Abstract
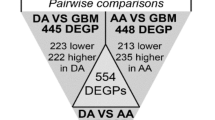
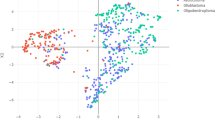
Glioma is a family of brain tumors with three main types exhibiting different progressions, which lack effective therapeutic options and specific molecular biomarkers. In this work, we propose a pipeline for multi-omics integrated analysis aimed at identifying features that could impact the development of different gliomas, assigned according to the latest classification guidelines. We estimate networks of genes and proteins based on human data, via the graphical lasso, as a network-based step towards variable selection. The estimated glioma networks were compared to disclose molecular relations that can be important for the development of a certain tumor type. Our outcomes were validated both mathematically, and through principal component analysis to determine if the selected subset of variables carries enough biological information to distinguish the three glioma types in a reduced dimensional subspace. The results highlight an overall agreement in variable selection across the two omics. Features exclusively selected by each glioma type appear as more representative of the pathological condition, making them suitable as potential diagnostic biomarkers. The comparison between glioma-type networks and with known protein-protein interactions reveals the presence of molecular relations that could be associated to a pathological condition. The 59 features identified by our analysis will be further considered to extend our work by integrating targeted biological evaluation.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
Notes
- 1.
DeltaCon distance was computed by a modified version of the delta_con R function (package rdsg) [6].
References
Brennan, C., et al.: The somatic genomic landscape of glioblastoma. Cell 155, 462–477 (2013). https://doi.org/10.1016/j.cell.2013.09.034
Dong, H., et al.: Proliferation, migration and invasion of pancreatic cancer cell lines. Cancers 15 (2023). https://doi.org/10.3390/cancers15041084
Friedman, J., Hastie, T., Tibshirani, R.: Sparse inverse covariance estimation with the graphical lasso. Biostatistics 9, 432–441 (2008). https://doi.org/10.1093/biostatistics/kxm045
Hawe, J.S., Theis, F.J., Heinig, M.: Inferring interaction networks from multi-omics data. Front. Genet. 10 (2019). https://doi.org/10.3389/fgene.2019.00535
Jarque, C.M.: Jarque-bera test. Int. Encycl. Stat. Sci. (2011)
Koutra, D., et al.: Deltacon: Principled massive-graph similarity function with attribution. ACM Trans. Knowl. Discov. Data 10 (2016). https://doi.org/10.1145/2824443
Louis, D.N., et al.: The 2021 WHO classification of tumors of the central nervous system: a summary. Neuro-Oncology 23, 1231–1251 (2021). https://doi.org/10.1093/neuonc/noab106
McLendon, R., et al.: Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455, 1061–1068 (2008). https://doi.org/10.1038/nature07385
Mendonça, M.L., et al.: Updating TCGA glioma classification through integration of molecular profiling data following the 2016 and 2021 WHO guidelines. bioRxiv p. 2023.02.19.529134 (2023). https://doi.org/10.1101/2023.02.19.529134
Qiao, L., Zhang, L., Chen, S., Shen, D.: Data-driven graph construction and graph learning: a review. Neurocomputing 312, 336–351 (2018). https://doi.org/10.1016/j.neucom.2018.05.084
Szklarczyk, D., Franceschini, A., Wyder, S., Forslund, K., et al.: String v10: protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 43, D447–D452 (2014)
Tantardini, M., Ieva, F., Tajoli, L., Piccardi, C.: Comparing methods for comparing networks. Sci. Rep. 9, 17557 (2019). https://doi.org/10.1038/s41598-019-53708-y
Tribius, S., Pidel, A., Casper, D.: Atm protein expression correlates with radioresistance in primary glioblastoma cells in culture. Int. J. Radiat. Oncol. Biol. Phys. 50, 511–523 (2001). https://doi.org/10.1016/S0360-3016(01)01489-4
TCGA, Network: Comprehensive, integrative genomic analysis of diffuse lower-grade gliomas. N. Engl. J. Med. 372, 2481–2498 (2015). https://doi.org/10.1056/NEJMoa1402121
Wang, D.: Discrepancy between mrna and protein abundance: Insight from information retrieval process in computers. Comput. Biol. Chem. 32, 462–468 (2008). https://doi.org/10.1016/j.compbiolchem.2008.07.014
Yang, J., Qianxue, C.: Expression of ku proteins in glioma and their association with radiotherapy sensitivity and prognosis. Acta Med. Mediterr. Int. Sci. J. Clin. Med. 4, 1065 (2018). https://doi.org/10.19193/0393-6384_2018_4_163
Yang, K., et al.: Glioma targeted therapy: insight into future of molecular approaches. Mol. Cancer 21, 39 (2022). https://doi.org/10.1186/s12943-022-01513-z
Zhao, T., et al.: The huge package for high-dimensional undirected graph estimation in r. J. Mach. Learn. Res.: JMLR 13, 1059–1062 (2012). https://doi.org/10.1093/nar/gku1003
Zhao, Z., Ju, Q., Ji, J., Li, Y., Zhao, Y.: N6-methyladenosine methylation regulator rbm15 is a potential prognostic biomarker and promotes cell proliferation in pancreatic adenocarcinoma. Front. Mol. Biosci. 9 (2022)
Acknowledgment
These results are based on data from the TCGA Research Network: https://www.cancer.gov/tcga.
This work is part of the MONET project PTDC/CCI-BIO/4180/2020, supported by FCT, with references CEECINST/00042/2021, UIDB/00297/2020, UIDP/00297/2020 (NOVA Math), UIDB/00667/2020 and UIDP/00667/2020 (UNIDEMI).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Coletti, R., Lopes, M.B. (2023). Multi-omics Data Integration and Network Inference for Biomarker Discovery in Glioma. In: Moniz, N., Vale, Z., Cascalho, J., Silva, C., Sebastião, R. (eds) Progress in Artificial Intelligence. EPIA 2023. Lecture Notes in Computer Science(), vol 14116. Springer, Cham. https://doi.org/10.1007/978-3-031-49011-8_20
Download citation
DOI: https://doi.org/10.1007/978-3-031-49011-8_20
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-49010-1
Online ISBN: 978-3-031-49011-8
eBook Packages: Computer ScienceComputer Science (R0)