Abstract
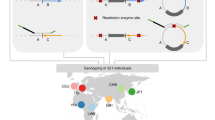
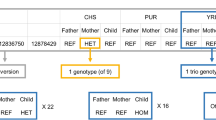
Structural rearrangements, including copy-number alterations and inversions, are increasingly recognized as an important contributor to human genetic variation. Copy number variants are readily measured via array-based techniques like comparative genomic hybridization, but copy-neutral variants such as inversion polymorphisms remain difficult to identify without whole genome sequencing. We introduce a method to identify inversion polymorphisms and estimate their frequency in a population using readily available single nucleotide polymorphism (SNP) data. Our method uses a probabilistic model to describe a population as a mixture of forward and inverted chromosomes and identifies putative inversions by characteristic differences in haplotype frequencies around inversion breakpoints. On simulated data, our method accurately predicts inversions with frequencies as low as 25% in the population and reliably estimates inversion frequencies over a wide range. On the human HapMap Phase 2 data, we predict between 88 and 142 inversion polymorphisms with frequency ranging from 20 to 92 percent. Many of these correspond to known inversions or have other evidence supporting them, and the predicted inversion frequencies largely agree with the limited information presently available.
Preview
Unable to display preview. Download preview PDF.
Similar content being viewed by others
References
Frazer, K., et al.: A second generation human haplotype map of over 3.1 million SNPs. Nature 449, 851–861 (2007)
Wellcome Trust Case Control Consortium: Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007)
Sharp, A., Cheng, Z., Eichler, E.: Structural variation of the human genome. Annu. Rev. Genomics Hum. Genet. 7, 407–442 (2006)
Walsh, T., McClellan, J., McCarthy, S., Addington, A., Pierce, S., Cooper, G., Nord, A., Kusenda, M., Malhotra, D., Bhandari, A., Stray, S., Rippey, C., Roccanova, P., Makarov, V., Lakshmi, B., Findling, R., Sikich, L., Stromberg, T., Merriman, B., Gogtay, N., Butler, P., Eckstrand, K., Noory, L., Gochman, P., Long, R., Chen, Z., Davis, S., Baker, C., Eichler, E., Meltzer, P., Nelson, S., Singleton, A., Lee, M., Rapoport, J., King, M., Sebat, J.: Rare structural variants disrupt multiple genes in neurodevelopmental pathways in schizophrenia. Science 320, 539–543 (2008)
Stefansson, H., Helgason, A., Thorleifsson, G., Steinthorsdottir, V., Masson, G., Barnard, J., Baker, A., Jonasdottir, A., Ingason, A., Gudnadottir, V., Desnica, N., Hicks, A., Gylfason, A., Gudbjartsson, D., Jonsdottir, G., Sainz, J., Agnarsson, K., Birgisdottir, B., Ghosh, S., Olafsdottir, A., Cazier, J., Kristjansson, K., Frigge, M., Thorgeirsson, T., Gulcher, J., Kong, A., Stefansson, K.: A common inversion under selection in Europeans. Nat. Genet. 37, 129–137 (2005)
Perry, G., Dominy, N., Claw, K., Lee, A., Fiegler, H., Redon, R., Werner, J., Villanea, F., Mountain, J., Misra, R., Carter, N., Lee, C., Stone, A.: Diet and the evolution of human amylase gene copy number variation. Nat. Genet. 39, 1256–1260 (2007)
Cooper, G., Zerr, T., Kidd, J., Eichler, E., Nickerson, D.: Systematic assessment of copy number variant detection via genome-wide SNP genotyping. Nat. Genet. 40, 1199–1203 (2008)
McCarroll, S., Kuruvilla, F., Korn, J., Cawley, S., Nemesh, J., Wysoker, A., Shapero, M., de Bakker, P., Maller, J., Kirby, A., Elliott, A., Parkin, M., Hubbell, E., Webster, T., Mei, R., Veitch, J., Collins, P., Handsaker, R., Lincoln, S., Nizzari, M., Blume, J., Jones, K., Rava, R., Daly, M., Gabriel, S., Altshuler, D.: Integrated detection and population-genetic analysis of SNPs and copy number variation. Nat. Genet. 40, 1166–1174 (2008)
