Abstract
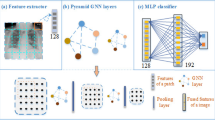
In recent years, the utilization of graph-based deep learning has gained prominence, yet its potential in the realm of medical diagnosis remains relatively unexplored. The challenge arises from the inherent irregular and unordered nature of physiological data, making it challenging to depict intricate patterns using conventional methods. Our research focuses on abnormality screening: classification of Chest X-Ray (CXR) as Tuberculosis positive or negative, using Graph Neural Networks (GNN) that uses Region Adjacency Graphs (RAGs) and each superpixel serves as a dedicated graph node. By integrating residual and concatenation structures, our approach effectively captures crucial features and relationships among superpixels, facilitating advancements in tuberculosis identification. Through the amalgamation of state-of-the-art neural network architectures and innovative graph-based representations, our work introduces a new perspective to medical image analysis.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Santosh, K.C., Allu, S., Rajaraman, S., Antani, S.: Advances in deep learning for tuberculosis screening using chest X-rays: the last 5 years review. J. Med. Syst. 46(11), 82 (2022)
Vajda, S., Karargyris, A., Jaeger, S., Santosh, K.C., Candemir, S., Xue, Z., Thoma, G.: Feature selection for automatic tuberculosis screening in frontal chest radiographs. J. Med. Syst. 42, 1–11 (2018)
Santosh, K.C., Antani, S.: Automated chest X-ray screening: can lung region symmetry help detect pulmonary abnormalities? IEEE Trans. Med. Imaging 37(5), 1168–1177 (2017)
Hirsh, A.E., Tsolaki, A.G., DeRiemer, K., Feldman, M.W., Small, P.M.: Stable association between strains of mycobacterium tuberculosis and their human host populations. Proc. Natl. Acad. Sci. 101(14), 4871–4876 (2004)
Mahbub, M.K., Biswas, M., Gaur, L., Alenezi, F., Santosh, K.C.: Deep features to detect pulmonary abnormalities in chest X-rays due to infectious diseaseX: Covid-19, pneumonia, and tuberculosis. Inf. Sci. 592, 389–401 (2022)
Ravimohan, S., Kornfeld, H., Weissman, D., Bisson, G.P.: Tuberculosis and lung damage: from epidemiology to pathophysiology. Eur. Respiratory Rev. 27(147) (2018)
Sharma, S.K., Mohan, A.: Tuberculosis: from an incurable scourge to a curable disease-journey over a millennium. Indian J. Med. Res. 137(3), 455 (2013)
Das, D., Santosh, K.C., Pal, U.: Cross-population train/test deep learning model: abnormality screening in chest x-rays. In: 2020 IEEE 33rd International Symposium on Computer-Based Medical Systems (CBMS), IEEE (2020)
Makkar, A., Santosh, K.C.: SecureFed: federated learning empowered medical imaging technique to analyze lung abnormalities in chest X-rays. Int. J. Mach. Learn. Cybern. 14, 1–12 (2023)
Rea, G., Sperandeo, M., Lieto, R., Bocchino, M., Quarato, C.M.I., Feragalli, B., Lacedonia, D.: Chest imaging in the diagnosis and management of pulmonary tuberculosis: the complementary role of thoraci ultrasound. Front. Med. 8, 753821 (2021)
Santosh, K.C., GhoshRoy, D., Nakarmi, S.: A systematic review on deep structured learning for covid-19 screening using chest CT from 2020 to 2022. In: Healthcare, vol. 11. no. 17, MDPI (2023)
Santosh, K.C., Vajda, S., Antani, S., Thoma, G.R.: Edge map analysis in chest X-rays for automatic pulmonary abnormality screening. Int. J. Comput. Assist. Radiol. Surg. 11, 1637–1646 (2016)
Oloko-Oba, M., Viriri, S.: A systematic review of deep learning techniques for tuberculosis detection from chest radiograph. Front. Med. 9, 830515 (2022)
Long, Jianwu. "A graph neural network for superpixel image classification. In: Journal of Physics: Conference Series, vol. 1871. no. 1. IOP Publishing (2021)
Bae, J., Yu, G., Lee, J., Vu, D., Anh, L., Kim, H., Kim, J.: Superpixel image classification with graph convolutional neural networks based on learnable positional embedding. Appl. Sci. 12(18), 9176 (2022)
