Abstract
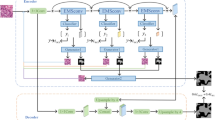
The Bethesda System for Reporting Thyroid Cytopathology (TBSRTC) has been widely accepted as a reliable criterion for thyroid cytology diagnosis, where extensive diagnostic information can be deduced from the allocation and boundary of cell nuclei. However, two major challenges hinder accurate nuclei segmentation from thyroid cytology. Firstly, unbalanced distribution of nuclei morphology across different TBSRTC categories can lead to a biased model. Secondly, the insufficiency of densely annotated images results in a less generalized model. In contrast, image-wise TBSRTC labels, while containing lightweight information, can be deeply explored for segmentation guidance. To this end, we propose a TBSRTC-category aware nuclei segmentation framework (TCSegNet). To top up the small amount of pixel-wise annotations and eliminate the category preference, a larger amount of image-wise labels are taken in as the complementary supervision signal in TCSegNet. This integration of data can effectively guide the pixel-wise nuclei segmentation task with a latent global context. We also propose a semi-supervised extension of TCSegNet that leverages images with only TBSRTC-category labels. To evaluate the proposed framework and also for further cytology cell studies, we curated and elaborately annotated a multi-label thyroid cytology benchmark, collected clinically from 2019 to 2022, which will be made public upon acceptance. Our TCSegNet outperforms state-of-the-art segmentation approaches with an improvement of 2.0% Dice and 2.7% IoU; besides, the semi-supervised extension can further boost this margin. In conclusion, our study explores the weak annotations by constructing an image-wise-label-guided nuclei segmentation framework, which has the potential medical importance to assist thyroid abnormality examination. Code is available at https://github.com/Junchao-Zhu/TCSegNet.
Similar content being viewed by others
References
Ferlay, J., et al.: Cancer incidence and mortality worldwide: sources, methods and major patterns in globocan 2012. Int. J. Cancer 136(5), E359–E386 (2015)
Haugen, B.R., et al.: 2015 American thyroid association management guidelines for adult patients with thyroid nodules and differentiated thyroid cancer: the american thyroid association guidelines task force on thyroid nodules and differentiated thyroid cancer. Thyroid 26(1), 1–133 (2016)
Cibas, E.S., Ali, S.Z.: The bethesda system for reporting thyroid cytopathology. Thyroid 19(11), 1159–1165 (2009)
Kumar, N., et al.: A multi-organ nucleus segmentation challenge. IEEE Trans. Med. Imaging 39(5), 1380–1391 (2019)
Veta, M., et al.: Prognostic value of automatically extracted nuclear morphometric features in whole slide images of male breast cancer. Mod. Pathol. 25(12), 1559–1565 (2012)
Kakudo, K.: Thyroid FNA Cytology: Differential Diagnoses and Pitfalls. Springer, Heidelberg (2019). https://doi.org/10.1007/978-981-13-1897-9
Xing, F., Yang, L.: Robust nucleus/cell detection and segmentation in digital pathology and microscopy images: a comprehensive review. IEEE Rev. Biomed. Eng. 9, 234–263 (2016)
Cibas, E.S., Ali, S.Z.: The 2017 bethesda system for reporting thyroid cytopathology. Thyroid 27(11), 1341–1346 (2017)
Graham, S., et al.: Hover-net: Simultaneous segmentation and classification of nuclei in multi-tissue histology images. Med. Image Anal. 58, 101563 (2019)
Gamper, J., Alemi Koohbanani, N., Benet, K., Khuram, A., Rajpoot, N.: PanNuke: an open pan-cancer histology dataset for nuclei instance segmentation and classification. In: Reyes-Aldasoro, C.C., Janowczyk, A., Veta, M., Bankhead, P., Sirinukunwattana, K. (eds.) ECDP 2019. LNCS, vol. 11435, pp. 11–19. Springer, Cham (2019). https://doi.org/10.1007/978-3-030-23937-4_2
Greenwald, N.F., et al.: Whole-cell segmentation of tissue images with human-level performance using large-scale data annotation and deep learning. Nat. Biotechnol. 40(4), 555–565 (2022)
Everingham, M., Eslami, S.A., Van Gool, L., Williams, C.K., Winn, J., Zisserman, A.: The pascal visual object classes challenge: a retrospective. Int. J. Comput. Vision 111, 98–136 (2015)