Perry, G., Ben-Dor, A., Tsalenko, A., Sampas, N., Rodriguez-Revenga, L., Tran, C., Scheffer, A., Steinfeld, I., Tsang, P., Yamada, N., Park, H., Kim, J., Seo, J., Yakhini, Z., Laderman, S., Bruhn, L., Lee, C.: The fine-scale and complex architecture of human copy-number variation. Am. J. Hum. Genet. 82, 685–695 (2008)
McCarroll, S., Hadnott, T., Perry, G., Sabeti, P., Zody, M., Barrett, J., Dallaire, S., Gabriel, S., Lee, C., Daly, M., Altshuler, D.: Common deletion polymorphisms in the human genome. Nat. Genet. 38, 86–92 (2006)
Conrad, D., Andrews, T., Carter, N., Hurles, M., Pritchard, J.: A high-resolution survey of deletion polymorphism in the human genome. Nat. Genet. 38, 75–81 (2006)
Corona, E., Raphael, B., Eskin, E.: Identification of deletion polymorphisms from haplotypes. In: Speed, T., Huang, H. (eds.) RECOMB 2007. LNCS (LNBI), vol. 4453, pp. 354–365. Springer, Heidelberg (2007)
Tuzun, E., Sharp, A.J., Bailey, J.A., Kaul, R., Morrison, V.A., Pertz, L.M., Haugen, E., Hayden, H., Albertson, D., Pinkel, D., Olson, M.V., Eichler, E.E.: Fine-scale structural variation of the human genome. Nat. Genet. 37, 727–732 (2005)
Korbel, J.O., Urban, A.E., Affourtit, J.P., Godwin, B., Grubert, F., Simons, J.F., Kim, P.M., Palejev, D., Carriero, N.J., Du, L., Taillon, B.E., Chen, Z., Tanzer, A., Saunders, A.C.E., Chi, J., Yang, F., Carter, N.P., Hurles, M.E., Weissman, S.M., Harkins, T.T., Gerstein, M.B., Egholm, M., Snyder, M.: Paired-end mapping reveals extensive structural variation in the human genome. Science 318(5849), 420–426 (2007)
Kidd, J.M., Cooper, G.M., Donahue, W.F., Hayden, H.S., Sampas, N., Graves, T., Hansen, N., Teague, B., Alkan, C., Antonacci, F., Haugen, E., Zerr, T., Yamada, N.A., Tsang, P., Newman, T.L., Tüzün, E., Cheng, Z., Ebling, H.M., Tusneem, N., David, R., Gillett, W., Phelps, K.A., Weaver, M., Saranga, D., Brand, A., Tao, W., Gustafson, E., McKernan, K., Chen, L., Malig, M., Smith, J.D., Korn, J.M., McCarroll, S.A., Altshuler, D.A., Peiffer, D.A., Dorschner, M., Stamatoyannopoulos, J., Schwartz, D., Nickerson, D.A., Mullikin, J.C., Wilson, R.K., Bruhn, L., Olson, M.V., Kaul, R., Smith, D.R., Eichler, E.E.: Mapping and sequencing of structural variation from eight human genomes. Nature 453, 56–64 (2008)
Levy, S., Sutton, G., Ng, P., Feuk, L., Halpern, A., Walenz, B., Axelrod, N., Huang, J., Kirkness, E., Denisov, G., Lin, Y., MacDonald, J., Pang, A., Shago, M., Stockwell, T., Tsiamouri, A., Bafna, V., Bansal, V., Kravitz, S., Busam, D., Beeson, K., McIntosh, T., Remington, K., Abril, J., Gill, J., Borman, J., Rogers, Y., Frazier, M., Scherer, S., Strausberg, R., Venter, J.: The diploid genome sequence of an individual human. PLoS Biol. 5, e254 (2007)
Wheeler, D., Srinivasan, M., Egholm, M., Shen, Y., Chen, L., McGuire, A., He, W., Chen, Y., Makhijani, V., Roth, G., Gomes, X., Tartaro, K., Niazi, F., Turcotte, C., Irzyk, G., Lupski, J., Chinault, C., Song, X., Liu, Y., Yuan, Y., Nazareth, L., Qin, X., Muzny, D., Margulies, M., Weinstock, G., Gibbs, R., Rothberg, J.: The complete genome of an individual by massively parallel DNA sequencing. Nature 452, 872–876 (2008)
Iafrate, A., Feuk, L., Rivera, M., Listewnik, M., Donahoe, P., Qi, Y., Scherer, S., Lee, C.: Detection of large-scale variation in the human genome. Nat. Genet. 36, 949–951 (2004)
Feuk, L., MacDonald, J., Tang, T., Carson, A., Li, M., Rao, G., Khaja, R., Scherer, S.: Discovery of human inversion polymorphisms by comparative analysis of human and chimpanzee DNA sequence assemblies. PLoS Genet. 1, e56 (2005)
Chaisson, M., Raphael, B., Pevzner, P.: Microinversions in mammalian evolution. Proc. Natl. Acad. Sci. U.S.A. 103, 19824–19829 (2006)
Kirkpatrick, M., Barton, N.: Chromosome inversions, local adaptation and speciation. Genetics 173, 419–434 (2006)
