analysis and visualization. Mol. Biosyst. 477–9 (2016)
Condorelli, D.F., Spampinato, G., Valenti, G., Musso, N., Castorina, S., Barresi, V.: Positive Caricature Transcriptomic Effects Associated with Broad Genomic Aberrations in Colorectal Cancer. Scientific Reports (2018)
Condorelli, D.F., Privitera, A.P., Barresi, V.: Chromosomal density of cancer up-regulated genes, aberrant enhancer activity and cancer fitness genes are associated with transcriptional cis-effects of broad copy number GAINs in colorectal cancer. Int. J. Mol. Sci. 20 (2019)
Sillars-Hardebol, A.H., Carvalho, B., Beliën, J.A.M., de Wit, M., Delis-van Diemen, P.M., Tijssen, M., et al.: BCL2L1has a functional role in colorectal cancer and its protein expression is associated with chromosome 20q GAIN. J. Pathol. 442–50 (2012)
Carvalho, B., Postma, C., Mongera, S., Hopmans, E., Diskin, S., van de Wiel, M.A., et al.: Multiple putative oncogenes at the chromosome 20q amplicon contribute to colorectal adenoma to carcinoma progression. Gut 58, 79–89 (2009)
Sillars-Hardebol, A.H., Carvalho, B., Tijssen, M., Beliën, J.A.M., de Wit, M., Delis-van Diemen, P.M., et al.: TPX2 and AURKA promote 20q amplicon-driven colorectal adenoma to carcinoma progression. Gut 61, 1568–1575 (2012)
Ptashkin, R.N., Pagan, C., Yaeger, R., Middha, S., Shia, J., O’Rourke, K.P., et al.: Chromosome 20q amplification defines a subtype of microsatellite stable, left-sided colon cancers with wild-type RAS/RAF and better overall survival. Mol. Cancer Res. (2017)
Voutsadakis, I.A.: Chromosome 20q11.21 amplifications in colorectal cancer. Cancer Genom. Proteom. 18, 487–96 (2021)
Bui, V.M.H., Mettling, C., Jou, J., Sun, H.S.: Genomic amplification of chromosome 20q13.33 is the early biomarker for the development of sporadic colorectal carcinoma. BMC Med. Genom. 13, 149 (2020)
Liu, Q., Guo, L., Qi, H., Lou, M., Wang, R., Hai, B., et al.: A MYBL2 complex for RRM2 transactivation and the synthetic effect of MYBL2 knockdown with WEE1 inhibition aGAINst colorectal cancer. Cell Death Dis. Nature Publ. Group 12, 1–11 (2021)
Song, S., Li, D., Yang, C., Yan, P., Bai, Y., Zhang, Y., et al.: Overexpression of NELFCD promotes colorectal cancer cells proliferation, migration, and invasion. Oncol. Targ. Ther. Dove Press 11, 8741 (2018)
Li, L., Li, P., Zhang, W., Zhou, H., Guo, E., Hu, G., et al.: FERMT1 contributes to the migration and invasion of nasopharyngeal carcinoma through epithelial–mesenchymal transition and cell cycle arrest. Cancer Cell Int. BioMed. Central 22, 1–14 (2022)
Yang, C., Li, D., Bai, Y., Song, S., Yan, P., Wu, R., et al.: DEAD-box helicase 27 plays a tumor-promoter role by regulating the stem cell-like activity of human colorectal cancer cells. Oncol. Targ. Ther. Dove Press 12, 233 (2019)
Wu, S., Zhang, W., Shen, D., Lu, J., Zhao, L.: PLCB4 upregulation is associated with unfavorable prognosis in pediatric acute myeloid leukemia. Oncol. Lett. Spandidos Publ. 18, 6057 (2019)
Belužić, L., Grbeša, I., Belužić, R., Park, J.H., Kong, H.K., Kopjar, N., et al.: Knock-down of AHCY and depletion of adenosine induces DNA damage and cell cycle arrest. Sci. Rep. Nature Publ. Group 8, 1–16 (2018)
Pimiento, J.M., Neill, K.G., Henderson-Jackson, E., Eschrich, S.A., Chen, D.T., Husain, K., et al.: Knockdown of CSE1L gene in colorectal cancer reduces tumorigenesis in vitro. Am. J. Pathol. (2016)
