Abstract
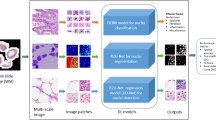
Pathologists generally diagnose whether or not cervical cancer cells have the potential to spread to other organs and assess the malignancy of cancer through whole slide histopathology images using virtual microscopy. In this process, the morphology of nuclei is one of the significant diagnostic indices, including the size, the orientation and arrangement of the nuclei. Therefore, accurate segmentation of nuclei is a crucial step in clinical diagnosis. However, several challenges exist, namely a single whole slide image (WSI) often occupies a large amount of memory, making it difficult to manipulate. More than that, due to the extremely high density and variant shapes, sizes and overlapping nuclei, as well as low contrast, weakly defined boundaries, different staining methods and image acquisition techniques, it is difficult to achieve accurate segmentation. A method is proposed, comprised of two main parts to achieve lesion localization and automatic segmentation of nuclei. Initially, a U-Net model was used to localize and segment lesions. Then, a multi-task cascade network was proposed to combine nuclei foreground and edge information to obtain instance segmentation results. Evaluation of the proposed method for lesion localization and nuclei segmentation using a dataset comprised of cervical tissue sections collected by experienced pathologists along with comparative experiments, demonstrates the outstanding performance of this method.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others

References
Mcguire, S.: World cancer report 2014. Geneva, Switzerland: world health organization, international agency for research on cancer, WHO Press, 2015. Adv. Nutr. 7(2), 418 (2016)
Canavan, T.P., Doshi, N.R.: Cervical cancer. Am. Fam. Physician 61(5), 1369 (2000)
LeCun, Y.: http://yann.lecun.com/exdb/lenet/. Accessed 16 Oct 2013
Saltzer, J.H.: End-to-end arguments in system design. ACM Trans. Comput. Syst. (TOCS) 2(4), 277–288 (1984)
Shelhamer, E., Long, J., Darrell, T.: Fully convolutional networks for semantic segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 39(4), 640–651 (2014)
Badrinarayanan, V., Kendall, A., Cipolla, R.: SegNet: a deep convolutional encoder-decoder architecture for image segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 12(39), 2481–2495 (2017)
Zheng, S., Jayasumana, S., Romera-Paredes, B., Vineet, V., Su, Z., Du, D., et al.: Conditional random fields as recurrent neural networks. In: IEEE International Conference on Computer Vision (ICCV), pp. 1529–1537 (2015)
Chen, L.C., Papandreou, G., Kokkinos, I., et al.: DeepLab: semantic image segmentation with deep convolutional nets, atrous convolution, and fully connected CRFs. IEEE Trans. Pattern Anal. Mach. Intell. 40(4), 834–848 (2018)
Ronneberger, O., Fischer, P., Brox, T.: U-Net: convolutional networks for biomedical image segmentation. In: Navab, N., Hornegger, J., Wells, W.M., Frangi, A.F. (eds.) MICCAI 2015. LNCS, vol. 9351, pp. 234–241. Springer, Cham (2015). https://doi.org/10.1007/978-3-319-24574-4_28
Chen, H., Qi, X., Yu, L., Heng, P.A.: DCAN: deep contour-aware networks for accurate gland segmentation. In: IEEE Conference on Computer Vision and Pattern Recognition (CVPR), pp. 2487–2496 (2016)
Dai, J., He, K., Sun, J.: Instance-aware semantic segmentation via multi-task network cascades. In: IEEE Conference on Computer Vision and Pattern Recognition (CVPR), pp. 3150–3158 (2015)
Li, Y., Qi, H., Dai, J., Ji, X., Wei, Y.: Fully convolutional instance-aware semantic segmentation. In: IEEE Conference on Computer Vision and Pattern Recognition (CVPR), pp. 4438–4446 (2017)
He, K., Gkioxari, G., Dollár, P., et al.: Mask R-CNN. In: IEEE International Conference on Computer Vision (ICCV), pp. 2980–2988 (2017)
Dai, J., Li, Y., He, K., et al.: R-FCN: Object detection via region-based fully convolutional networks. Advances in Neural Information Processing Systems 29 (NIPS) (2016)
Vincent, P., Larochelle, H., Lajoie, I., Bengio, Y., Manzagol, P.A.: Stacked denoising autoencoders: learning useful representations in a deep network with a local denoising criterion. J. Mach. Learn. Res. 11(12), 3371–3408 (2010)
Cho, K., Van Merrienboer, B., Gulcehre, C., Bahdanau, D., Bougares, F., Schwenk, H.: Learning phrase representations using RNN encoder-decoder for statistical machine translation. Computer Science (2014)
Acknowledgements
This work is supported by National Key Scientific Instruments and Equipment Development Program of China (2013YQ03065101) and partially supported by National Natural Science Foundation (NNSF) of China under Grant 61503243 and National Science Foundation (NSF) of China under the Grant 61521063.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Singapore Pte Ltd.
About this paper
Cite this paper
Yang, Q. et al. (2019). Cervical Nuclei Segmentation in Whole Slide Histopathology Images Using Convolution Neural Network. In: Yap, B., Mohamed, A., Berry, M. (eds) Soft Computing in Data Science. SCDS 2018. Communications in Computer and Information Science, vol 937. Springer, Singapore. https://doi.org/10.1007/978-981-13-3441-2_8
Download citation
DOI: https://doi.org/10.1007/978-981-13-3441-2_8
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-13-3440-5
Online ISBN: 978-981-13-3441-2
eBook Packages: Computer ScienceComputer Science (R0)