Abstract
The investigation of network dynamics is a major issue in systems and synthetic biology. One of the essential steps in a dynamics investigation is the parameter estimation in the model that expresses biological phenomena. Indeed, various techniques for parameter optimization have been devised and implemented in both free and commercial software. While the computational time for parameter estimation has been greatly reduced, due to improvements in calculation algorithms and the advent of high performance computers, the accuracy of parameter estimation has not been addressed.
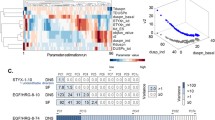
We previously proposed an approach for accurate parameter optimization by using Differential Elimination, which is an algebraic approach for rewriting a system of differential equations into another equivalent system. The equivalent system has the same solution as the original system, and it includes high-order derivatives, which contain information about the form of the observed time-series data. The introduction of an equivalent system into the numerical parameter optimizing procedure resulted in the drastic improvement of the estimation accuracy, since our approach evaluates the difference of not only the values but also the forms between the measured and estimated data, while the classical numerical approach evaluates only the value difference. In this report, we describe the detailed procedure of our approach for accurate parameter estimation in dynamic systems. The ability of our approach is illustrated in terms of the parameter estimation accuracy, in comparison with classical methods.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Preview
Unable to display preview. Download preview PDF.
Similar content being viewed by others
References
Kitano, H.: System Biology: A Brief Overview. Science 295(5560), 1662–1664 (2002)
Nocedal, J., Wright, S.J.: Numerical Optimization. Springer, New York (1999)
Denis-Vidal, L., Joly-Blanchard, G., Noiret, C.: System identifiability (symbolic computation) and parameter estimation (numerical computation). Numerical Algorithms 34, 282–292 (2003)
Boulier, F., Denis-Vidal, F., Henin, T., Lemaire, F.: LÉPISME. In: Proceedings of the ICPSS Conference (2004), http://hal.archives-ouvertes.fr/hal-00140368
Ritt, J.F.: Differential Algebra. Dover Publications Inc., New York (1950)
Kolchin, E.R.: Differential Algebra and Algebraic Groups. Academic Press, New York (1973)
Seidenberg, A.: An elimination theory for differential algebra. Univ. California Publ. Math. (New Series) 3, 31–65 (1956)
Wu, W.T.: On the foundation of algebraic differential geometry. Mechanization of Mathematics, Research Preprints 3, 2–27 (1989)
Boulier, F., Lazard, D., Ollivier, F., Petitot, M.: Representation for the radical of a finitely generated differential ideal. In: Proceedings of ISSAC 1995, pp. 158–166 (1995)
Boulier, F., Lazard, D., Ollivier, F., Petitot, M.: Computing representations for radicals of finitely generated differential ideals. Journal of AAECC 20(1), 73–121 (2009); (1997 Techrep. IT306 of the LIFL)
Boulier, F.: The BLAD libraries (2004), http://www.lifl.fr/~boulier/BLAD
Boulier, F.: Differential Elimination and Biological Modeling. Johann Radon Institute for Computational and Applied Mathematics (RICAM) Book Series 2, 111–139 (2007)
Boulier, F., Lemaire, F.: Differential Algebra and System Modeling in Cellular Biology. In: Horimoto, K., Regensburger, G., Rosenkranz, M., Yoshida, H. (eds.) AB 2008. LNCS, vol. 5147, pp. 22–39. Springer, Heidelberg (2008)
Nakatsui, M., Horimoto, K.: Parameter Optimization in the network dynamics including unmeasured variables by the symbolic-numeric approach. In: Proc. of the Third International Symposium on Optimization and Systems Biology (OSB 2009), pp. 245–253 (2009)
Nakatsui, M., Horimoto, K.: Improvement of Estimation Accuracy in Parameter Optimization by Symbolic Computation. In: Proceedings of IEEE Multi-Conference on Systems and Control (in press)
Nakatsui, M., Horimoto, K., Okamoto, M., Tokumoto, Y., Miyake, J.: Parameter Optimization by Using Differential Elimination: a General Approach for Introducing Constraints into Objective Functions. BMC Systems Biology (in press)
Nakatsui, M., Horimoto, K., Lemaire, F., Ürgüplü, A., Sedoglavic, F., Boulier, F.: Brute force meets Bruno force in parameter optimization: Introduction of novel constraints for parameter accuracy improvement by symbolic computation. IET Systems Biology (in press)
Powell, M.J.D.: An efficient method for finding the minimum of a function of several variables without calculating derivatives. Computer Journal 7, 142–162 (1954)
Powell, M.J.D.: On the calculation of orthogonal vectors. Computer Journal 11, 302–304 (1968)
Holland, J.H.: Adaptation in Natural and Artificial Systems. The University of Michigan Press, Ann Arbor (1975)
Goldberg, D.D.: Genetic Algorithms in Search, Optimization and Machine Learning. Addison-Wesley Longman Publishing Co., Inc., Boston (1989)
Eberhart, R., Kennedy, J.: A New Optimizer Using Particle Swarm Theory. In: Proc. of Sixth International Symposium on Micro Machine and Human Science (Nagoya Japan), pp. 39–43. IEEE Service Center, Piscataway (1995)
Kennedy, J., Eberhart, R.: Particle swarm optimization. In: Proc. IEEE International Conference on Neural Networks (Perth, Australia), pp. IV:1942– IV:1948. IEEE Service Center, Piscataway (1995)
Jonikow, C.Z., Michalewicz, Z.: An Experimental Comparison of Binary and Floating Point Representations in Genetic Algorithms. In: Proceedings of the Fourth International Conference on Genetic Algorithms, pp. 31–36 (1991)
Ono, I., Kobayashi, S.: A real-coded genetic algorithm for function optimization using unimodal distribution crossover. In: Proc 7th ICGA, pp. 249–253 (1997)
Satoh, H., Ono, I., Kobayashi, S.: A new generation alternation model of genetic algorithm and its assessment. J. of Japanese Society for Artificial Intelligence 15(2), 743–744 (1997)
Novák, B., Tyson, J.J.: Design principles of biochemical oscillators. Nat. Rev. Mol. Cell Biol. 9(12), 981–991 (2008)
Kwon, Y.K., Cho, K.H.: Quantitative analysis of robustness and fragility in biological networks based on feedback dynamics. Bioinformatics 24(7), 987–994 (2008)
Tyson, J.J., Chen, K.C., Novák, B.: Sniffers, buzzers, toggles and blinkers: dynamics of regulatory and signaling pathways in the cell. Curr. Opin. Cell. Biol. 15(2), 221–231 (2003)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2012 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Nakatsui, M., Sedoglavic, A., Lemaire, F., Boulier, F., Ürgüplü, A., Horimoto, K. (2012). A General Procedure for Accurate Parameter Estimation in Dynamic Systems Using New Estimation Errors. In: Horimoto, K., Nakatsui, M., Popov, N. (eds) Algebraic and Numeric Biology. Lecture Notes in Computer Science, vol 6479. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-28067-2_9
Download citation
DOI: https://doi.org/10.1007/978-3-642-28067-2_9
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-28066-5
Online ISBN: 978-3-642-28067-2
eBook Packages: Computer ScienceComputer Science (R0)