Dwivedi, V.P., Luu, A.T., Laurent, T., Bengio, Y., Bresson, X.: Graph neural networks with learnable structural and positional representations. arXiv preprint arXiv:2110.07875 (2021)
Gori, M., Monfardini, G., Scarselli, F.: A new model for learning in graph domains. In: Proceedings, 2005 IEEE International Joint Conference on Neural Networks, vol. 2, IEEE (2005)
Bruna, J., Zaremba, W., Szlam, A., LeCun, Y.: Spectral networks and locally connected networks on graphs. arXiv preprint arXiv:1312.6203 (2013)
Kipf, T.N., Welling, M.: Semi-supervised classification with graph convolutional networks. arXiv preprint arXiv:1609.02907 (2016)
Shi, J., Malik, J.: Normalized cuts and image segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 22(8), 888–905 (2000)
Avelar, P.H.C., Tavares, A.R., Da Silveira, T.L.T., Jung, C.R., Lamb, L.C.: Superpixel image classification with graph attention networks. In: 2020 33rd SIBGRAPI Conference on Graphics, Patterns and Images (SIBGRAPI), IEEE (2020)
Stutz, D., Hermans, A., Leibe, B.: Superpixels: an evaluation of the state-of-the-art. Comput. Vis. Image Underst. 166, 1–27 (2018)
Monti, F., Boscaini, D., Masci, J., Rodolà, E., Svoboda, J., Bronstein, M. M.: Geometric deep learning on graphs and manifolds using mixture model CNNs. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (2017)
LeCun, Y.: The MNIST database of handwritten digits. https://yann.lecun.com/exdb/mnist/ (1998)
Fey, M., Lenssen, J. E., Weichert, F., Muller, H.: Splinecnn: fast geometric deep learning with continuous b-spline kernels. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (20180
Chaudhuri, U., Banerjee, B., Bhattacharya, A.: Siamese graph convolutional network for content based remote sensing image retrieval. Comput. Vis. Image Underst. 184, 22–30 (2019)
Knyazev, B., Taylor, G.W., Amer, M.: Understanding attention and generalization in graph neural networks. In: Advances in Neural Information Processing Systems, vol. 32 (2019)
Keyulu, X., Weihua, H., Jure, L., Stefanie, J.: How powerful are graph neural networks? In: International Conference on Learning Representations (ICLR) (2019)
Knyazev, B., Lin, X., Amer, M.R., Taylor, G.W.: Image classification with hierarchical multigraph networks. arXiv preprint arXiv:1907.09000 (2019)
B. P. Health. Belarus Tuberculosis Portal (2020). Accessed 9 Jun 2020. https://tuberculosis.by/
Achanta, R., Shaji, A., Smith, K., Lucchi, A., Fua, P., Süsstrunk, S.: SLIC superpixels compared to state-of-the-art superpixel methods. IEEE Trans. Pattern Anal. Mach. Intell. 34(11), 2274–2282 (2012)
Defferrard, M., Bresson, X., Vandergheynst, P.: Convolutional neural networks on graphs with fast localized spectral filtering. In: Advances in Neural Information Processing Systems, vol. 29 (2016)
Veličković, P., Cucurull, G., Casanova, A., Romero, A., Liò, P., Bengio, Y.: Graph attention networks. arXiv preprint arXiv:1710.10903 (2017)
Santosh, K.C.: AI-driven tools for coronavirus outbreak: need of active learning and cross-population train/test models on multitudinal/multimodal data. J. Med. Syst. 44(5), 1–5 (2020). https://doi.org/10.1007/s10916-020-01562-1
Nakarmi, S. and Santosh, K.C.: Active learning to minimize the risk from future epidemics. In: 2023 IEEE Conference on Artificial Intelligence (CAI), IEEE (2023)
Zhang, W., et al.:. Information gain propagation: a new way to graph active learning with soft labels. arXiv preprint arXiv:2203.01093 (2022)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2024 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Pradhan, R., Santosh, K. (2024). Analyzing Pulmonary Abnormality with Superpixel Based Graph Neural Network in Chest X-Ray. In: Santosh, K., et al. Recent Trends in Image Processing and Pattern Recognition. RTIP2R 2023. Communications in Computer and Information Science, vol 2027. Springer, Cham. https://doi.org/10.1007/978-3-031-53085-2_9
Download citation
DOI: https://doi.org/10.1007/978-3-031-53085-2_9
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-53084-5
Online ISBN: 978-3-031-53085-2
eBook Packages: Computer ScienceComputer Science (R0)