Abstract
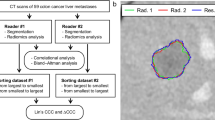
We present an analysis of 81 patients with colorectal liver metastases from two major cancer centers prospectively enrolled in an imaging trial to assess reproducibility of radiomic features in contrast-enhanced CT. All scans were reconstructed with different slice thicknesses and levels of iterative reconstruction. Radiomic features were extracted from the liver parenchyma and largest metastasis from each reconstruction, using different levels of resampling and methods of feature aggregation. The prognostic value of reproducible features was tested using Cox proportional hazards to model overall survival in an independent, public data set of 197 hepatic resection patients with colorectal liver metastases. Our results show that larger differences in slice thickness reduced the concordance of features (\(p<10^{-6}\)). Extracting features with 2.5D aggregation and no axial resampling produced the most robust features, and the best test-set performance in the survival model on the independent data set (C-index = 0.65). Across all feature extraction methods, restricting the survival models to use reproducible features had no statistically significant effect on the test set performance (\(p=0.98\)). In conclusion, our results show that feature extraction settings can positively impact the robustness of radiomics features to variations in slice thickness, without negatively effecting prognostic performance.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Berenguer, R., et al.: Radiomics of CT features may be nonreproducible and redundant: influence of CT acquisition parameters. Radiology 288(2), 407–415 (2018). https://doi.org/10.1148/radiol.2018172361
Carrasco, J.L., Jover, L.: Estimating the generalized concordance correlation coefficient through variance components. Biometrics 59(4), 849–858 (2003). https://doi.org/10.1111/j.0006-341x.2003.00099.x
Clark, K., et al.: The cancer imaging archive (TCIA): maintaining and operating a public information repository. J. Digit. Imaging 26(6), 1045–1057 (2013). https://doi.org/10.1007/s10278-013-9622-7
Emaminejad, N., Wahi-Anwar, M.W., Kim, G.H.J., Hsu, W., Brown, M., McNitt-Gray, M.: Reproducibility of lung nodule radiomic features: Multivariable and univariable investigations that account for interactions between CT acquisition and reconstruction parameters. Med. Phys. 48(6), 2906–2919 (2021). https://doi.org/10.1002/mp.14830
Erdal, B.S., et al.: Are quantitative features of lung nodules reproducible at different CT acquisition and reconstruction parameters? PLoS ONE 15, e0240184 (2020). https://doi.org/10.1371/journal.pone.0240184
Ger, R.B., et al.: Comprehensive investigation on controlling for CT imaging variabilities in radiomics studies. Sci. Rep. 8(1), 13047 (2018). https://doi.org/10.1038/s41598-018-31509-z
Gillies, R.J., Kinahan, P.E., Hricak, H.: Radiomics: images are more than pictures, they are data. Radiology 278, 563–577 (2016). https://doi.org/10.1148/radiol.2015151169
van Griethuysen, J.J., et al.: Computational radiomics system to decode the radiographic phenotype. Cancer Res. 77(21), e104–e107 (2017). https://doi.org/10.1158/0008-5472.can-17-0339
Shafiq-ul Hassan, M., Latifi, K., Zhang, G., Ullah, G., Gillies, R., Moros, E.: Voxel size and gray level normalization of CT radiomic features in lung cancer. Sci. Rep. 8(1), 10545 (2018). https://doi.org/10.1038/s41598-018-28895-9
Shafiq-ul Hassan, M., et al.: Intrinsic dependencies of CT radiomic features on voxel size and number of gray levels. Med. Phys. 44(3), 1050–1062 (2017). https://doi.org/10.1002/mp.12123
Horvat, N., et al.: A primer on texture analysis in abdominal radiology. Abdom. Radiol. (NY) 47, 2972–2985 (2022). https://doi.org/10.1007/s00261-021-03359-3
Ibrahim, A., et al.: MaasPenn radiomics reproducibility score: a novel quantitative measure for evaluating the reproducibility of CT-based handcrafted radiomic features. Cancers (Basel) 14(7), 1599 (2022). https://doi.org/10.3390/cancers14071599
Isensee, F., Jaeger, P.F., Kohl, S.A.A., Petersen, J., Maier-Hein, K.H.: nnU-Net: a self-configuring method for deep learning-based biomedical image segmentation. Nat. Methods 18(2), 203–211 (2021). https://doi.org/10.1038/s41592-020-01008-z
Kikinis, R., Pieper, S.D., Vosburgh, K.G.: 3D slicer: a platform for subject-specific image analysis, visualization, and clinical support. In: Jolesz, F.A. (ed.) Intraoperative Imaging and Image-Guided Therapy, pp. 277–289. Springer, New York (2014). https://doi.org/10.1007/978-1-4614-7657-3_19
Kim, Y.J., Lee, H.J., Kim, K.G., Lee, S.H.: The effect of CT scan parameters on the measurement of CT radiomic features: a lung nodule phantom study. Comput. Math. Methods Med. 2019, 1–12 (2019). https://doi.org/10.1155/2019/8790694
Larue, R.T.H.M., et al.: Influence of gray level discretization on radiomic feature stability for different CT scanners, tube currents and slice thicknesses: a comprehensive phantom study. Acta Oncol. 56(11), 1544–1553 (2017). https://doi.org/10.1080/0284186x.2017.1351624