El Khoury, W., Nasr, Z.: Deregulation of ribosomal proteins in human cancers. Biosci Rep. 41 (2021)
Wang, Y., Pan, S., He, X., Wang, Y., Huang, H., Chen, J., et al.: CPNE1 Enhances Colorectal Cancer Cell Growth, Glycolysis, and Drug Resistance Through Regulating the AKT-GLUT1/HK2 Pathway, Vol. 14. Onco Targets Ther. Dove Press, p. 699 (2021)
Chen, J., Elfiky, A., Han, M., Chen, C., Saif, M.W.: The role of Src in colon cancer and its therapeutic implications. Clin. Colorectal Cancer. 13, 5–13 (2014)
Jin, W.: Regulation of Src Family Kinases during Colorectal Cancer Development and Its Clinical Implications. Cancers, pp. 12 (2020)
Yao, C., Li, G., Cai, M., Qian, Y., Wang, L., Xiao, L., et al.: Prostate cancer downregulated SIRP-α modulates apoptosis and proliferation through p38-MAPK/NF-κB/COX-2 signaling. Oncol. Lett. 13, 4995–5001 (2017)
Sanidas, I., Polytarchou, C., Hatziapostolou, M., Ezell, S.A., Kottakis, F., Hu, L., et al.: Phosphoproteomics screen reveals akt isoform-specific signals linking RNA processing to lung cancer. Mol. Cell. 53, 577–590 (2014)
Paronetto, M.P., Passacantilli, I., Sette, C.: Alternative splicing and cell survival: from tissue homeostasis to disease. Cell Death Differ. 23, 1919–1929 (2016)
Coomer, A.O., Black, F., Greystoke, A., Munkley, J., Elliott, D.J.: Alternative splicing in lung cancer. Biochim. Biophys. Acta Gene Regul. Mech. 1862, 194388 (2019)
Yoshimoto, T., Matsubara, D., Soda, M., Ueno, T., Amano, Y., Kihara, A., et al.: Mucin 21 is a key molecule involved in the incohesive growth pattern in lung adenocarcinoma. Cancer Sci. 110, 3006–3011 (2019)
Hou, L., Lin, T., Wang, Y., Liu, B., Wang, M.: Collagen type 1 alpha 1 chain is a novel predictive biomarker of poor progression-free survival and chemoresistance in metastatic lung cancer. J. Cancer. 12, 5723–5731 (2021)
Ruan, J.S., Zhou, H., Yang, L., Wang, L., Jiang, Z.S., Wang, S.M.: CCNA2 facilitates epithelial-to-mesenchymal transition via the integrin αvβ3 signaling in NSCLC. Int. J. Clin. Exp. Pathol. 10, 8324–8333 (2017)
Serveaux-Dancer, M., Jabaudon, M., Creveaux, I., Belville, C., Blondonnet, R., Gross, C., et al.: Pathological implications of receptor for advanced glycation end-product gene polymorphism. Dis. Markers. 2019, 2067353 (2019)
Zhang, W., Fan, J., Chen, Q., Lei, C., Qiao, B., Liu, Q.: SPP1 and AGER as potential prognostic biomarkers for lung adenocarcinoma. Oncol. Lett. 15, 7028–7036 (2018)
Yuan, L., et al.: SFTPA1 is a potential prognostic biomarker correlated with immune cell infiltration and response to immunotherapy in lung adenocarcinoma. Cancer Immunol. Immunother. 71(2), 399–415 (2021). https://doi.org/10.1007/s00262-021-02995-4
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Cosentini, I., Barresi, V., Condorelli, D.F., Ferro, A., Pulvirenti, A., Alaimo, S. (2023). COMBO: A Computational Framework to Analyze RNA-seq and Methylation Data Through Heterogeneous Multi-layer Networks. In: Cherifi, H., Mantegna, R.N., Rocha, L.M., Cherifi, C., Miccichè, S. (eds) Complex Networks and Their Applications XI. COMPLEX NETWORKS 2016 2022. Studies in Computational Intelligence, vol 1077. Springer, Cham. https://doi.org/10.1007/978-3-031-21127-0_21
Download citation
DOI: https://doi.org/10.1007/978-3-031-21127-0_21
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-21126-3
Online ISBN: 978-3-031-21127-0
eBook Packages: EngineeringEngineering (R0)