Abstract
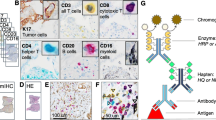
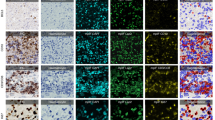
Immunohistochemistry (IHC) staining is a widely used technique in the diagnosis of abnormal cells such as cancer. For instance, it can be used to determine the distribution and localization of the differentially expressed biomarkers of immune cells (such as T-cells or B-cells) in cancerous tissue for an immune response study. Typically, the immunological data of interest includes the type, density and location of the immune cells within the tumor samples; this data is of particular interest to pathologists for accurate patient survival prediction. However, to manually count each subset of immune cells under a bright-field microscope for each piece of IHC stained tissue is usually extremely tedious and time consuming. This makes automatic detection very attractive, but it can be very challenging due to the wide variety of cell appearances resulting from different tissue types, block cuttings, and staining processes. This paper presents a novel method for automatic immune cell counting on digitally scanned images of IHC stained slides. The method first uses a sparse color unmixing technique to separate the IHC image into multiple color channels that correspond to different cell structures. Since the immune cell biomarkers that we are interested in are membrane markers, the detection problem is formulated into a deep learning framework using the membrane image channel. The algorithm is evaluated on a clinical data set containing a large number of IHC slides and demonstrates more effective detection than the existing technique and the result is also in accordance with the human observer’s output.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Preview
Unable to display preview. Download preview PDF.
Similar content being viewed by others
References
Galon, J., et al.: Type, Density, and Location of Immune Cells Within Human Colorectal Tumors Predict Clinical Outcome. Science 313(5795), 1960–1964 (2006)
Parvin, B., et al.: Iterative Voting for Inference of Structural Saliency and Characterization of Subcellular Events. IEEE Trans. Image Processing 16(3), 615–623 (2007)
Xin, Q., et al.: Iterative Voting for Inference of Structural Saliency and Characterization of Subcellular Events. IEEE Trans. Biomedical Engineering 59(3), 754–765 (2011)
Arteta, C., Lempitsky, V., Noble, J.A., Zisserman, A.: Learning to Detect Cells Using Non-overlapping Extremal Regions. In: Ayache, N., Delingette, H., Golland, P., Mori, K. (eds.) MICCAI 2012, Part I. LNCS, vol. 7510, pp. 348–356. Springer, Heidelberg (2012)
Mualla, F., et al.: Automatic Cell Detection in Bright-Field Microscope Images Using SIFT, Random Forests, and Hierarchical Clustering. IEEE Trans. Medical Imaging 32(12), 2274–2286 (2013)
Niazi, M.K.K., et al.: An Automated Method for Counting Cytotoxic T-cells from CD8 Stained Images of Renal Biopsies. In: SPIE, vol. 8676 (2013)
LeCun, Y., et al.: Gradient-based Learning Applied to Document Recognition. Proceedings of the IEEE 86(11), 2278–2324 (1998)
Cireşan, D.C., Giusti, A., Gambardella, L.M., Schmidhuber, J.: Mitosis Detection in Breast Cancer Histology Images with Deep Neural Networks. In: Mori, K., Sakuma, I., Sato, Y., Barillot, C., Navab, N. (eds.) MICCAI 2013, Part II. LNCS, vol. 8150, pp. 411–418. Springer, Heidelberg (2013)
Ruifrok, A.C., et al.: Quantification of Histochemical Staining by Color Deconvolution. Anal. Quant. Cytol. Histol. 23, 291–299 (2001)
Kesheva, N.: A Survey of Spectral Unmixing Algorithms. Lincoln Laboratory Journal 14(1), 55–78 (2003)
Lindeberg, T.: Edge Detection and Ridge Detection with Automatic Scale Selection. In: CVPR, pp. 465–470 (1996)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer International Publishing Switzerland
About this paper
Cite this paper
Chen, T., Chefd’hotel, C. (2014). Deep Learning Based Automatic Immune Cell Detection for Immunohistochemistry Images. In: Wu, G., Zhang, D., Zhou, L. (eds) Machine Learning in Medical Imaging. MLMI 2014. Lecture Notes in Computer Science, vol 8679. Springer, Cham. https://doi.org/10.1007/978-3-319-10581-9_3
Download citation
DOI: https://doi.org/10.1007/978-3-319-10581-9_3
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-10580-2
Online ISBN: 978-3-319-10581-9
eBook Packages: Computer ScienceComputer Science (R0)