Ligero, M., et al.: Minimizing acquisition-related radiomics variability by image resampling and batch effect correction to allow for large-scale data analysis. Eur. Radiol. 31(3), 1460–1470 (2020). https://doi.org/10.1007/s00330-020-07174-0
Lin, L.I.K.: A concordance correlation coefficient to evaluate reproducibility. Biometrics 45(1), 255 (1989). https://doi.org/10.2307/2532051
Lu, L., Ehmke, R.C., Schwartz, L.H., Zhao, B.: Assessing agreement between radiomic features computed for multiple CT imaging settings. PLoS ONE 11(12), e0166550 (2016). https://doi.org/10.1371/journal.pone.0166550
Meyer, M., et al.: Reproducibility of CT radiomic features within the same patient: influence of radiation dose and CT reconstruction settings. Radiology 293(3), 583–591 (2019). https://doi.org/10.1148/radiol.2019190928
Park, S., et al.: Deep learning algorithm for reducing CT slice thickness: effect on reproducibility of radiomic features in lung cancer. Korean J. Radiol. 20(10), 1431 (2019). https://doi.org/10.3348/kjr.2019.0212
Perrin, T., et al.: Short-term reproducibility of radiomic features in liver parenchyma and liver malignancies on contrast-enhanced CT imaging. Abdom. Radiol. (NY) 43(12), 3271–3278 (2018). https://doi.org/10.1007/s00261-018-1600-6
Sanchez, L.E., Rundo, L., Gill, A.B., Hoare, M., Serrao, E.M., Sala, E.: Robustness of radiomic features in CT images with different slice thickness, comparing liver tumour and muscle. Sci. Rep. 11(1), 8262 (2021). https://doi.org/10.1038/s41598-021-87598-w
Shen, C., Liu, Z., Guan, M., Song, J., Lian, Y., Wang, S., Tang, Z., Dong, D., Kong, L., Wang, M., Shi, D., Tian, J.: 2D and 3D CT radiomics features prognostic performance comparison in non-small cell lung cancer. Transl. Oncol. 10(6), 886–894 (2017). https://doi.org/10.1016/j.tranon.2017.08.007
Simpson, A.L., et al.: Computed tomography image texture: a noninvasive prognostic marker of hepatic recurrence after hepatectomy for metastatic colorectal cancer. Ann. Surg. Oncol. 24(9), 2482–2490 (2017). https://doi.org/10.1245/s10434-017-5896-1
Simpson, A.L., et al.: Preoperative CT and survival data for patients undergoing resection of colorectal liver metastases (Colorectal-Liver-Metastases) (Version 2) [Data set]. The Cancer Imaging Archive (2023). https://doi.org/10.7937/QXK2-QG03
van Timmeren, J.E., et al.: Test-retest data for radiomics feature stability analysis: generalizable or study-specific? Tomography 2(4), 361–365 (2016). https://doi.org/10.18383/j.tom.2016.00208
Traverso, A., Wee, L., Dekker, A., Gillies, R.: Repeatability and reproducibility of radiomic features: a systematic review. Int. J. Radiat. Oncol. Biol. Phys. 102(4), 1143–1158 (2018). https://doi.org/10.1016/j.ijrobp.2018.05.053
Varghese, B.A., et al.: Reliability of CT-based texture features: phantom study. J. Appl. Clin. Med. Phys. 20(8), 155–163 (2019). https://doi.org/10.1002/acm2.12666
Yang, S., Wu, N., Zhang, L., Li, M.: Evaluation of the linear interpolation method in correcting the influence of slice thicknesses on radiomic feature values in solid pulmonary nodules: a prospective patient study. Ann. Transl. Med. 9(4), 279–279 (2021). https://doi.org/10.21037/atm-20-2992
Zhao, B.: Understanding sources of variation to improve the reproducibility of radiomics. Front. Oncol. 11, 826 (2021). https://doi.org/10.3389/fonc.2021.633176
Zhao, B., Tan, Y., Tsai, W.Y., Schwartz, L.H., Lu, L.: Exploring variability in CT characterization of tumors: a preliminary phantom study. Transl. Oncol. 7(1), 88–93 (2014). https://doi.org/10.1593/tlo.13865
Zwanenburg, A., Leger, S., Vallières, M., Löck, S.: Image biomarker standardisation initiative: Reference manual. arXiv:1612.07003 [cs.CV] (2016). https://doi.org/10.48550/ARXIV.1612.07003
Zwanenburg, A., et al.: The image biomarker standardization initiative: standardized quantitative radiomics for high-throughput image-based phenotyping. Radiology 295(2), 328–338 (2020). https://doi.org/10.1148/radiol.2020191145
Acknowledgements
This work was supported in part by National Institutes of Health grant R01 CA233888.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
1 Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Peoples, J.J. et al. (2023). Examining the Effects of Slice Thickness on the Reproducibility of CT Radiomics for Patients with Colorectal Liver Metastases. In: Sudre, C.H., Baumgartner, C.F., Dalca, A., Mehta, R., Qin, C., Wells, W.M. (eds) Uncertainty for Safe Utilization of Machine Learning in Medical Imaging. UNSURE 2023. Lecture Notes in Computer Science, vol 14291. Springer, Cham. https://doi.org/10.1007/978-3-031-44336-7_5
Download citation
DOI: https://doi.org/10.1007/978-3-031-44336-7_5
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-44335-0
Online ISBN: 978-3-031-44336-7
eBook Packages: Computer ScienceComputer Science (R0)