Hoffmann, A., Sgrò, C., Weeks, A.: Chromosomal inversion polymorphisms and adaptation. Trends Ecol. Evol. (Amst.) 19, 482–488 (2004)
Bansal, V., Bashir, A., Bafna, V.: Evidence for large inversion polymorphisms in the human genome from HapMap data. Genome Res. 17, 219–230 (2007)
Patil, N., Berno, A., Hinds, D., Barrett, W., Doshi, J., Hacker, C., Kautzer, C., Lee, D., Marjoribanks, C., McDonough, D., Nguyen, B., Norris, M., Sheehan, J., Shen, N., Stern, D., Stokowski, R., Thomas, D., Trulson, M., Vyas, K., Frazer, K., Fodor, S., Cox, D.: Blocks of limited haplotype diversity revealed by high-resolution scanning of human chromosome 21. Science 294, 1719–1723 (2001)
Daly, M., Rioux, J., Schaffner, S., Hudson, T., Lander, E.: High-resolution haplotype structure in the human genome. Nat. Genet. 29, 229–232 (2001)
Pritchard, J., Stephens, M., Donnelly, P.: Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000)
Price, A., Patterson, N., Plenge, R., Weinblatt, M., Shadick, N., Reich, D.: Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006)
Sridhar, S., Rao, S., Halperin, E.: An efficient and accurate graph-based approach to detect population substructure. In: Speed, T., Huang, H. (eds.) RECOMB 2007. LNCS (LNBI), vol. 4453, pp. 503–517. Springer, Heidelberg (2007)
Dempster, A.P., Laird, N.M., Rubin, D.B.: Maximum likelihood from incomplete data via the EM algorithm. J. Roy. Statist. Soc. Ser. B 39(1), 1–38 (1977)
Falush, D., Stephens, M., Pritchard, J.: Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164, 1567–1587 (2003)
Schaffner, S.F., Foo, C., Gabriel, S., Reich, D., Daly, M.J., Altshuler, D.: Calibrating a coalescent simulation of human genome sequence variation. Genome Res. 15(11), 1576–1583 (2005)
Sindi, S.S., Helman, E., Bashir, A., Raphael, B.J.: A geometric approach for classification and comparison of structural variants. In: Bioinformatics. Proc. ISMB/ECCB 2009 (in press, 2009)
Koolen, D., Vissers, L., Pfundt, R., de Leeuw, N., Knight, S., Regan, R., Kooy, R., Reyniers, E., Romano, C., Fichera, M., Schinzel, A., Baumer, A., Anderlid, B., Schoumans, J., Knoers, N., van Kessel, A., Sistermans, E., Veltman, J., Brunner, H., de Vries, B.: A new chromosome 17q21.31 microdeletion syndrome associated with a common inversion polymorphism. Nat. Genet. 38, 999–1001 (2006)
Zhang, K., Deng, M., Chen, T., Waterman, M., Sun, F.: A dynamic programming algorithm for haplotype block partitioning. Proc. Natl. Acad. Sci. U.S.A. 99, 7335–7339 (2002)
Anderson, E., Novembre, J.: Finding haplotype block boundaries by using the minimum-description-length principle. Am. J. Hum. Genet. 73, 336–354 (2003)
Wang, N., Akey, J., Zhang, K., Chakraborty, R., Jin, L.: Distribution of recombination crossovers and the origin of haplotype blocks: the interplay of population history, recombination, and mutation. Am. J. Hum. Genet. 71, 1227–1234 (2002)
Kimmel, G., Shamir, R.: GERBIL: Genotype resolution and block identification using likelihood. Proc. Natl. Acad. Sci. U.S.A. 102, 158–162 (2005)
1000 Genomes Project. Technical report (2008), http://www.1000genomes.org
Feuk, L., Carson, A.R., Scherer, S.W.: Structural variation in the human genome. Nat. Rev. Genet. 7(2), 85–97 (2006)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2009 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Sindi, S.S., Raphael, B.J. (2009). Identification and Frequency Estimation of Inversion Polymorphisms from Haplotype Data. In: Batzoglou, S. (eds) Research in Computational Molecular Biology. RECOMB 2009. Lecture Notes in Computer Science(), vol 5541. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-02008-7_30
Download citation
DOI: https://doi.org/10.1007/978-3-642-02008-7_30
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-02007-0
Online ISBN: 978-3-642-02008-7
eBook Packages: Computer ScienceComputer Science (R0)