Zhou, Y., Onder, O.F., Dou, Q., Tsougenis, E., Chen, H., Heng, P.-A.: CIA-Net: robust nuclei instance segmentation with contour-aware information aggregation. In: Chung, A.C.S., Gee, J.C., Yushkevich, P.A., Bao, S. (eds.) IPMI 2019. LNCS, vol. 11492, pp. 682–693. Springer, Cham (2019). https://doi.org/10.1007/978-3-030-20351-1_53
Huang, J., Shen, Y., Shen, D., Ke, J.: CA2.5-net nuclei segmentation framework with a microscopy cell benchmark collection. In: de Bruijne, M., et al. (eds.) MICCAI 2021. LNCS, vol. 12908, pp. 445–454. Springer, Cham (2021). https://doi.org/10.1007/978-3-030-87237-3_43
Ke, J., et al.: Clusterseg: a crowd cluster pinpointed nucleus segmentation framework with cross-modality datasets. Med. Image Anal. 85, 102758 (2023)
Peng, Z., et al.: Conformer: local features coupling global representations for visual recognition. In: Proceedings of the IEEE/CVF International Conference on Computer Vision, pp. 367–376 (2021)
Hou, Q., Cheng, M.M., Hu, X., Borji, A., Tu, Z., Torr, P.H.: Deeply supervised salient object detection with short connections. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp. 3203–3212 (2017)
Tarvainen, A., Valpola, H.: Mean teachers are better role models: weight-averaged consistency targets improve semi-supervised deep learning results. Adv. Neural Inf. Process. Syst. 30, 1–10 (2017)
Laine, S., Aila, T.: Temporal ensembling for semi-supervised learning. arXiv preprint arXiv:1610.02242 (2016)
He, K., Gkioxari, G., Dollár, P., Girshick, R.: Mask r-cnn. In: Proceedings of the IEEE International Conference on Computer Vision, pp. 2961–2969 (2017)
Cao, H., et al.: Swin-unet: unet-like pure transformer for medical image segmentation. In: Computer Vision-ECCV 2022 Workshops: Tel Aviv, Israel, 23–27 October 2022, Proceedings, Part III, pp. 205–218. Springer, Heidelberg (2023). https://doi.org/10.1007/978-3-031-25066-8_9
Badrinarayanan, V., Kendall, A., Cipolla, R.: Segnet: a deep convolutional encoder-decoder architecture for image segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 39(12), 2481–2495 (2017)
Zhou, Z., Rahman Siddiquee, M.M., Tajbakhsh, N., Liang, J.: UNet++: a nested u-net architecture for medical image segmentation. In: Stoyanov, D., et al. (eds.) DLMIA/ML-CDS -2018. LNCS, vol. 11045, pp. 3–11. Springer, Cham (2018). https://doi.org/10.1007/978-3-030-00889-5_1
Zou, Y., et al.: Pseudoseg: designing pseudo labels for semantic segmentation. arXiv preprint arXiv:2010.09713 (2020)
Chen, X., Yuan, Y., Zeng, G., Wang, J.: Semi-supervised semantic segmentation with cross pseudo supervision. In: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, pp. 2613–2622 (2021)
Luo, X., Hu, M., Song, T., Wang, G., Zhang, S.: Semi-supervised medical image segmentation via cross teaching between cnn and transformer. In: International Conference on Medical Imaging with Deep Learning, pp. 820–833. PMLR (2022)
Chen, Z., Zhu, L., Wan, L., Wang, S., Feng, W., Heng, P.A.: A multi-task mean teacher for semi-supervised shadow detection. In: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, pp. 5611–5620 (2020)
Liu, W., Rabinovich, A., Berg, A.C.: Parsenet: looking wider to see better. arXiv preprint arXiv:1506.04579 (2015)
Acknowledgement
This work was supported by National Natural Science Foundation of China (Grant No. 62102247) and Natural Science Foundation of Shanghai (No. 23ZR1430700).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Zhu, J., Shen, Y., Zhang, H., Ke, J. (2023). An Anti-biased TBSRTC-Category Aware Nuclei Segmentation Framework with a Multi-label Thyroid Cytology Benchmark. In: Greenspan, H., et al. Medical Image Computing and Computer Assisted Intervention – MICCAI 2023. MICCAI 2023. Lecture Notes in Computer Science, vol 14225. Springer, Cham. https://doi.org/10.1007/978-3-031-43987-2_56
Download citation
DOI: https://doi.org/10.1007/978-3-031-43987-2_56
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-43986-5
Online ISBN: 978-3-031-43987-2
eBook Packages: Computer ScienceComputer Science (R0)